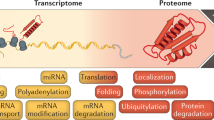
Key Points
-
Changes in mRNA-decay rates account for a large proportion of regulated gene expression.
-
The predominant pathway initiates with deadenylation. Subsequently, the mRNA can either undergo decapping and 5′→3′ decay, or 3′→5′ decay. Many of the enzymes and regulatory factors for these processes have now been described.
-
Recently, cytoplasmic structures, known as P bodies, which are sites of mRNA decay, have been identified.
-
There are minor pathways of decay for certain mRNAs. These include pathways initiated by deadenylation-independent decapping and endoribonucleolytic cleavage.
-
There are several surveillance mechanisms to degrade aberrant and unwanted mRNAs. These include nonsense-mediated decay, non-stop decay and no-go decay.
-
Turnover of mRNAs is a highly regulated process that responds to cellular signals. Sequences within the mRNA, such as AU-rich elements, can determine the rate of decay under various conditions.
-
RNA-binding proteins recognize sequence elements and modulate the rate of decay, some by recruiting the decay machinery, others by remodelling the conformation of messenger ribonucleoprotein.
-
The process of mRNA decay is just one aspect of mRNA metabolism, but it is intimately linked with other processes, including translation, transcription and mRNA localization.
Abstract
When considering the control of gene expression, the focus has traditionally been on transcriptional regulation. Recently, however, the large contribution made by mRNA decay has become difficult to ignore. Large-scale analyses indicate that as many as half of all changes in the amounts of mRNA in some responses can be attributed to altered rates of decay. In this article, we discuss some of the mechanisms that are used by the cell to mediate and regulate this intriguing process.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Amrani, N., Sachs, M. S. & Jacobson, A. Early nonsense: mRNA decay solves a translational problem. Nature Rev. Mol. Cell Biol. 7, 415–425 (2006).
Conti, E. & Izaurralde, E. Nonsense-mediated mRNA decay: molecular insights and mechanistic variations across species. Curr. Opin. Cell Biol. 17, 316–325 (2005).
Anderson, P. & Kedersha, N. RNA granules. J. Cell Biol. 172, 803–808 (2006).
Houseley, J., LaCava, J. & Tollervey, D. RNA-quality control by the exosome. Nature Rev. Mol. Cell Biol. 7, 529–539 (2006).
Eulalio, A., Behm-Ansmant, I. & Izaurralde, E. P bodies: at the crossroads of post-transcriptional pathways. Nature Rev. Mol. Cell Biol. 8, 9–22 (2006).
Khabar, K. S. The AU-rich transcriptome: more than interferons and cytokines, and its role in disease. J. Interferon Cytokine Res. 25, 1–10 (2005).
He, F. et al. Genome-wide analysis of mRNAs regulated by the nonsense-mediated and 5′ to 3′ mRNA decay pathways in yeast. Mol. Cell 12, 1439–1452 (2003).
Houalla, R. et al. Microarray detection of novel nuclear RNA substrates for the exosome. Yeast 23, 439–454 (2006).
Brown, C. E., Tarun, S. Z. Jr, Boeck, R. & Sachs, A. B. PAN3 encodes a subunit of the Pab1p-dependent poly(A) nuclease in Saccharomyces cerevisiae. Mol. Cell Biol. 16, 5744–5753 (1996).
Yamashita, A. et al. Concerted action of poly(A) nucleases and decapping enzyme in mammalian mRNA turnover. Nature Struct. Mol. Biol. 12, 1054–1063 (2005). The authors used a β-globin reporter to show that there are two phases of poly(A) shortening in mammalian cells, the first performed by PAN2–PAN3 and the second by CCR4–NOT.
Tucker, M. et al. The transcription factor associated Ccr4 and Caf1 proteins are components of the major cytoplasmic mRNA deadenylase in Saccharomyces cerevisiae. Cell 104, 377–386 (2001).
Tucker, M., Staples, R. R., Valencia-Sanchez, M. A., Muhlrad, D. & Parker, R. Ccr4p is the catalytic subunit of a Ccr4p/Pop2p/Notp mRNA deadenylase complex in Saccharomyces cerevisiae. EMBO J. 21, 1427–1436 (2002).
Dupressoir, A. et al. Identification of four families of yCCR4- and Mg2+-dependent endonuclease-related proteins in higher eukaryotes, and characterization of orthologs of yCCR4 with a conserved leucine-rich repeat essential for hCAF1/hPOP2 binding. BMC Genomics 2, 9 (2001).
Dehlin, E., Wormington, M., Korner, C. G. & Wahle, E. Cap-dependent deadenylation of mRNA. EMBO J. 19, 1079–1086 (2000).
Gao, M., Fritz, D. T., Ford, L. P. & Wilusz, J. Interaction between a poly(A)-specific ribonuclease and the 5′ cap influences mRNA deadenylation rates in vitro. Mol. Cell 5, 479–488 (2000).
Martinez, J., Ren, Y. G., Nilsson, P., Ehrenberg, M. & Virtanen, A. The mRNA cap structure stimulates rate of poly(A) removal and amplifies processivity of degradation. J. Biol. Chem. 276, 27923–27929 (2001).
Balatsos, N. A., Nilsson, P., Mazza, C., Cusack, S. & Virtanen, A. Inhibition of mRNA deadenylation by the nuclear cap binding complex (CBC). J. Biol. Chem. 281, 4517–4522 (2006).
Korner, C. G. et al. The deadenylating nuclease (DAN) is involved in poly(A) tail removal during the meiotic maturation of Xenopus oocytes. EMBO J. 17, 5427–5437 (1998).
Milone, J., Wilusz, J. & Bellofatto, V. Characterization of deadenylation in trypanosome extracts and its inhibition by poly(A)-binding protein Pab1p. RNA 10, 448–457 (2004).
Opyrchal, M., Anderson, J. R., Sokoloski, K. J., Wilusz, C. J. & Wilusz, J. A cell-free mRNA stability assay reveals conservation of the enzymes and mechanisms of mRNA decay between mosquito and mammalian cell lines. Insect Biochem. Mol. Biol. 35, 1321–1334 (2005).
Reverdatto, S. V., Dutko, J. A., Chekanova, J. A., Hamilton, D. A. & Belostotsky, D. A. mRNA deadenylation by PARN is essential for embryogenesis in higher plants. RNA 10, 1200–1214 (2004).
Chiba, Y. et al. AtPARN is an essential poly(A) ribonuclease in Arabidopsis. Gene 328, 95–102 (2004).
Temme, C., Zaessinger, S., Meyer, S., Simonelig, M. & Wahle, E. A complex containing the CCR4 and CAF1 proteins is involved in mRNA deadenylation in Drosophila. EMBO J. 23, 2862–2871 (2004).
Moser, M. J., Holley, W. R., Chatterjee, A. & Mian, I. S. The proofreading domain of Escherichia coli DNA polymerase I and other DNA and/or RNA exonuclease domains. Nucleic Acids Res. 25, 5110–5118 (1997).
Liu, H., Rodgers, N. D., Jiao, X. & Kiledjian, M. The scavenger mRNA decapping enzyme DcpS is a member of the HIT family of pyrophosphatases. EMBO J. 21, 4699–4708 (2002).
Steiger, M., Carr-Schmid, A., Schwartz, D. C., Kiledjian, M. & Parker, R. Analysis of recombinant yeast decapping enzyme. RNA 9, 231–238 (2003).
Fenger-Gron, M., Fillman, C., Norrild, B. & Lykke-Andersen, J. Multiple processing body factors and the ARE-binding protein TTP activate mRNA decapping. Mol. Cell 20, 905–915 (2005).
Yu, J. H., Yang, W. H., Gulick, T., Bloch, K. D. & Bloch, D. B. Ge-1 is a central component of the mammalian cytoplasmic mRNA processing body. RNA 11, 1795–1802 (2005).
Tharun, S. et al. Yeast Sm-like proteins function in mRNA decapping and decay. Nature 404, 515–518 (2000).
Tharun, S. & Parker, R. Targeting an mRNA for decapping: displacement of translation factors and association of the Lsm1p–7p complex on deadenylated yeast mRNAs. Mol. Cell 8, 1075–1083 (2001).
Kshirsagar, M. & Parker, R. Identification of Edc3p as an enhancer of mRNA decapping in Saccharomyces cerevisiae. Genetics 166, 729–739 (2004).
Mangus, D. A., Amrani, N. & Jacobson, A. Pbp1p, a factor interacting with Saccharomyces cerevisiae poly(A)-binding protein, regulates polyadenylation. Mol. Cell Biol. 18, 7383–7396 (1998).
Yang, W. H., Yu, J. H., Gulick, T., Bloch, K. D. & Bloch, D. B. RNA-associated protein 55 (RAP55) localizes to mRNA processing bodies and stress granules. RNA 12, 547–554 (2006).
Wilusz, C. J. & Wilusz, J. Eukaryotic Lsm proteins: lessons from bacteria. Nature Struct. Mol. Biol. 12, 1031–1036 (2005).
Schwartz, D., Decker, C. J. & Parker, R. The enhancer of decapping proteins, Edc1p and Edc2p, bind RNA and stimulate the activity of the decapping enzyme. RNA 9, 239–251 (2003).
Coller, J. M., Tucker, M., Sheth, U., Valencia-Sanchez, M. A. & Parker, R. The DEAD box helicase, Dhh1p, functions in mRNA decapping and interacts with both the decapping and deadenylase complexes. RNA 7, 1717–1727 (2001).
Bonnerot, C., Boeck, R. & Lapeyre, B. The two proteins Pat1p (Mrt1p) and Spb8p interact in vivo, are required for mRNA decay, and are functionally linked to Pab1p. Mol. Cell Biol. 20, 5939–5946 (2000).
Coller, J. & Parker, R. General translational repression by activators of mRNA decapping. Cell 122, 875–886 (2005). Highlights the correlation between translation and mRNA decay. The authors show that yeast Dhh1 and Pat1 are important for translational repression that occurs prior to transport to P bodies and decapping.
Ferraiuolo, M. A. et al. A role for the eIF4E-binding protein 4E-T in P-body formation and mRNA decay. J. Cell Biol. 170, 913–924 (2005).
Andrei, M. A. et al. A role for eIF4E and eIF4E-transporter in targeting mRNPs to mammalian processing bodies. RNA 11, 717–727 (2005).
Cougot, N., Babajko, S. & Seraphin, B. Cytoplasmic foci are sites of mRNA decay in human cells. J. Cell Biol. 165, 31–40 (2004).
Sheth, U. & Parker, R. Decapping and decay of messenger RNA occur in cytoplasmic processing bodies. Science 300, 805–808 (2003).
Bashkirov, V. I., Scherthan, H., Solinger, J. A., Buerstedde, J. M. & Heyer, W. D. A mouse cytoplasmic exoribonuclease (mXRN1p) with preference for G4 tetraplex substrates. J. Cell Biol. 136, 761–773 (1997).
Unterholzner, L. & Izaurralde, E. SMG7 acts as a molecular link between mRNA surveillance and mRNA decay. Mol. Cell 16, 587–596 (2004).
Sheth, U. & Parker, R. Targeting of aberrant mRNAs to cytoplasmic processing bodies. Cell 125, 1095–1109 (2006). Shows that NMD substrates are localized to P bodies in S. cerevisiae . Evidence indicates that P-body localization is not sufficient for mRNA decay and that another step is required to trigger degradation.
Sen, G. L. & Blau, H. M. Argonaute 2/RISC resides in sites of mammalian mRNA decay known as cytoplasmic bodies. Nature Cell Biol. 7, 633–636 (2005).
Graham, A. C., Kiss, D. L. & Andrulis, E. D. Differential distribution of exosome subunits at the nuclear lamina and in cytoplasmic foci. Mol. Biol. Cell 17, 1399–1409 (2006).
Kedersha, N. et al. Stress granules and processing bodies are dynamically linked sites of mRNP remodeling. J. Cell Biol. 169, 871–884 (2005).
Teixeira, D., Sheth, U., Valencia-Sanchez, M. A., Brengues, M. & Parker, R. Processing bodies require RNA for assembly and contain nontranslating mRNAs. RNA. 11, 371–382 (2005).
Segal, S. P., Dunckley, T. & Parker, R. Sbp1p affects translational repression and decapping in Saccharomyces cerevisiae. Mol. Cell Biol. 26, 5120–5130 (2006).
Hilgers, V., Teixeira, D. & Parker, R. Translation-independent inhibition of mRNA deadenylation during stress in Saccharomyces cerevisiae. RNA 12, 1835–1845 (2006).
Bhattacharyya, S. N., Habermacher, R., Martine, U., Closs, E. I. & Filipowicz, W. Relief of microRNA-mediated translational repression in human cells subjected to stress. Cell 125, 1111–1124 (2006). Shows a link between miRNA-mediated decay and ARE-dependent mRNA stabilization mediated by HuR. Furthermore, it shows that mRNAs can be released from P bodies in mammalian cells under the appropriate conditions.
Brengues, M., Teixeira, D. & Parker, R. Movement of eukaryotic mRNAs between polysomes and cytoplasmic processing bodies. Science 310, 486–489 (2005). Shows that mRNAs can be released from P bodies in S. cerevisiae and return to polysomes for translation.
Badis, G., Saveanu, C., Fromont-Racine, M. & Jacquier, A. Targeted mRNA degradation by deadenylation-independent decapping. Mol. Cell 15, 5–15 (2004).
Muhlrad, D. & Parker, R. The yeast EDC1 mRNA undergoes deadenylation-independent decapping stimulated by Not2p, Not4p, and Not5p. EMBO J. 24, 1033–1045 (2005).
Gatfield, D. & Izaurralde, E. Nonsense-mediated messenger RNA decay is initiated by endonucleolytic cleavage in Drosophila. Nature 429, 575–578 (2004). Describes a novel route for NMD that is independent of the EJC in D. melanogaster.
Stevens, A. et al. β-Globin mRNA decay in erythroid cells: UG site-preferred endonucleolytic cleavage that is augmented by a premature termination codon. Proc. Natl Acad. Sci. USA 99, 12741–12746 (2002).
Liu, J. et al. Argonaute2 is the catalytic engine of mammalian RNAi. Science 305, 1437–1441 (2004).
Song, J. J., Smith, S. K., Hannon, G. J. & Joshua-Tor, L. Crystal structure of Argonaute and its implications for RISC slicer activity. Science 305, 1434–1437 (2004).
Chernokalskaya, E. et al. A polysomal ribonuclease involved in the destabilization of albumin mRNA is a novel member of the peroxidase gene family. RNA 4, 1537–1548 (1998).
Yang, F. & Schoenberg, D. R. Endonuclease-mediated mRNA decay involves the selective targeting of PMR1 to polyribosome-bound substrate mRNA. Mol. Cell 14, 435–445 (2004).
Yang, F. et al. Polysome-bound endonuclease PMR1 is targeted to stress granules via stress-specific binding to TIA-1. Mol. Cell Biol. 26, 8803–8813 (2006).
Hollien, J. & Weissman, J. S. Decay of endoplasmic reticulum-localized mRNAs during the unfolded protein response. Science 313, 104–107 (2006). Shows that ER-localized mRNAs are degraded by an endonuclease, probably IRE1, during the unfolded-protein response.
Yoshida, H., Matsui, T., Yamamoto, A., Okada, T. & Mori, K. XBP1 mRNA is induced by ATF6 and spliced by IRE1 in response to ER stress to produce a highly active transcription factor. Cell 107, 881–891 (2001).
Xiao, S., Scott, F., Fierke, C. A. & Engelke, D. R. Eukaryotic ribonuclease P: a plurality of ribonucleoprotein enzymes. Annu. Rev. Biochem. 71, 165–189 (2002).
Gill, T., Cai, T., Aulds, J., Wierzbicki, S. & Schmitt, M. E. RNase MRP cleaves the CLB2 mRNA to promote cell cycle progression: novel method of mRNA degradation. Mol. Cell Biol. 24, 945–953 (2004).
Gill, T., Aulds, J. & Schmitt, M. E. A specialized processing body that is temporally and asymmetrically regulated during the cell cycle in Saccharomyces cerevisiae. J. Cell Biol. 173, 35–45 (2006). Shows that a specialized P body that contains RNAse MRP is involved in degradation of the cyclin-B mRNA in S. cerevisiae.
Thiel, C. T. et al. Severely incapacitating mutations in patients with extreme short stature identify RNA-processing endoribonuclease RMRP as an essential cell growth regulator. Am. J. Hum. Genet. 77, 795–806 (2005).
Yang, F., Peng, Y. & Schoenberg, D. R. Endonuclease-mediated mRNA decay requires tyrosine phosphorylation of polysomal ribonuclease 1 (PMR1) for the targeting and degradation of polyribosome-bound substrate mRNA. J. Biol. Chem. 279, 48993–49002 (2004).
Le, H. H., Izaurralde, E., Maquat, L. E. & Moore, M. J. The spliceosome deposits multiple proteins 20–24 nucleotides upstream of mRNA exon–exon junctions. EMBO J. 19, 6860–6869 (2000).
Gatfield, D., Unterholzner, L., Ciccarelli, F. D., Bork, P. & Izaurralde, E. Nonsense-mediated mRNA decay in Drosophila : at the intersection of the yeast and mammalian pathways. EMBO J. 22, 3960–3970 (2003).
Buhler, M., Steiner, S., Mohn, F., Paillusson, A. & Muhlemann, O. EJC-independent degradation of nonsense immunoglobulin-μ mRNA depends on 3′ UTR length. Nature Struct. Mol. Biol. 13, 462–464 (2006). Provides the best evidence for EJC-independent NMD in mammalian cells.
Amrani, N. et al. A faux 3′-UTR promotes aberrant termination and triggers nonsense-mediated mRNA decay. Nature 432, 112–118 (2004).
Kashima, I. et al. Binding of a novel SMG-1–Upf1–eRF1–eRF3 complex (SURF) to the exon junction complex triggers Upf1 phosphorylation and nonsense-mediated mRNA decay. Genes Dev. 20, 355–367 (2006).
Chiu, S. Y., Serin, G., Ohara, O. & Maquat, L. E. Characterization of human Smg5/7a: a protein with similarities to Caenorhabditis elegans SMG5 and SMG7 that functions in the dephosphorylation of Upf1. RNA 9, 77–87 (2003).
Ohnishi, T. et al. Phosphorylation of hUPF1 induces formation of mRNA surveillance complexes containing hSMG-5 and hSMG-7. Mol. Cell 12, 1187–1200 (2003).
Mitchell, P. & Tollervey, D. An NMD pathway in yeast involving accelerated deadenylation and exosome-mediated 3′→5′ degradation. Mol. Cell 11, 1405–1413 (2003).
Lejeune, F., Li, X. & Maquat, L. E. Nonsense-mediated mRNA decay in mammalian cells involves decapping, deadenylating, and exonucleolytic activities. Mol. Cell 12, 675–687 (2003).
Mendell, J. T., Sharifi, N. A., Meyers, J. L., Martinez-Murillo, F. & Dietz, H. C. Nonsense surveillance regulates expression of diverse classes of mammalian transcripts and mutes genomic noise. Nature Genet. 36, 1073–1078 (2004).
Kaygun, H. & Marzluff, W. F. Regulated degradation of replication-dependent histone mRNAs requires both ATR and Upf1. Nature Struct. Mol. Biol. 12, 794–800 (2005). Shows that Upf1 can induce histone-mRNA decay during the cell cycle independent of other NMD factors.
Kim, Y. K., Furic, L., DesGroseillers, L. & Maquat, L. E. Mammalian Staufen1 recruits Upf1 to specific mRNA 3′UTRs so as to elicit mRNA decay. Cell 120, 195–208 (2005). Describes another non-NMD role for the UPF1 protein.
van Hoof, A., Frischmeyer, P. A., Dietz, H. C. & Parker, R. Exosome-mediated recognition and degradation of mRNAs lacking a termination codon. Science 295, 2262–2264 (2002).
Frischmeyer, P. A. et al. An mRNA surveillance mechanism that eliminates transcripts lacking termination codons. Science 295, 2258–2261 (2002).
Inada, T. & Aiba, H. Translation of aberrant mRNAs lacking a termination codon or with a shortened 3′-UTR is repressed after initiation in yeast. EMBO J. 24, 1584–1595 (2005).
Doma, M. K. & Parker, R. Endonucleolytic cleavage of eukaryotic mRNAs with stalls in translation elongation. Nature 440, 561–564 (2006). The first description of no-go decay, which initiates with an endonucleolytic-cleavage event.
Onouchi, H. et al. Nascent peptide-mediated translation elongation arrest coupled with mRNA degradation in the CGS1 gene of Arabidopsis. Genes Dev. 19, 1799–1810 (2005).
Fan, J. et al. Global analysis of stress-regulated mRNA turnover by using cDNA arrays. Proc. Natl Acad. Sci. USA 99, 10611–10616 (2002).
Cheadle, C. et al. Control of gene expression during T cell activation: alternate regulation of mRNA transcription and mRNA stability. BMC Genomics 6, 75 (2005).
Stoecklin, G., Lu, M., Rattenbacher, B. & Moroni, C. A constitutive decay element promotes tumor necrosis factor α mRNA degradation via an AU-rich element-independent pathway. Mol. Cell Biol. 23, 3506–3515 (2003).
Moraes, K. C., Wilusz, C. J. & Wilusz, J. CUG-BP binds to RNA substrates and recruits PARN deadenylase. RNA 12, 1084–1091 (2006).
Ueno, S. & Sagata, N. Requirement for both EDEN and AUUUA motifs in translational arrest of Mos mRNA upon fertilization of Xenopus eggs. Dev. Biol. 250, 156–167 (2002).
Anderson, J. R. et al. Sequence-specific RNA binding mediated by the RNase PH domain of components of the exosome. RNA 12, 1810–1816 (2006).
Chen, C. Y. et al. AU binding proteins recruit the exosome to degrade ARE-containing mRNAs. Cell 107, 451–464 (2001).
Gherzi, R. et al. A KH domain RNA binding protein, KSRP, promotes ARE-directed mRNA turnover by recruiting the degradation machinery. Mol. Cell 14, 571–583 (2004).
Tran, H., Schilling, M., Wirbelauer, C., Hess, D. & Nagamine, Y. Facilitation of mRNA deadenylation and decay by the exosome-bound, DExH protein RHAU. Mol. Cell 13, 101–111 (2004).
Chou, C. F. et al. Tethering KSRP, a decay-promoting AU-rich element-binding protein, to mRNAs elicits mRNA decay. Mol. Cell Biol. 26, 3695–3706 (2006).
Lykke-Andersen, J. & Wagner, E. Recruitment and activation of mRNA decay enzymes by two ARE-mediated decay activation domains in the proteins TTP and BRF-1. Genes Dev. 19, 351–361 (2005).
Lai, W. S., Kennington, E. A. & Blackshear, P. J. Tristetraprolin and its family members can promote the cell-free deadenylation of AU-rich element-containing mRNAs by poly(A) ribonuclease. Mol. Cell Biol. 23, 3798–3812 (2003).
Lal, A. et al. Concurrent versus individual binding of HuR and AUF1 to common labile target mRNAs. EMBO J. 23, 3092–3102 (2004).
Linker, K. et al. Involvement of KSRP in the post-transcriptional regulation of human iNOS expression-complex interplay of KSRP with TTP and HuR. Nucleic Acids Res. 33, 4813–4827 (2005).
Briata, P. et al. p38-dependent phosphorylation of the mRNA decay-promoting factor KSRP controls the stability of select myogenic transcripts. Mol. Cell 20, 891–903 (2005). Excellent example of how mRNP structure can be regulated through modification of RNA-binding proteins, leading to changes in the rate of mRNA decay.
Figueroa, A. et al. Role of HuR in skeletal myogenesis through coordinate regulation of muscle differentiation genes. Mol. Cell Biol. 23, 4991–5004 (2003).
Rigby, W. F. et al. Structure/function analysis of tristetraprolin (TTP): p38 stress-activated protein kinase and lipopolysaccharide stimulation do not alter TTP function. J. Immunol. 174, 7883–7893 (2005).
Johnson, B. A., Stehn, J. R., Yaffe, M. B. & Blackwell, T. K. Cytoplasmic localization of tristetraprolin involves 14-3-3-dependent and -independent mechanisms. J. Biol. Chem. 277, 18029–18036 (2002).
Stoecklin, G. et al. MK2-induced tristetraprolin:14-3-3 complexes prevent stress granule association and ARE-mRNA decay. EMBO J. 23, 1313–1324 (2004).
Shen, Z. J., Esnault, S. & Malter, J. S. The peptidyl-prolyl isomerase Pin1 regulates the stability of granulocyte-macrophage colony-stimulating factor mRNA in activated eosinophils. Nature Immunol. 6, 1280–1287 (2005). Shows that regulation of an RNA-binding protein through isomerization leads to changes in mRNA decay.
Goldstrohm, A. C., Hook, B. A., Seay, D. J. & Wickens, M. PUF proteins bind Pop2p to regulate messenger RNAs. Nature Struct. Mol. Biol. 13, 533–539 (2006).
Gerber, A. P., Herschlag, D. & Brown, P. O. Extensive association of functionally and cytotopically related mRNAs with Puf family RNA-binding proteins in yeast. PLoS. Biol. 2, e79 (2004).
Foat, B. C., Houshmandi, S. S., Olivas, W. M. & Bussemaker, H. J. Profiling condition-specific, genome-wide regulation of mRNA stability in yeast. Proc. Natl Acad. Sci. USA 102, 17675–17680 (2005).
Kiledjian, M., Wang, X. & Liebhaber, S. A. Identification of two KH domain proteins in the α-globin mRNP stability complex. EMBO J. 14, 4357–4364 (1995).
Yu, J. & Russell, J. E. Structural and functional analysis of an mRNP complex that mediates the high stability of human β-globin mRNA. Mol. Cell Biol. 21, 5879–5888 (2001).
Lindquist, J. N., Parsons, C. J., Stefanovic, B. & Brenner, D. A. Regulation of α1(I) collagen messenger RNA decay by interactions with αCP at the 3′-untranslated region. J. Biol. Chem. 279, 23822–23829 (2004).
Ostareck, D. H. et al. mRNA silencing in erythroid differentiation: hnRNP K and hnRNP E1 regulate 15-lipoxygenase translation from the 3′ end. Cell 89, 597–606 (1997).
Kong, J., Sumaroka, M., Eastmond, D. L. & Liebhaber, S. A. Shared stabilization functions of pyrimidine-rich determinants in the erythroid 15-lipoxygenase and α-globin mRNAs. Mol. Cell Biol. 26, 5603–5614 (2006).
Wang, Z., Day, N., Trifillis, P. & Kiledjian, M. An mRNA stability complex functions with poly(A)-binding protein to stabilize mRNA in vitro. Mol. Cell Biol. 19, 4552–4560 (1999).
Kim, J. H. & Richter, J. D. Opposing polymerase–deadenylase activities regulate cytoplasmic polyadenylation. Mol. Cell 24, 173–183 (2006). Intriguing finding that a deadenylase and a polyadenylase are part of the same complex that is involved in translational regulation.
Wilczynska, A., Aigueperse, C., Kress, M., Dautry, F. & Weil, D. The translational regulator CPEB1 provides a link between dcp1 bodies and stress granules. J. Cell Sci. 118, 981–992 (2005).
Panasenko, O. et al. The yeast CCR4–Not complex controls ubiquitination of the nascent associated polypeptide complex. J. Biol. Chem. 281, 31389–31398 (2006).
Lotan, R. et al. The RNA polymerase II subunit Rpb4p mediates decay of a specific class of mRNAs. Genes Dev. 19, 3004–3016 (2005).
Lin, M. D., Fan, S. J., Hsu, W. S. & Chou, T. B. Drosophila decapping protein 1, dDcp1, is a component of the oskar mRNP complex and directs its posterior localization in the oocyte. Dev. Cell 10, 601–613 (2006).
Nakamura, A., Amikura, R., Hanyu, K. & Kobayashi, S. Me31B silences translation of oocyte-localizing RNAs through the formation of cytoplasmic RNP complex during Drosophila oogenesis. Development 128, 3233–3242 (2001).
Semotok, J. L. et al. Smaug recruits the CCR4/POP2/NOT deadenylase complex to trigger maternal transcript localization in the early Drosophila embryo. Curr. Biol. 15, 284–294 (2005).
Valencia-Sanchez, M. A., Liu, J., Hannon, G. J. & Parker, R. Control of translation and mRNA degradation by miRNAs and siRNAs. Genes Dev. 20, 515–524 (2006).
Kim, V. N. Small RNAs just got bigger: Piwi-interacting RNAs (piRNAs) in mammalian testes. Genes Dev. 20, 1993–1997 (2006).
Zamore, P. D. & Haley, B. Ribo-gnome: the big world of small RNAs. Science 309, 1519–1524 (2005).
Behm-Ansmant, I. et al. mRNA degradation by miRNAs and GW182 requires both CCR4:NOT deadenylase and DCP1:DCP2 decapping complexes. Genes Dev. 20, 1885–1898 (2006).
Wu, L., Fan, J. & Belasco, J. G. MicroRNAs direct rapid deadenylation of mRNA. Proc. Natl Acad. Sci. USA 103, 4034–4039 (2006).
Giraldez, A. J. et al. Zebrafish MiR-430 promotes deadenylation and clearance of maternal mRNAs. Science 312, 75–79 (2006). References 127 and 128 link miRNA function to the normal pathway of mRNA decay.
Jing, Q. et al. Involvement of microRNA in AU-rich element-mediated mRNA instability. Cell 120, 623–634 (2005).
Das, B., Butler, J. S. & Sherman, F. Degradation of normal mRNA in the nucleus of Saccharomyces cerevisiae. Mol. Cell Biol. 23, 5502–5515 (2003).
Milligan, L., Torchet, C., Allmang, C., Shipman, T. & Tollervey, D. A nuclear surveillance pathway for mRNAs with defective polyadenylation. Mol. Cell Biol. 25, 9996–10004 (2005).
Bousquet-Antonelli, C., Presutti, C. & Tollervey, D. Identification of a regulated pathway for nuclear pre-mRNA turnover. Cell 102, 765–775 (2000).
Kufel, J., Bousquet-Antonelli, C., Beggs, J. D. & Tollervey, D. Nuclear pre-mRNA decapping and 5′ degradation in yeast require the Lsm2–8p complex. Mol. Cell Biol. 24, 9646–9657 (2004).
Kuai, L., Das, B. & Sherman, F. A nuclear degradation pathway controls the abundance of normal mRNAs in Saccharomyces cerevisiae. Proc. Natl Acad. Sci. USA 102, 13962–13967 (2005).
Roth, K. M., Wolf, M. K., Rossi, M. & Butler, J. S. The nuclear exosome contributes to autogenous control of NAB2 mRNA levels. Mol. Cell Biol. 25, 1577–1585 (2005).
LaCava, J. et al. RNA degradation by the exosome is promoted by a nuclear polyadenylation complex. Cell 121, 713–724 (2005).
Vanacova, S. et al. A new yeast poly(A) polymerase complex involved in RNA quality control. PLoS Biol. 3, e189 (2005).
Liu, Q., Greimann, J. C. & Lima, C. D. Reconstitution, activities, and structure of the eukaryotic RNA exosome. Cell 127, 1223–1237 (2006).
Dziembowski, A., Lorentzen, E., Conti, E. & Seraphin, B. A single subunit, Dis3, is essentially responsible for yeast exosome core activity. Nature Struct. Mol. Biol. 17 Dec 2006 (doi: 10.1038/nsmb1184).
Acknowledgements
Research in the Wilusz laboratory is supported by grants from the National Institutes of Health. We thank N. Kedersha for supplying Figure 2 and for her insights on P bodies. We are also grateful to members of the Wilusz laboratory and S. Vasudevan for reading and commenting on the manuscript. We apologize to those authors whose work we were unable to cite due to lack of space.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
Related links
DATABASES
Saccharomyces Genome database
FURTHER INFORMATION
Glossary
- mRNA-surveillance pathway
-
Mechanism used by the cell to identify and destroy aberrant mRNAs.
- Poly(A) tail
-
A homopolymeric stretch of ∼25–200 adenine nucleotides that is present at the 3′ end of most eukaryotic mRNAs.
- Exonuclease
-
Also known as an exoribonuclease. An enzyme that catalyses the removal of mononucleotides from either the 5′ or 3′ end of an RNA molecule.
- Polysome
-
Two or more ribosomes that are bound to different sites on the same mRNA.
- 5′ cap
-
The 7-methylguanosine structure found at the 5′ end of a eukaryotic mRNA.
- Decapping
-
The removal of the mRNA 5′-cap structure.
- Exosome
-
A large complex of 3′→5′ exonucleases that functions in the nucleus and the cytoplasm in several different RNA-processing and RNA-degradation pathways.
- Transcriptome
-
The pool of mRNAs that is present in the cell under a given condition.
- Deadenylase
-
An enzyme that removes the 3′-poly(A) tail from RNA in a 3′→5′ direction.
- RNase D
-
Escherichia coli RNase D is a 3′→5′ exoribonuclease that is required for processing of tRNA precursors. Many eukaryotic proteins with similar exoribonuclease domains have been identified.
- RNA helicase
-
An enzyme that catalyses the unwinding of the RNA secondary structure.
- RNase PH
-
E. coli RNase PH is a phosphorolytic 3′→5′ exoribonuclease that is involved in the maturation of tRNA precursors.
- Sm-like domain
-
A protein domain that is similar to that found in the Sm proteins associated with the small nuclear RNAs of the splicing machinery.
- Endonuclease
-
Also known as an endoribonuclease. An enzyme that catalyses the hydrolysis of ester linkages within an RNA molecule by creating internal breaks.
- Stress granule
-
A cytoplasmic structure that develops in mammalian cells during certain types of cellular stress. It is believed to be a site where translationally repressed mRNAs are stored or sorted for decay. P bodies often assemble close to stress granules.
- Cartilage hair hypoplasia
-
A recessively inherited developmental disorder characterized by short stature, fine and sparse hair, immune deficiency and a predisposition to malignancy. The defect is caused by mutations in RNase MRP.
- Messenger ribonucleoprotein
-
(mRNP). A complex that comprises an mRNA and associated RNA-binding proteins.
- 14-3-3 proteins
-
A large family of small, acidic proteins that interact predominantly with proteins that are phosphorylated on serine or threonine residues. 14-3-3 proteins have been implicated in many intracellular-signalling events through their ability to regulate catalytic activity, subcellular localization and protein–protein interactions.
- A site
-
The site at which the ribosome binds an aminoacylated tRNA for addition to the elongating peptide chain.
- Heterogeneous nuclear ribonucleoprotein
-
(hnRNP). A historic term used to describe protein complexes that associate with mRNAs and pre-mRNAs in the nucleus.
- KH domain
-
An RNA-binding domain that was first identified in the hnRNP K protein.
Rights and permissions
About this article
Cite this article
Garneau, N., Wilusz, J. & Wilusz, C. The highways and byways of mRNA decay. Nat Rev Mol Cell Biol 8, 113–126 (2007). https://doi.org/10.1038/nrm2104
Issue Date:
DOI: https://doi.org/10.1038/nrm2104
This article is cited by
-
The multifaceted role of Fragile X-Related Protein 1 (FXR1) in cellular processes: an updated review on cancer and clinical applications
Cell Death & Disease (2024)
-
Short poly(A) tails are protected from deadenylation by the LARP1–PABP complex
Nature Structural & Molecular Biology (2023)
-
Biomolecular condensates in kidney physiology and disease
Nature Reviews Nephrology (2023)
-
Genome-wide probing of eukaryotic nascent RNA structure elucidates cotranscriptional folding and its antimutagenic effect
Nature Communications (2023)
-
Avidity-based bright and photostable light-up aptamers for single-molecule mRNA imaging
Nature Chemical Biology (2023)