Key Points
-
Recent progress in high-throughput sequencing has uncovered an astounding landscape of small RNAs in eukaryotic cells. Various small RNAs of distinctive characteristics have been found and can be classified into three classes based on their biogenesis mechanism and the type of Argonaute (Ago) protein that they are associated with.
-
MicroRNAs (miRNAs) are generated from local hairpin structures by the action of two RNase III-type proteins, Drosha and Dicer. Mature ∼22-nucleotide (nt) miRNAs are then bound by Ago-subfamily proteins. miRNAs target mRNAs and thereby function as post-transcriptional regulators.
-
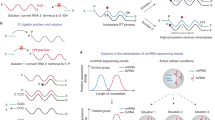
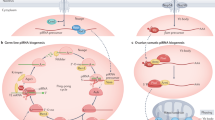
Piwi-interacting RNAs (piRNAs), which are 24–31 nt in length, are associated with Piwi-subfamily proteins. The biogenesis of piRNAs does not depend on Dicer. At least some piRNAs are involved in transposon silencing through heterochromatin formation or RNA destabilization.
-
Endogenous small interfering RNAs (endo-siRNAs), such as miRNAs, associate with Ago-subfamily proteins. However, endo-siRNAs differ from miRNAs in that they are derived from long double-stranded RNAs and are dependent only on Dicer but not on Drosha. They are also slightly shorter (∼21 nt) than miRNAs. At least some of the endo-siRNAs have been shown to function as post-transcriptional regulators that target RNAs.
-
There are numerous other small RNAs that are generated through non-canonical pathways. Many of them are difficult to classify and their biogenesis pathways remain poorly understood, but they may have species-specific functions that are not yet fully appreciated.
Abstract
Small RNAs of 20–30 nucleotides can target both chromatin and transcripts, and thereby keep both the genome and the transcriptome under extensive surveillance. Recent progress in high-throughput sequencing has uncovered an astounding landscape of small RNAs in eukaryotic cells. Various small RNAs of distinctive characteristics have been found and can be classified into three classes based on their biogenesis mechanism and the type of Argonaute protein that they are associated with: microRNAs (miRNAs), endogenous small interfering RNAs (endo-siRNAs or esiRNAs) and Piwi-interacting RNAs (piRNAs). This Review summarizes our current knowledge of how these intriguing molecules are generated in animal cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Lee, R. C., Feinbaum, R. L. & Ambros, V. The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 75, 843–854 (1993).
Wightman, B., Ha, I. & Ruvkun, G. Posttranscriptional regulation of the heterochronic gene lin-14 by lin-4 mediates temporal pattern formation in C. elegans. Cell 75, 855–862 (1993).
Lau, N. C., Lim, L. P., Weinstein, E. G. & Bartel, D. P. An abundant class of tiny RNAs with probable regulatory roles in Caenorhabditis elegans. Science 294, 858–862 (2001).
Lee, R. C. & Ambros, V. An extensive class of small RNAs in Caenorhabditis elegans. Science 294, 862–864 (2001).
Lagos-Quintana, M., Rauhut, R., Lendeckel, W. & Tuschl, T. Identification of novel genes coding for small expressed RNAs. Science 294, 853–858 (2001).
Lu, C. et al. Elucidation of the small RNA component of the transcriptome. Science 309, 1567–1569 (2005).
Margulies, M. et al. Genome sequencing in microfabricated high-density picolitre reactors. Nature 437, 376–380 (2005).
Lai, E. C., Tomancak, P., Williams, R. W. & Rubin, G. M. Computational identification of Drosophila microRNA genes. Genome Biol. 4, R42 (2003).
Nam, J. W. et al. Human microRNA prediction through a probabilistic co-learning model of sequence and structure. Nucleic Acids Res. 33, 3570–3581 (2005).
Li, S. C., Pan, C. Y. & Lin, W. C. Bioinformatic discovery of microRNA precursors from human ESTs and introns. BMC Genomics 7, 164 (2006).
Huang, T. H. et al. MiRFinder: an improved approach and software implementation for genome-wide fast microRNA precursor scans. BMC Bioinformatics 8, 341 (2007).
Chu, C. Y. & Rana, T. M. Small RNAs: regulators and guardians of the genome. J. Cell Physiol. 213, 412–419 (2007).
Filipowicz, W., Bhattacharyya, S. N. & Sonenberg, N. Mechanisms of post-transcriptional regulation by microRNAs: are the answers in sight? Nature Rev. Genet. 9, 102–114 (2008).
Vagin, V. V. et al. A distinct small RNA pathway silences selfish genetic elements in the germline. Science 313, 320–324 (2006). The first paper to describe that piRNAs are produced in a Dicer-independent manner. The authors also reported that D. melanogaster piRNAs are modified at their 3′ ends.
Watanabe, T. et al. Endogenous siRNAs from naturally formed dsRNAs regulate transcripts in mouse oocytes. Nature 453, 539–543 (2008).
Tam, O. H. et al. Pseudogene-derived small interfering RNAs regulate gene expression in mouse oocytes. Nature 453, 534–538 (2008). References 15 and 16 contributed to the identification of many endo-siRNAs in mouse oocytes and also to the proposal of the functions of pseudogenes in silencing the parental genes.
Babiarz, J. E., Ruby, J. G., Wang, Y., Bartel, D. P. & Blelloch, R. Mouse ES cells express endogenous shRNAs, siRNAs, and other Microprocessor-independent, Dicer-dependent small RNAs. Genes Dev. 22, 2773–2785 (2008).
Okamura, K. et al. The Drosophila hairpin RNA pathway generates endogenous short interfering RNAs. Nature 453, 803–806 (2008).
Kawamura, Y. et al. Drosophila endogenous small RNAs bind to Argonaute 2 in somatic cells. Nature 453, 793–797 (2008).
Czech, B. et al. An endogenous small interfering RNA pathway in Drosophila. Nature 453, 798–802 (2008).
Jones-Rhoades, M. W., Bartel, D. P. & Bartel, B. MicroRNAs and their regulatory roles in plants. Annu. Rev. Plant Biol. 57, 19–53 (2006).
Mallory, A. C. & Vaucheret, H. Functions of microRNAs and related small RNAs in plants. Nature Genet. 38, S31–S36 (2006).
Zhang, B., Pan, X., Cobb, G. P. & Anderson, T. A. Plant microRNA: a small regulatory molecule with big impact. Dev. Biol. 289, 3–16 (2006).
Verdel, A. & Moazed, D. RNAi-directed assembly of heterochromatin in fission yeast. FEBS Lett. 579, 5872–5878 (2005).
Kim, V. N. MicroRNA biogenesis: coordinated cropping and dicing. Nature Rev. Mol. Cell Biol. 6, 376–385 (2005).
Kim, D. H., Saetrom, P., Snove, O. Jr. & Rossi, J. J. MicroRNA-directed transcriptional gene silencing in mammalian cells. Proc. Natl Acad. Sci. USA 105, 16230–16235 (2008).
Grimson, A. et al. Early origins and evolution of microRNAs and Piwi-interacting RNAs in animals. Nature 455, 1193–1197 (2008).
Ibanez-Ventoso, C., Vora, M. & Driscoll, M. Sequence relationships among C. elegans, D. melanogaster and human microRNAs highlight the extensive conservation of microRNAs in biology. PLoS ONE 3, e2818 (2008).
Chapman, E. J. & Carrington, J. C. Specialization and evolution of endogenous small RNA pathways. Nature Rev. Genet. 8, 884–896 (2007).
Millar, A. A. & Waterhouse, P. M. Plant and animal microRNAs: similarities and differences. Funct. Integr. Genomics 5, 129–135 (2005).
Ventura, A. et al. Targeted deletion reveals essential and overlapping functions of the miR-17 through 92 family of miRNA clusters. Cell 132, 875–886 (2008).
Lee, Y., Jeon, K., Lee, J. T., Kim, S. & Kim, V. N. MicroRNA maturation: stepwise processing and subcellular localization. EMBO J. 21, 4663–4670 (2002).
Lee, Y. et al. MicroRNA genes are transcribed by RNA polymerase II. EMBO J. 23, 4051–4060 (2004).
Cai, X., Hagedorn, C. H. & Cullen, B. R. Human microRNAs are processed from capped, polyadenylated transcripts that can also function as mRNAs. RNA 10, 1957–1966 (2004).
Borchert, G. M., Lanier, W. & Davidson, B. L. RNA polymerase III transcribes human microRNAs. Nature Struct. Mol. Biol. 13, 1097–1101 (2006).
Lee, Y. S. & Dutta, A. MicroRNAs in cancer. Annu. Rev. Pathol. 25 Sep 2008 (doi:10.1146annurev.pathol.4.110807.092222).
Lee, Y. et al. The nuclear RNase III Drosha initiates microRNA processing. Nature 425, 415–419 (2003).
Han, J. et al. The Drosha–DGCR8 complex in primary microRNA processing. Genes Dev. 18, 3016–3027 (2004).
Denli, A. M., Tops, B. B., Plasterk, R. H., Ketting, R. F. & Hannon, G. J. Processing of primary microRNAs by the Microprocessor complex. Nature 432, 231–235 (2004).
Gregory, R. I. et al. The Microprocessor complex mediates the genesis of microRNAs. Nature 432, 235–240 (2004).
Landthaler, M., Yalcin, A. & Tuschl, T. The human DiGeorge syndrome critical region gene 8 and its D. melanogaster homolog are required for miRNA biogenesis. Curr. Biol. 14, 2162–2167 (2004).
Wang, Y., Medvid, R., Melton, C., Jaenisch, R. & Blelloch, R. DGCR8 is essential for microRNA biogenesis and silencing of embryonic stem cell self-renewal. Nature Genet. 39, 380–385 (2007).
Filippov, V., Solovyev, V., Filippova, M. & Gill, S. S. A novel type of RNase III family proteins in eukaryotes. Gene 245, 213–221 (2000).
Wu, H., Xu, H., Miraglia, L. J. & Crooke, S. T. Human RNase III is a 160-kDa protein involved in preribosomal RNA processing. J. Biol. Chem. 275, 36957–36965 (2000).
Fortin, K. R., Nicholson, R. H. & Nicholson, A. W. Mouse ribonuclease III. cDNA structure, expression analysis, and chromosomal location. BMC Genomics 3, 26 (2002).
Han, J. et al. Molecular basis for the recognition of primary microRNAs by the Drosha–DGCR8 complex. Cell 125, 887–901 (2006).
Zeng, Y. & Cullen, B. R. Efficient processing of primary microRNA hairpins by Drosha requires flanking nonstructured RNA sequences. J. Biol. Chem. 280, 27595–27603 (2005).
Kim, Y. K. & Kim, V. N. Processing of intronic microRNAs. EMBO J. 26, 775–783 (2007).
Morlando, M. et al. Primary microRNA transcripts are processed co-transcriptionally. Nature Struct. Mol. Biol. 15, 902–909 (2008).
Pawlicki, J. M. & Steitz, J. A. Primary microRNA transcript retention at sites of transcription leads to enhanced microRNA production. J. Cell Biol. 182, 61–76 (2008).
Dye, M. J., Gromak, N. & Proudfoot, N. J. Exon tethering in transcription by RNA polymerase II. Mol. Cell 21, 849–859 (2006).
Han, J. et al. Posttranscriptional crossregulation between Drosha and DGCR8. Cell 136, 75–84 (2009).
Ruby, J. G., Jan, C. H. & Bartel, D. P. Intronic microRNA precursors that bypass Drosha processing. Nature 448, 83–86 (2007).
Okamura, K., Hagen, J. W., Duan, H., Tyler, D. M. & Lai, E. C. The mirtron pathway generates microRNA-class regulatory RNAs in Drosophila. Cell 130, 89–100 (2007).
Berezikov, E., Chung, W. J., Willis, J., Cuppen, E. & Lai, E. C. Mammalian mirtron genes. Mol. Cell 28, 328–336 (2007).
Ender, C. et al. A human snoRNA with microRNA-like functions. Mol. Cell 32, 519–528 (2008).
Kim, V. N. MicroRNA precursors in motion: exportin-5 mediates their nuclear export. Trends Cell Biol. 14, 156–159 (2004).
Lund, E., Guttinger, S., Calado, A., Dahlberg, J. E. & Kutay, U. Nuclear export of microRNA precursors. Science 303, 95–98 (2004).
Yi, R., Doehle, B. P., Qin, Y., Macara, I. G. & Cullen, B. R. Overexpression of exportin 5 enhances RNA interference mediated by short hairpin RNAs and microRNAs. RNA 11, 220–226 (2005).
Bohnsack, M. T., Czaplinski, K. & Gorlich, D. Exportin 5 is a RanGTP-dependent dsRNA-binding protein that mediates nuclear export of pre-miRNAs. RNA 10, 185–191 (2004).
Bohnsack, M. T. et al. Exp5 exports eEF1A via tRNA from nuclei and synergizes with other transport pathways to confine translation to the cytoplasm. EMBO J. 21, 6205–6215 (2002).
Calado, A., Treichel, N., Muller, E. C., Otto, A. & Kutay, U. Exportin-5-mediated nuclear export of eukaryotic elongation factor 1A and tRNA. EMBO J. 21, 6216–6224 (2002).
Gwizdek, C. et al. Exportin-5 mediates nuclear export of minihelix-containing RNAs. J. Biol. Chem. 278, 5505–5508 (2003).
Basyuk, E., Suavet, F., Doglio, A., Bordonne, R. & Bertrand, E. Human let-7 stem-loop precursors harbor features of RNase III cleavage products. Nucleic Acids Res. 31, 6593–6597 (2003).
Zeng, Y. & Cullen, B. R. Structural requirements for pre-microRNA binding and nuclear export by Exportin 5. Nucleic Acids Res. 32, 4776–4785 (2004).
Bernstein, E., Caudy, A. A., Hammond, S. M. & Hannon, G. J. Role for a bidentate ribonuclease in the initiation step of RNA interference. Nature 409, 363–366 (2001).
Grishok, A. et al. Genes and mechanisms related to RNA interference regulate expression of the small temporal RNAs that control C. elegans developmental timing. Cell 106, 23–34 (2001).
Hutvagner, G. et al. A cellular function for the RNA-interference enzyme Dicer in the maturation of the let-7 small temporal RNA. Science 293, 834–838 (2001).
Ketting, R. F. et al. Dicer functions in RNA interference and in synthesis of small RNA involved in developmental timing in C. elegans. Genes Dev. 15, 2654–2659 (2001).
Knight, S. W. & Bass, B. L. A role for the RNase III enzyme DCR-1 in RNA interference and germ line development in Caenorhabditis elegans. Science 293, 2269–2271 (2001).
Lee, Y. S. et al. Distinct roles for Drosophila Dicer-1 and Dicer-2 in the siRNA/miRNA silencing pathways. Cell 117, 69–81 (2004).
Xie, Z. et al. Genetic and functional diversification of small RNA pathways in plants. PLoS Biol. 2, e104 (2004).
Forstemann, K. et al. Normal microRNA maturation and germ-line stem cell maintenance requires Loquacious, a double-stranded RNA-binding domain protein. PLoS Biol. 3, e236 (2005).
Jiang, F. et al. Dicer-1 and R3D1-L catalyze microRNA maturation in Drosophila. Genes Dev. 19, 1674–1679 (2005).
Saito, K., Ishizuka, A., Siomi, H. & Siomi, M. C. Processing of pre-microRNAs by the Dicer-1–Loquacious complex in Drosophila cells. PLoS Biol. 3, e235 (2005).
Chendrimada, T. P. et al. TRBP recruits the Dicer complex to Ago2 for microRNA processing and gene silencing. Nature 436, 740–744 (2005).
Haase, A. D. et al. TRBP, a regulator of cellular PKR and HIV-1 virus expression, interacts with Dicer and functions in RNA silencing. EMBO Rep. 6, 961–967 (2005).
Lee, Y. et al. The role of PACT in the RNA silencing pathway. EMBO J. 25, 522–532 (2006).
Garcia, M. A., Meurs, E. F. & Esteban, M. The dsRNA protein kinase PKR: virus and cell control. Biochimie 89, 799–811 (2007).
Aza-Blanc, P. et al. Identification of modulators of TRAIL-induced apoptosis via RNAi-based phenotypic screening. Mol. Cell 12, 627–637 (2003).
Schwarz, D. S. et al. Asymmetry in the assembly of the RNAi enzyme complex. Cell 115, 199–208 (2003).
Khvorova, A., Reynolds, A. & Jayasena, S. D. Functional siRNAs and miRNAs exhibit strand bias. Cell 115, 209–216 (2003).
Maniataki, E. & Mourelatos, Z. A human, ATP-independent, RISC assembly machine fueled by pre-miRNA. Genes Dev. 19, 2979–2990 (2005).
MacRae, I. J., Ma, E., Zhou, M., Robinson, C. V. & Doudna, J. A. In vitro reconstitution of the human RISC-loading complex. Proc. Natl Acad. Sci. USA 105, 512–517 (2008).
Gregory, R. I., Chendrimada, T. P., Cooch, N. & Shiekhattar, R. Human RISC couples microRNA biogenesis and posttranscriptional gene silencing. Cell 123, 631–640 (2005).
Tomari, Y., Matranga, C., Haley, B., Martinez, N. & Zamore, P. D. A protein sensor for siRNA asymmetry. Science 306, 1377–1380 (2004).
Preall, J. B. & Sontheimer, E. J. RNAi: RISC gets loaded. Cell 123, 543–545 (2005).
Liu, Q. et al. R2D2, a bridge between the initiation and effector steps of the Drosophila RNAi pathway. Science 301, 1921–1925 (2003).
Matranga, C., Tomari, Y., Shin, C., Bartel, D. P. & Zamore, P. D. Passenger-strand cleavage facilitates assembly of siRNA into Ago2-containing RNAi enzyme complexes. Cell 123, 607–620 (2005).
Miyoshi, K., Tsukumo, H., Nagami, T., Siomi, H. & Siomi, M. C. Slicer function of Drosophila Argonautes and its involvement in RISC formation. Genes Dev. 19, 2837–2848 (2005).
Leuschner, P. J., Ameres, S. L., Kueng, S. & Martinez, J. Cleavage of the siRNA passenger strand during RISC assembly in human cells. EMBO Rep. 7, 314–320 (2006).
Diederichs, S. & Haber, D. A. Dual role for argonautes in microRNA processing and posttranscriptional regulation of microRNA expression. Cell 131, 1097–1108 (2007).
Tomari, Y., Du, T. & Zamore, P. D. Sorting of Drosophila small silencing RNAs. Cell 130, 299–308 (2007).
Forstemann, K., Horwich, M. D., Wee, L., Tomari, Y. & Zamore, P. D. Drosophila microRNAs are sorted into functionally distinct argonaute complexes after production by dicer-1. Cell 130, 287–297 (2007).
Steiner, F. A. et al. Structural features of small RNA precursors determine Argonaute loading in Caenorhabditis elegans. Nature Struct. Mol. Biol. 14, 927–933 (2007).
Liu, J. et al. Argonaute2 is the catalytic engine of mammalian RNAi. Science 305, 1437–1441 (2004).
Meister, G. et al. Human Argonaute2 mediates RNA cleavage targeted by miRNAs and siRNAs. Mol. Cell 15, 185–197 (2004).
Azuma-Mukai, A. et al. Characterization of endogenous human Argonautes and their miRNA partners in RNA silencing. Proc. Natl Acad. Sci. USA 105, 7964–7969 (2008).
Seitz, H., Ghildiyal, M. & Zamore, P. D. Argonaute loading improves the 5′ precision of both microRNAs and their miRNA strands in flies. Curr. Biol. 18, 147–151 (2008).
Landgraf, P. et al. A mammalian microRNA expression atlas based on small RNA library sequencing. Cell 129, 1401–1414 (2007).
Jiang, Q. et al. miR2Disease: a manually curated database for microRNA deregulation in human disease. Nucleic Acids Res. 37, D98–D104 (2008).
Chen, J. F. et al. The role of microRNA-1 and microRNA-133 in skeletal muscle proliferation and differentiation. Nature Genet. 38, 228–233 (2006).
Kim, H. K., Lee, Y. S., Sivaprasad, U., Malhotra, A. & Dutta, A. Muscle-specific microRNA miR-206 promotes muscle differentiation. J. Cell Biol. 174, 677–687 (2006).
Rao, P. K., Kumar, R. M., Farkhondeh, M., Baskerville, S. & Lodish, H. F. Myogenic factors that regulate expression of muscle-specific microRNAs. Proc. Natl Acad. Sci. USA 103, 8721–8726 (2006).
He, L., He, X., Lowe, S. W. & Hannon, G. J. microRNAs join the p53 network — another piece in the tumour-suppression puzzle. Nature Rev. Cancer 7, 819–822 (2007).
He, L. et al. A microRNA polycistron as a potential human oncogene. Nature 435, 828–833 (2005).
Chang, T. C. et al. Widespread microRNA repression by Myc contributes to tumorigenesis. Nature Genet. 40, 43–50 (2008).
Bueno, M. J. et al. Genetic and epigenetic silencing of microRNA-203 enhances ABL1 and BCR-ABL1 oncogene expression. Cancer Cell 13, 496–506 (2008).
Davis, B. N., Hilyard, A. C., Lagna, G. & Hata, A. SMAD proteins control DROSHA-mediated microRNA maturation. Nature 454, 56–61 (2008).
Guil, S. & Caceres, J. F. The multifunctional RNA-binding protein hnRNP A1 is required for processing of miR-18a. Nature Struct. Mol. Biol. 14, 591–596 (2007).
Michlewski, G., Guil, S., Semple, C. A. & Caceres, J. F. Posttranscriptional regulation of miRNAs harboring conserved terminal loops. Mol. Cell 32, 383–393 (2008).
Pasquinelli, A. E. et al. Conservation of the sequence and temporal expression of let-7 heterochronic regulatory RNA. Nature 408, 86–89 (2000).
Suh, M. R. et al. Human embryonic stem cells express a unique set of microRNAs. Dev. Biol. 270, 488–498 (2004).
Thomson, J. M. et al. Extensive post-transcriptional regulation of microRNAs and its implications for cancer. Genes Dev. 20, 2202–2207 (2006).
Wulczyn, F. G. et al. Post-transcriptional regulation of the let-7 microRNA during neural cell specification. FASEB J. 21, 415–426 (2007).
Newman, M. A., Thomson, J. M. & Hammond, S. M. Lin-28 interaction with the let-7 precursor loop mediates regulated microRNA processing. RNA 14, 1539–1549 (2008).
Viswanathan, S. R., Daley, G. Q. & Gregory, R. I. Selective blockade of microRNA processing by Lin28. Science 320, 97–100 (2008).
Rybak, A. et al. A feedback loop comprising lin-28 and let-7 controls pre-let-7 maturation during neural stem-cell commitment. Nature Cell Biol. 10, 987–993 (2008).
Heo, I. et al. Lin28 mediates the terminal uridylation of let-7 precursor microRNA. Mol. Cell 32, 276–284 (2008).
Balzer, E. & Moss, E. G. Localization of the developmental timing regulator Lin28 to mRNP complexes, P-bodies and stress granules. RNA Biol. 4, 16–25 (2007).
Hwang, H. W., Wentzel, E. A. & Mendell, J. T. A hexanucleotide element directs microRNA nuclear import. Science 315, 97–100 (2007).
Kennedy, S., Wang, D. & Ruvkun, G. A conserved siRNA-degrading RNase negatively regulates RNA interference in C. elegans. Nature 427, 645–649 (2004).
Ramachandran, V. & Chen, X. Degradation of microRNAs by a family of exoribonucleases in Arabidopsis. Science 321, 1490–1492 (2008).
Yang, W. et al. Modulation of microRNA processing and expression through RNA editing by ADAR deaminases. Nature Struct. Mol. Biol. 13, 13–21 (2006).
Kawahara, Y., Zinshteyn, B., Chendrimada, T. P., Shiekhattar, R. & Nishikura, K. RNA editing of the microRNA-151 precursor blocks cleavage by the Dicer–TRBP complex. EMBO Rep. 8, 763–769 (2007).
Kawahara, Y. et al. Frequency and fate of microRNA editing in human brain. Nucleic Acids Res. 36, 5270–5280 (2008).
Obernosterer, G., Leuschner, P. J., Alenius, M. & Martinez, J. Post-transcriptional regulation of microRNA expression. RNA 12, 1161–1167 (2006).
Lee, E. J. et al. Systematic evaluation of microRNA processing patterns in tissues, cell lines, and tumors. RNA 14, 35–42 (2008).
Michael, M. Z., O'Connor, S. M., van Holst Pellekaan, N. G., Young, G. P. & James, R. J. Reduced accumulation of specific microRNAs in colorectal neoplasia. Mol. Cancer Res. 1, 882–891 (2003).
Kefas, B. et al. microRNA-7 inhibits the epidermal growth factor receptor and the Akt pathway and is down-regulated in glioblastoma. Cancer Res. 68, 3566–3572 (2008).
Martinez, N. J. et al. A C. elegans genome-scale microRNA network contains composite feedback motifs with high flux capacity. Genes Dev. 22, 2535–2549 (2008).
Tokumaru, S., Suzuki, M., Yamada, H., Nagino, M. & Takahashi, T. let-7 regulates Dicer expression and constitutes a negative feedback loop. Carcinogenesis 29, 2073–2077 (2008).
Forman, J. J., Legesse-Miller, A. & Coller, H. A. A search for conserved sequences in coding regions reveals that the let-7 microRNA targets Dicer within its coding sequence. Proc. Natl Acad. Sci. USA 105, 14879–14884 (2008).
Piskounova, E. et al. Determinants of microRNA processing inhibition by the developmentally regulated RNA-binding protein Lin28. J. Biol. Chem. 283, 21310–21314 (2008).
Bracken, C. P. et al. A double-negative feedback loop between ZEB1–SIP1 and the microRNA-200 family regulates epithelial–mesenchymal transition. Cancer Res. 68, 7846–7854 (2008).
Aravin, A. A. et al. Double-stranded RNA-mediated silencing of genomic tandem repeats and transposable elements in the D. melanogaster germline. Curr. Biol. 11, 1017–1027 (2001).
Aravin, A. A. et al. The small RNA profile during Drosophila melanogaster development. Dev. Cell 5, 337–350 (2003). This was the herald of studies for a novel class of small RNAs of 24–29 nt (rasiRNAs or piRNAs) that are specifically expressed in germ lines.
Cox, D. N. et al. A novel class of evolutionarily conserved genes defined by piwi are essential for stem cell self-renewal. Genes Dev. 12, 3715–3727 (1998).
Cox, D. N., Chao, A. & Lin, H. piwi encodes a nucleoplasmic factor whose activity modulates the number and division rate of germline stem cells. Development 127, 503–514 (2000).
Szakmary, A., Cox, D. N., Wang, Z. & Lin, H. Regulatory relationship among piwi, pumilio, and bag-of-marbles in Drosophila germline stem cell self-renewal and differentiation. Curr. Biol. 15, 171–178 (2005).
Kalmykova, A. I., Klenov, M. S. & Gvozdev, V. A. Argonaute protein PIWI controls mobilization of retrotransposons in the Drosophila male germline. Nucleic Acids Res. 33, 2052–2059 (2005).
Sarot, E., Payen-Groschene, G., Bucheton, A. & Pelisson, A. Evidence for a piwi-dependent RNA silencing of the gypsy endogenous retrovirus by the Drosophila melanogaster flamenco gene. Genetics 166, 1313–1321 (2004).
Harris, A. N. & Macdonald, P. M. aubergine encodes a Drosophila polar granule component required for pole cell formation and related to eIF2C. Development 128, 2823–2832 (2001).
Vagin, V. V. et al. The RNA interference proteins and vasa locus are involved in the silencing of retrotransposons in the female germline of Drosophila melanogaster. RNA Biol. 1, 54–58 (2004).
Saito, K. et al. Specific association of Piwi with rasiRNAs derived from retrotransposon and heterochromatic regions in the Drosophila genome. Genes Dev. 20, 2214–2222 (2006).
Brennecke, J. et al. Discrete small RNA-generating loci as master regulators of transposon activity in Drosophila. Cell 128, 1089–1103 (2007).
Gunawardane, L. S. et al. A slicer-mediated mechanism for repeat-associated siRNA 5′ end formation in Drosophila. Science 315, 1587–1590 (2007). The ping-pong pathway for piRNA biogenesis was proposed on the basis of these D. melanogaster studies.
Nishida, K. M. et al. Gene silencing mechanisms mediated by Aubergine–piRNA complexes in Drosophila male gonad. RNA 13, 1911–1922 (2007). Shows that Suppressor of Stellate piRNAs are directly associated with AUB in D. melanogaster testes and that the complex can slice the Stellate transcripts.
Aravin, A. et al. A novel class of small RNAs bind to MILI protein in mouse testes. Nature 442, 203–207 (2006).
Girard, A., Sachidanandam, R., Hannon, G. J. & Carmell, M. A. A germline-specific class of small RNAs binds mammalian Piwi proteins. Nature 442, 199–202 (2006).
Grivna, S. T., Beyret, E., Wang, Z. & Lin, H. A novel class of small RNAs in mouse spermatogenic cells. Genes Dev. 20, 1709–1714 (2006).
Watanabe, T. et al. Identification and characterization of two novel classes of small RNAs in the mouse germline: retrotransposon-derived siRNAs in oocytes and germline small RNAs in testes. Genes Dev. 20, 1732–1743 (2006).
Houwing, S. et al. A role for Piwi and piRNAs in germ cell maintenance and transposon silencing in zebrafish. Cell 129, 69–82 (2007).
Carmell, M. A. et al. MIWI2 is essential for spermatogenesis and repression of transposons in the mouse male germline. Dev. Cell 12, 503–514 (2007).
Aravin, A. A., Sachidanandam, R., Girard, A., Fejes-Toth, K. & Hannon, G. J. Developmentally regulated piRNA clusters implicate MILI in transposon control. Science 316, 744–747 (2007).
Kuramochi-Miyagawa, S. et al. DNA methylation of retrotransposon genes is regulated by Piwi family members MILI and MIWI2 in murine fetal testes. Genes Dev. 22, 908–917 (2008).
Zamore, P. D. RNA silencing: genomic defence with a slice of pi. Nature 446, 864–865 (2007).
Pane, A., Wehr, K. & Schupbach, T. zucchini and squash encode two putative nucleases required for rasiRNA production in the Drosophila germline. Dev. Cell 12, 851–862 (2007).
Lim, A. K. & Kai, T. Unique germ-line organelle, nuage, functions to repress selfish genetic elements in Drosophila melanogaster. Proc. Natl Acad. Sci. USA 104, 6714–6719 (2007).
Findley, S. D., Tamanaha, M., Clegg, N. J. & Ruohola-Baker, H. Maelstrom, a Drosophila spindle-class gene, encodes a protein that colocalizes with Vasa and RDE1/AGO1 homolog, Aubergine, in nuage. Development 130, 859–871 (2003).
Aravin, A. A., Hannon, G. J. & Brennecke, J. The Piwi–piRNA pathway provides an adaptive defense in the transposon arms race. Science 318, 761–764 (2007).
Brennecke, J. et al. An epigenetic role for maternally inherited piRNAs in transposon silencing. Science 322, 1387–1392 (2008).
Lau, N. C. et al. Characterization of the piRNA complex from rat testes. Science 313, 363–367 (2006).
Aravin, A. A. et al. A piRNA pathway primed by individual transposons is linked to de novo DNA methylation in mice. Mol. Cell 31, 785–799 (2008).
Saito, K., Sakaguchi, Y., Suzuki, T., Siomi, H. & Siomi, M. C. Pimet, the Drosophila homolog of HEN1, mediates 2′-O-methylation of Piwi-interacting RNAs at their 3′ ends. Genes Dev. 21, 1603–1608 (2007).
Horwich, M. D. et al. The Drosophila RNA methyltransferase, DmHen1, modifies germline piRNAs and single-stranded siRNAs in RISC. Curr. Biol. 17, 1265–1272 (2007).
Kirino, Y. & Mourelatos, Z. Mouse Piwi-interacting RNAs are 2′-O-methylated at their 3′ termini. Nature Struct. Mol. Biol. 14, 347–348 (2007).
Ohara, T., Sakaguchi, Y., Suzuki, T., Ueda, H. & Miyauchi, K. The 3′ termini of mouse Piwi-interacting RNAs are 2′-O-methylated. Nature Struct. Mol. Biol. 14, 349–350 (2007).
Yu, B. et al. Methylation as a crucial step in plant microRNA biogenesis. Science 307, 932–935 (2005).
Ghildiyal, M. et al. Endogenous siRNAs derived from transposons and mRNAs in Drosophila somatic cells. Science 320, 1077–1081 (2008).
Chung, W. J., Okamura, K., Martin, R. & Lai, E. C. Endogenous RNA interference provides a somatic defense against Drosophila transposons. Curr. Biol. 18, 795–802 (2008).
Okamura, K., Balla, S., Martin, R., Liu, N. & Lai, E. C. Two distinct mechanisms generate endogenous siRNAs from bidirectional transcription in Drosophila melanogaster. Nature Struct. Mol. Biol. 15, 581–590 (2008).
van Rij, R. P. et al. The RNA silencing endonuclease Argonaute 2 mediates specific antiviral immunity in Drosophila melanogaster. Genes Dev. 20, 2985–2995 (2006).
Wang, X. H. et al. RNA interference directs innate immunity against viruses in adult Drosophila. Science 312, 452–454 (2006).
Nishikura, K. Editor meets silencer: crosstalk between RNA editing and RNA interference. Nature Rev. Mol. Cell Biol. 7, 919–931 (2006).
Fukuda, T. et al. DEAD-box RNA helicase subunits of the Drosha complex are required for processing of rRNA and a subset of microRNAs. Nature Cell Biol. 9, 604–611 (2007).
Shiohama, A., Sasaki, T., Noda, S., Minoshima, S. & Shimizu, N. Nucleolar localization of DGCR8 and identification of eleven DGCR8-associated proteins. Exp. Cell Res. 313, 4196–4207 (2007).
Yu, B. et al. The FHA domain proteins DAWDLE in Arabidopsis and SNIP1 in humans act in small RNA biogenesis. Proc. Natl Acad. Sci. USA 105, 10073–10078 (2008).
Neumuller, R. A. et al. Mei-P26 regulates microRNAs and cell growth in the Drosophila ovarian stem cell lineage. Nature 454, 241–245 (2008).
Krol, J. et al. Ribonuclease dicer cleaves triplet repeat hairpins into shorter repeats that silence specific targets. Mol. Cell 25, 575–586 (2007).
Ma, J. B., Ye, K. & Patel, D. J. Structural basis for overhang-specific small interfering RNA recognition by the PAZ domain. Nature 429, 318–322 (2004).
Lingel, A., Simon, B., Izaurralde, E. & Sattler, M. Nucleic acid 3′-end recognition by the Argonaute2 PAZ domain. Nature Struct. Mol. Biol. 11, 576–577 (2004).
Wang, Y. et al. Structure of an argonaute silencing complex with a seed-containing guide DNA and target RNA duplex. Nature 456, 921–926 (2008).
Wang, Y., Sheng, G., Juranek, S., Tuschl, T. & Patel, D. J. Structure of the guide-strand-containing argonaute silencing complex. Nature 456, 209–213 (2008).
Song, J. J., Smith, S. K., Hannon, G. J. & Joshua-Tor, L. Crystal structure of Argonaute and its implications for RISC slicer activity. Science 305, 1434–1437 (2004).
Parker, J. S., Roe, S. M. & Barford, D. Structural insights into mRNA recognition from a PIWI domain–siRNA guide complex. Nature 434, 663–666 (2005).
Ma, J. B. et al. Structural basis for 5′-end-specific recognition of guide RNA by the A. fulgidus Piwi protein. Nature 434, 666–670 (2005).
Brower-Toland, B. et al. Drosophila PIWI associates with chromatin and interacts directly with HP1a. Genes Dev. 21, 2300–2311 (2007).
Blaszczyk, J. et al. Crystallographic and modeling studies of RNase III suggest a mechanism for double-stranded RNA cleavage. Structure 9, 1225–1236 (2001).
Zhang, H., Kolb, F. A., Jaskiewicz, L., Westhof, E. & Filipowicz, W. Single processing center models for human Dicer and bacterial RNase III. Cell 118, 57–68 (2004).
Yeom, K. H., Lee, Y., Han, J., Suh, M. R. & Kim, V. N. Characterization of DGCR8/Pasha, the essential cofactor for Drosha in primary miRNA processing. Nucleic Acids Res. 34, 4622–4629 (2006).
Sohn, S. Y. et al. Crystal structure of human DGCR8 core. Nature Struct. Mol. Biol. 14, 847–853 (2007).
Faller, M., Matsunaga, M., Yin, S., Loo, J. A. & Guo, F. Heme is involved in microRNA processing. Nature Struct. Mol. Biol. 14, 23–29 (2007).
Song, J. J. et al. The crystal structure of the Argonaute2 PAZ domain reveals an RNA binding motif in RNAi effector complexes. Nature Struct. Biol. 10, 1026–1032 (2003).
Yan, K. S. et al. Structure and conserved RNA binding of the PAZ domain. Nature 426, 468–474 (2003).
Macrae, I. J. et al. Structural basis for double-stranded RNA processing by Dicer. Science 311, 195–198 (2006).
Calin, G. A. et al. Frequent deletions and down-regulation of micro- RNA genes miR15 and miR16 at 13q14 in chronic lymphocytic leukemia. Proc. Natl Acad. Sci. USA 99, 15524–15529 (2002).
Tam, W. Identification and characterization of human BIC, a gene on chromosome 21 that encodes a noncoding RNA. Gene 274, 157–167 (2001).
Acknowledgements
We are grateful to H. Siomi, T. Watanabe, Y. Watanabe and S. Kuramochi-Miyagawa for helpful discussions and comments. This work was supported by the Creative Research Initiatives Program (V.N.K.), the BK21 Research Funds from the Ministry of Education (J.H.) from Science and Technology of Korea, New Energy and Industrial Technology Development Organization (NEDO) grants (M.C.S.) and Core Research for Evolutionary Science and Technology (CREST) from the Japan Science and Technology Agency (JST) (M.C.S.). M.C.S. is Associate Professor of Global COE for Human Metabolomics Systems Biology by the Ministry of Education, Culture, Science and Technology, Japan.
Author information
Authors and Affiliations
Corresponding authors
Related links
Related links
DATABASES
FURTHER INFORMATION
Glossary
- Heterochromatin
-
Highly condensed regions of the genome in which transcription is generally limited.
- RNase III-type protein
-
An endonuclease that cleaves double-stranded RNAs and creates 5′-phosphate and 3′-hydroxyl termini, leaving 2-nucleotide 3′ overhangs.
- Bilaterian
-
An animal that has a front, a back, an upside and a downside (bilateral symmetry).
- Paralogue
-
A gene or protein with a highly similar sequence to another that is encoded in the same genome.
- Polycistronic transcription unit
-
An RNA transcript that includes regions that represent multiple gene products.
- Stem-loop structure
-
A lollipop-shaped structure that is formed when a single-stranded nucleic acid molecule loops back on itself to form a complementary double helix (stem) topped by a loop.
- Mirtron
-
A microRNA that is generated from a short spliced intron without Drosha-mediated cleavage.
- dsRBD
-
(Double-stranded-RNA-binding domain). A protein domain that binds to the A-form double-stranded RNA helix. Proteins that contain a dsRBD function in RNA localization, editing, translational repression and post-transcriptional gene silencing.
- Retrotransposon
-
A transposon that mobilizes through RNA intermediates; the DNA elements are transcribed into RNA and then reverse-transcribed into DNA, which is inserted at new sites in the genome.
- RNA-dependent RNA polymerase
-
An RNA polymerase that transcribes RNAs from RNA templates.
Rights and permissions
About this article
Cite this article
Kim, V., Han, J. & Siomi, M. Biogenesis of small RNAs in animals. Nat Rev Mol Cell Biol 10, 126–139 (2009). https://doi.org/10.1038/nrm2632
Issue Date:
DOI: https://doi.org/10.1038/nrm2632
This article is cited by
-
Molecular functions of microRNAs in colorectal cancer: recent roles in proliferation, angiogenesis, apoptosis, and chemoresistance
Naunyn-Schmiedeberg's Archives of Pharmacology (2024)
-
Epigenetic and metabolic reprogramming in inflammatory bowel diseases: diagnostic and prognostic biomarkers in colorectal cancer
Cancer Cell International (2023)
-
Characterization of MicroRNA expression profiles in the ovarian tissue of goats during the sexual maturity period
Journal of Ovarian Research (2023)
-
The epigenetic regulatory mechanism of PIWI/piRNAs in human cancers
Molecular Cancer (2023)
-
Employing nanoparticle tracking analysis of salivary neuronal exosomes for early detection of neurodegenerative diseases
Translational Neurodegeneration (2023)