Key Points
-
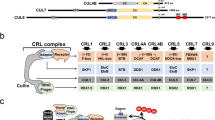
The cullin–RING ligases (CRLs) comprise a superfamily of ubiquitin ligases that are implicated in the regulation of a diverse array of eukaryotic functions.
-
The various cullin proteins function as a rigid scaffold for the assembly of this modular class of ligase. All cullins associate with a RING protein through their C-terminal domain, whereas the N-terminal region recruits a wide variety of receptor proteins that confer substrate specificity.
-
The cullin–RING module is often referred to as the catalytic core, because it is common to all CRLs. It recruits ubiquitin-conjugating enzymes (E2s) and activates the transfer of ubiquitin from the E2 to the substrate through an as-yet-unclear mechanism that does not involve a CRL–ubiquitin-thioester intermediate.
-
The substrate receptors for CRLs are generally linked to the catalytic core through adaptor proteins that are specific for each cullin-family member. Numerous substrate receptors can be recruited to each CRL core, which increases the diversity of proteins that can be targeted for ubiquitylation.
-
In most cases, the recognition of a substrate by a CRL requires post-translational modification of the substrate. This further increases the repertoire of substrates that can be targeted to a given CRL and also links protein ubiquitylation and turnover to numerous signalling pathways.
-
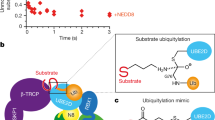
CRL activity can be regulated by numerous mechanisms, which include the turnover of substrate receptors, the reversible attachment of the ubiquitin-like protein NEDD8 to cullins, and the sequestration of cullins by CAND1.
Abstract
Cullin–RING complexes comprise the largest known class of ubiquitin ligases. Owing to the great diversity of their substrate-receptor subunits, it is possible that there are hundreds of distinct cullin–RING ubiquitin ligases in eukaryotic cells, which establishes these enzymes as key mediators of post-translational protein regulation. In this review, we focus on the composition, regulation and function of cullin–RING ligases, and describe how these enzymes can be characterized by a set of general principles.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Feldman, R. M., Correll, C. C., Kaplan, K. B. & Deshaies, R. J. A complex of Cdc4p, Skp1p, and Cdc53p/cullin catalyzes ubiquitination of the phosphorylated CDK inhibitor Sic1p. Cell 91, 221–230 (1997).
Skowyra, D., Craig, K. L., Tyers, M., Elledge, S. J. & Harper, J. W. F-box proteins are receptors that recruit phosphorylated substrates to the SCF ubiquitin-ligase complex. Cell 91, 209–219 (1997). References 1 and 2 report the discovery of the prototypical SCF ubiquitin ligase.
Zachariae, W. et al. Mass spectrometric analysis of the anaphase-promoting complex from yeast: identification of a subunit related to cullins. Science 279, 1216–1219 (1998).
Yu, H. et al. Identification of a cullin homology region in a subunit of the anaphase-promoting complex. Science 279, 1219–1222 (1998).
Nikolaev, A. Y., Li, M., Puskas, N., Qin, J. & Gu, W. Parc: a cytoplasmic anchor for p53. Cell 112, 29–40 (2003).
Zheng, N. et al. Structure of the Cul1–Rbx1–Skp1–F-boxSkp2 SCF ubiquitin ligase complex. Nature 416, 703–709 (2002). Describes the structure of an SCF ubiquitin ligase.
Tan, P. et al. Recruitment of a ROC1–CUL1 ubiquitin ligase by Skp1 and HOS to catalyze the ubiquitination of IκBα. Mol. Cell 3, 527–533 (1999).
Ohta, T., Michel, J. J., Schottelius, A. J. & Xiong, Y. ROC1, a homolog of APC11, represents a family of cullin partners with an associated ubiquitin ligase activity. Mol. Cell 3, 535–541 (1999).
Kamura, T. et al. Rbx1, a component of the VHL tumor suppressor complex and SCF ubiquitin ligase. Science 284, 657–661 (1999).
Seol, J. H. et al. Cdc53/cullin and the essential Hrt1 RING-H2 subunit of SCF define a ubiquitin ligase module that activates the E2 enzyme Cdc34. Genes Dev. 13, 1614–1626 (1999). References 7–10 discuss the discovery of the RING-H2 subunit of cullin ligases.
Schulman, B. A. et al. Insights into SCF ubiquitin ligases from the structure of the Skp1–Skp2 complex. Nature 408, 381–386 (2000).
Michel, J. J. & Xiong, Y. Human CUL-1, but not other cullin family members, selectively interacts with SKP1 to form a complex with SKP2 and cyclin A. Cell Growth Differ. 9, 435–449 (1998).
Dias, D. C., Dolios, G., Wang, R. & Pan, Z. Q. CUL7: a DOC domain-containing cullin selectively binds Skp1–Fbx29 to form an SCF-like complex. Proc. Natl Acad. Sci. USA 99, 16601–16606 (2002).
Stebbins, C. E., Kaelin, W. G. Jr. & Pavletich, N. P. Structure of the VHL–ElonginC–ElonginB complex: implications for VHL tumor suppressor function. Science 284, 455–461 (1999).
Geyer, R., Wee, S., Anderson, S., Yates, J. & Wolf, D. A. BTB/POZ domain proteins are putative substrate adaptors for cullin 3 ubiquitin ligases. Mol. Cell 12, 783–790 (2003).
Pintard, L. et al. The BTB protein MEL-26 is a substrate-specific adaptor of the CUL-3 ubiquitin-ligase. Nature 425, 311–316 (2003).
Xu, L. et al. BTB proteins are substrate-specific adaptors in an SCF-like modular ubiquitin ligase containing CUL-3. Nature 425, 316–321 (2003).
Furukawa, M., He, Y. J., Borchers, C. & Xiong, Y. Targeting of protein ubiquitination by BTB–Cullin 3–Roc1 ubiquitin ligases. Nature Cell Biol. 5, 1001–1007 (2003). References 15–18 describe the identification of BTB domains as adaptors for CUL3-containing ubiquitin ligases.
Shiyanov, P., Nag, A. & Raychaudhuri, P. Cullin 4A associates with the UV-damaged DNA-binding protein DDB. J. Biol. Chem. 274, 35309–35312 (1999).
Ulane, C. M. & Horvath, C. M. Paramyxoviruses SV5 and HPIV2 assemble STAT protein ubiquitin ligase complexes from cellular components. Virology 304, 160–166 (2002).
Wertz, I. E. et al. Human De-etiolated-1 regulates c-Jun by assembling a CUL4A ubiquitin ligase. Science 303, 1371–1374 (2004).
Bai, C. et al. SKP1 connects cell cycle regulators to the ubiquitin proteolysis machinery through a novel motif, the F-box. Cell 86, 263–274 (1996). Reports the discovery of the F-box motif.
Kamura, T. et al. The Elongin BC complex interacts with the conserved SOCS-box motif present in members of the SOCS, ras, WD-40 repeat, and ankyrin repeat families. Genes Dev. 12, 3872–3881 (1998). Shows that elongin BC recruits SOCS-box proteins to CUL2.
Kus, B. M., Caldon, C. E., Andorn-Broza, R. & Edwards, A. M. Functional interaction of 13 yeast SCF complexes with a set of yeast E2 enzymes in vitro. Proteins 54, 455–467 (2004).
Kaiser, P., Sia, R. A., Bardes, E. G., Lew, D. J. & Reed, S. I. Cdc34 and the F-box protein Met30 are required for degradation of the Cdk-inhibitory kinase Swe1. Genes Dev. 12, 2587–2597 (1998).
Skowyra, D. et al. Reconstitution of G1 cyclin ubiquitination with complexes containing SCFGrr1 and Rbx1. Science 284, 662–665 (1999).
Kaplun, L., Ivantsiv, Y., Bakhrat, A. & Raveh, D. DNA damage response-mediated degradation of Ho endonuclease via the ubiquitin system involves its nuclear export. J. Biol. Chem. 278, 48727–48734 (2003).
Kaplan, K. B., Hyman, A. A. & Sorger, P. K. Regulating the yeast kinetochore by ubiquitin-dependent degradation and Skp1p-mediated phosphorylation. Cell 91, 491–500 (1997).
Galan, J. M. et al. Skp1p and the F-box protein Rcy1p form a non-SCF complex involved in recycling of the SNARE Snc1p in yeast. Mol. Cell. Biol. 21, 3105–3117 (2001).
Seol, J. H., Shevchenko, A. & Deshaies, R. J. Skp1 forms multiple protein complexes, including RAVE, a regulator of V-ATPase assembly. Nature Cell Biol. 3, 384–391 (2001).
Pintard, L., Willems, A. & Peter, M. Cullin-based ubiquitin ligases: Cul3–BTB complexes join the family. EMBO J. 23, 1681–1687 (2004).
Russell, I. D., Grancell, A. S. & Sorger, P. K. The unstable F-box protein p58-Ctf13 forms the structural core of the CBF3 kinetochore complex. J. Cell Biol. 145, 933–950 (1999).
Suzuki, H. et al. Homodimer of two F-box proteins βTrCP1 or βTrCP2 binds to IκBα for signal-dependent ubiquitination. J. Biol. Chem. 275, 2877–2884 (2000).
Seibert, V. et al. Combinatorial diversity of fission yeast SCF ubiquitin ligases by homo- and heterooligomeric assemblies of the F-box proteins Pop1p and Pop2p. BMC Biochem. 3, 22 (2002).
Dixon, C. et al. Overproduction of polypeptides corresponding to the amino terminus of the F-box proteins Cdc4p and Met30p inhibits ubiquitin ligase activities of their SCF complexes. Eukaryot. Cell 2, 123–133 (2003).
Wu, G. et al. Structure of a β-TrCP1–Skp1–β-catenin complex: destruction motif binding and lysine specificity of the SCFβ-TrCP1 ubiquitin ligase. Mol. Cell 11, 1445–1456 (2003).
Orlicky, S., Tang, X., Willems, A., Tyers, M. & Sicheri, F. Structural basis for phosphodependent substrate selection and orientation by the SCFCdc4 ubiquitin ligase. Cell 112, 243–256 (2003).
Ganoth, D. et al. The cell-cycle regulatory protein Cks1 is required for SCFSkp2-mediated ubiquitinylation of p27. Nature Cell Biol. 3, 321–324 (2001).
Spruck, C. et al. A CDK-independent function of mammalian Cks1: targeting of SCFSkp2 to the CDK inhibitor p27Kip1. Mol. Cell 7, 639–650 (2001).
Zhou, P. & Howley, P. M. Ubiquitination and degradation of the substrate recognition subunits of SCF ubiquitin-protein ligases. Mol. Cell 2, 571–580 (1998).
Galan, J. M. & Peter, M. Ubiquitin-dependent degradation of multiple F-box proteins by an autocatalytic mechanism. Proc. Natl Acad. Sci. USA 96, 9124–9129 (1999).
Schoenfeld, A. R., Davidowitz, E. J. & Burk, R. D. Elongin BC complex prevents degradation of von Hippel–Lindau tumor suppressor gene products. Proc. Natl Acad. Sci. USA 97, 8507–8512 (2000).
Ayad, N. G. et al. Tome-1, a trigger of mitotic entry, is degraded during G1 via the APC. Cell 113, 101–113 (2003).
Bashir, T., Dorrello, N. V., Amador, V., Guardavaccaro, D. & Pagano, M. Control of the SCFSkp2–Cks1 ubiquitin ligase by the APC/CCdh1 ubiquitin ligase. Nature 428, 190–193 (2004).
Wei, W. et al. Degradation of the SCF component Skp2 in cell-cycle phase G1 by the anaphase-promoting complex. Nature 428, 194–198 (2004).
Margottin-Goguet, F. et al. Prophase destruction of Emi1 by the SCFβTrCP/Slimb ubiquitin ligase activates the anaphase promoting complex to allow progression beyond prometaphase. Dev. Cell 4, 813–826 (2003).
Guardavaccaro, D. et al. Control of meiotic and mitotic progression by the F box protein β-Trcp1 in vivo. Dev. Cell 4, 799–812 (2003).
Deshaies, R. J. SCF and Cullin/Ring H2-based ubiquitin ligases. Annu. Rev. Cell Dev. Biol. 15, 435–467 (1999).
Li, Y., Gazdoiu, S., Pan, Z. Q. & Fuchs, S. Y. Stability of homologue of Slimb F-box protein is regulated by availability of its substrate. J. Biol. Chem. 279, 11074–11080 (2004).
Kaiser, P., Flick, K., Wittenberg, C. & Reed, S. I. Regulation of transcription by ubiquitination without proteolysis: Cdc34/SCFMet30-mediated inactivation of the transcription factor Met4. Cell 102, 303–314 (2000).
Flick, K. et al. Proteolysis-independent regulation of the transcription factor Met4 by a single Lys48-linked ubiquitin chain. Nature Cell Biol. 6, 634–641 (2004).
Kuras, L. et al. Dual regulation of the met4 transcription factor by ubiquitin-dependent degradation and inhibition of promoter recruitment. Mol. Cell 10, 69–80 (2002).
Jiang, J. Degrading Ci: who is Cul-pable? Genes Dev. 16, 2315–2321 (2002).
Fong, A. & Sun, S. C. Genetic evidence for the essential role of β-transducin repeat-containing protein in the inducible processing of NF-κB2/p100. J. Biol. Chem. 277, 22111–22114 (2002).
von der Lehr, N. et al. The F-box protein Skp2 participates in c-Myc proteosomal degradation and acts as a cofactor for c-Myc-regulated transcription. Mol. Cell 11, 1189–1200 (2003).
Kim, S. Y., Herbst, A., Tworkowski, K. A., Salghetti, S. E. & Tansey, W. P. Skp2 regulates Myc protein stability and activity. Mol. Cell 11, 1177–1188 (2003).
Salghetti, S. E., Caudy, A. A., Chenoweth, J. G. & Tansey, W. P. Regulation of transcriptional activation domain function by ubiquitin. Science 293, 1651–1653 (2001).
Cardozo, T. & Pagano, M. The SCF ubiquitin ligase: insights into a molecular machine. Nature Rev. Mol. Cell Biol. 5, 739–751 (2004).
Pagano, M. & Benmaamar, R. When protein destruction runs amok, malignancy is on the loose. Cancer Cell 4, 251–256 (2003).
Guardavaccaro, D. & Pagano, M. Oncogenic aberrations of cullin-dependent ubiquitin ligases. Oncogene 23, 2037–2049 (2004).
Reed, S. I. Ratchets and clocks: the cell cycle, ubiquitylation and protein turnover. Nature Rev. Mol. Cell Biol. 4, 855–864 (2003).
Verma, R. et al. Phosphorylation of Sic1p by G1 Cdk required for its degradation and entry into S phase. Science 278, 455–460 (1997). Shows that phosphorylation underlies the targeting of substrates to SCF for ubiquitylation.
Lanker, S., Valdivieso, M. H. & Wittenberg, C. Rapid degradation of the G1 cyclin Cln2 induced by CDK-dependent phosphorylation. Science 271, 1597–1601 (1996).
Clurman, B. E., Sheaff, R. J., Thress, K., Groudine, M. & Roberts, J. M. Turnover of cyclin E by the ubiquitin-proteasome pathway is regulated by cdk2 binding and cyclin phosphorylation. Genes Dev. 10, 1979–1990 (1996).
Won, K. A. & Reed, S. I. Activation of cyclin E/CDK2 is coupled to site-specific autophosphorylation and ubiquitin-dependent degradation of cyclin E. EMBO J. 15, 4182–4193 (1996).
Sheaff, R. J., Groudine, M., Gordon, M., Roberts, J. M. & Clurman, B. E. Cyclin E–CDK2 is a regulator of p27Kip1. Genes Dev. 11, 1464–1478 (1997).
Yoshida, Y. et al. E3 ubiquitin ligase that recognizes sugar chains. Nature 418, 438–442 (2002).
Ivan, M. et al. HIFα targeted for VHL-mediated destruction by proline hydroxylation: implications for O2 sensing. Science 292, 464–468 (2001).
Jaakkola, P. et al. Targeting of HIF-α to the von Hippel–Lindau ubiquitylation complex by O2-regulated prolyl hydroxylation. Science 292, 468–472 (2001). References 68 and 69 show that HIF1α recognition by VHL requires proline hydroxylation, which indicates a new mechanism for regulating transcription through the sensing of oxygen levels.
Schwob, E., Bohm, T., Mendenhall, M. D. & Nasmyth, K. The B-type cyclin kinase inhibitor p40SIC1 controls the G1 to S transition in S. cerevisiae. Cell 79, 233–244 (1994). The genetic dissection of the G1-to-S transition in S. cerevisiae points the way to the discovery of SCF.
Verma, R., McDonald, H., Yates, J. R. 3rd & Deshaies, R. J. Selective degradation of ubiquitinated Sic1 by purified 26S proteasome yields active S phase cyclin–Cdk. Mol. Cell 8, 439–448 (2001).
Meimoun, A. et al. Degradation of the transcription factor Gcn4 requires the kinase Pho85 and the SCFCDC4 ubiquitin-ligase complex. Mol. Biol. Cell 11, 915–927 (2000).
Chi, Y. et al. Negative regulation of Gcn4 and Msn2 transcription factors by Srb10 cyclin-dependent kinase. Genes Dev. 15, 1078–1092 (2001).
Shemer, R., Meimoun, A., Holtzman, T. & Kornitzer, D. Regulation of the transcription factor Gcn4 by Pho85 cyclin PCL5. Mol. Cell. Biol. 22, 5395–5404 (2002).
Furstenthal, L., Swanson, C., Kaiser, B. K., Eldridge, A. G. & Jackson, P. K. Triggering ubiquitination of a CDK inhibitor at origins of DNA replication. Nature Cell Biol. 3, 715–722 (2001).
Montagnoli, A. et al. Ubiquitination of p27 is regulated by Cdk-dependent phosphorylation and trimeric complex formation. Genes Dev. 13, 1181–1189 (1999).
Cheng, M., Sexl, V., Sherr, C. J. & Roussel, M. F. Assembly of cyclin D-dependent kinase and titration of p27Kip1 regulated by mitogen-activated protein kinase kinase (MEK1). Proc. Natl Acad. Sci. USA 95, 1091–1096 (1998).
Welcker, M. et al. Multisite phosphorylation by Cdk2 and GSK3 controls cyclin E degradation. Mol. Cell 12, 381–392 (2003).
Klein, P., Pawson, T. & Tyers, M. Mathematical modeling suggests cooperative interactions between a disordered polyvalent ligand and a single receptor site. Curr. Biol. 13, 1669–1678 (2003).
Noureddine, M. A., Donaldson, T. D., Thacker, S. A. & Duronio, R. J. Drosophila Roc1a encodes a RING-H2 protein with a unique function in processing the Hh signal transducer Ci by the SCF E3 ubiquitin ligase. Dev. Cell 2, 757–770 (2002).
Furukawa, M., Ohta, T. & Xiong, Y. Activation of UBC5 ubiquitin-conjugating enzyme by the RING finger of ROC1 and assembly of active ubiquitin ligases by all cullins. J. Biol. Chem. 277, 15758–15765 (2002).
Strack, P. et al. SCFβ-TRCP and phosphorylation dependent ubiquitination of IκBα catalyzed by Ubc3 and Ubc4. Oncogene 19, 3529–3536 (2000).
Kamura, T., Conrad, M. N., Yan, Q., Conaway, R. C. & Conaway, J. W. The Rbx1 subunit of SCF and VHL E3 ubiquitin ligase activates Rub1 modification of cullins Cdc53 and Cul2. Genes Dev. 13, 2928–2933 (1999).
Liu, Y., Mathias, N., Steussy, C. N. & Goebl, M. G. Intragenic suppression among CDC34 (UBC3) mutations defines a class of ubiquitin-conjugating catalytic domains. Mol. Cell. Biol. 15, 5635–5644 (1995).
Ptak, C. et al. Functional and physical characterization of the cell cycle ubiquitin-conjugating enzyme CDC34 (UBC3). Identification of a functional determinant within the tail that facilitates CDC34 self-association. J. Biol. Chem. 269, 26539–26545 (1994).
Varelas, X., Ptak, C. & Ellison, M. J. Cdc34 self-association is facilitated by ubiquitin thiolester formation and is required for its catalytic activity. Mol. Cell. Biol. 23, 5388–5400 (2003).
Mathias, N., Steussy, C. N. & Goebl, M. G. An essential domain within Cdc34p is required for binding to a complex containing Cdc4p and Cdc53p in Saccharomyces cerevisiae. J. Biol. Chem. 273, 4040–4045 (1998).
Ptak, C. et al. Creation of a pluripotent ubiquitin-conjugating enzyme. Mol. Cell. Biol. 21, 6537–6548 (2001).
Deffenbaugh, A. E. et al. Release of ubiquitin-charged Cdc34-S∼Ub from the RING domain is essential for ubiquitination of the SCFCdc4-bound substrate Sic1. Cell 114, 611–622 (2003).
Petroski, M. D. & Deshaies, R. J. Context of multiubiquitin chain attachment influences the rate of Sic1 degradation. Mol. Cell 11, 1435–1444 (2003).
Blondel, M. et al. Nuclear-specific degradation of Far1 is controlled by the localization of the F-box protein Cdc4. EMBO J. 19, 6085–6097 (2000).
Hori, T. et al. Covalent modification of all members of human cullin family proteins by NEDD8. Oncogene 18, 6829–6834 (1999).
Osaka, F. et al. Covalent modifier NEDD8 is essential for SCF ubiquitin-ligase in fission yeast. EMBO J. 19, 3475–3484 (2000).
Podust, V. N. et al. A Nedd8 conjugation pathway is essential for proteolytic targeting of p27Kip1 by ubiquitination. Proc. Natl Acad. Sci. USA 97, 4579–4584 (2000).
Read, M. A. et al. Nedd8 modification of cul-1 activates SCFβ-TrCP-dependent ubiquitination of IκBα. Mol. Cell. Biol. 20, 2326–2333 (2000).
Wu, K., Chen, A. & Pan, Z. Q. Conjugation of Nedd8 to CUL1 enhances the ability of the ROC1–CUL1 complex to promote ubiquitin polymerization. J. Biol. Chem. 275, 32317–32324 (2000).
Kawakami, T. et al. NEDD8 recruits E2-ubiquitin to SCF E3 ligase. EMBO J. 20, 4003–4012 (2001).
Ohh, M. et al. An intact NEDD8 pathway is required for Cullin-dependent ubiquitylation in mammalian cells. EMBO Rep. 3, 177–182 (2002).
Pintard, L. et al. Neddylation and deneddylation of CUL-3 is required to target MEI-1/Katanin for degradation at the meiosis-to-mitosis transition in C. elegans. Curr. Biol. 13, 911–921 (2003).
Ou, C. Y., Lin, Y. F., Chen, Y. J. & Chien, C. T. Distinct protein degradation mechanisms mediated by Cul1 and Cul3 controlling Ci stability in Drosophila eye development. Genes Dev. 16, 2403–2414 (2002).
Tateishi, K., Omata, M., Tanaka, K. & Chiba, T. The NEDD8 system is essential for cell cycle progression and morphogenetic pathway in mice. J. Cell Biol. 155, 571–579 (2001).
Lammer, D. et al. Modification of yeast Cdc53p by the ubiquitin-related protein rub1p affects function of the SCFCdc4 complex. Genes Dev. 12, 914–926 (1998).
Liakopoulos, D., Doenges, G., Matuschewski, K. & Jentsch, S. A novel protein modification pathway related to the ubiquitin system. EMBO J. 17, 2208–2214 (1998). References 102 and 103 report the identification of the neddylation pathway, as well as evidence that modification of CUL1/Cdc53 by NEDD8 stimulates its function.
Lyapina, S. et al. Promotion of NEDD–CUL1 conjugate cleavage by COP9 signalosome. Science 292, 1382–1385 (2001).
Cope, G. A. et al. Role of predicted metalloprotease motif of Jab1/Csn5 in cleavage of Nedd8 from Cul1. Science 298, 608–611 (2002). References 104 and 105 describe how the CSN regulates the neddylated state of CUL1 through the cleavage of NEDD8 by its CSN5 subunit.
Ambroggio, X. I., Rees, D. C. & Deshaies, R. J. JAMM: a metalloprotease-like zinc site in the proteasome and signalosome. PLoS Biol. 2, E2 (2004).
Tran, H. J., Allen, M. D., Lowe, J. & Bycroft, M. Structure of the Jab1/MPN domain and its implications for proteasome function. Biochemistry 42, 11460–11465 (2003).
Cope, G. A. & Deshaies, R. J. COP9 signalosome: a multifunctional regulator of SCF and other cullin-based ubiquitin ligases. Cell 114, 663–671 (2003).
Zhou, C. et al. Fission yeast COP9/signalosome suppresses cullin activity through recruitment of the deubiquitylating enzyme Ubp12p. Mol. Cell 11, 927–938 (2003).
Liu, C. et al. Cop9/signalosome subunits and Pcu4 regulate ribonucleotide reductase by both checkpoint-dependent and-independent mechanisms. Genes Dev. 17, 1130–1140 (2003).
Schwechheimer, C. et al. Interactions of the COP9 signalosome with the E3 ubiquitin ligase SCFTIRI in mediating auxin response. Science 292, 1379–1382 (2001).
Doronkin, S., Djagaeva, I. & Beckendorf, S. K. The COP9 signalosome promotes degradation of Cyclin E during early Drosophila oogenesis. Dev. Cell 4, 699–710 (2003).
Liu, J., Furukawa, M., Matsumoto, T. & Xiong, Y. NEDD8 modification of CUL1 dissociates p120CAND1, an inhibitor of CUL1–SKP1 binding and SCF ligases. Mol. Cell 10, 1511–1518 (2002).
Zheng, J. et al. CAND1 binds to unneddylated CUL1 and regulates the formation of SCF ubiquitin E3 ligase complex. Mol. Cell 10, 1519–1526 (2002). References 113 and 114 report the identification of CAND1 as a regulator of CRL assembly.
Hwang, J. W., Min, K. W., Tamura, T. A. & Yoon, J. B. TIP120A associates with unneddylated cullin 1 and regulates its neddylation. FEBS Lett. 541, 102–108 (2003).
Min, K. W. et al. TIP120A associates with cullins and modulates ubiquitin ligase activity. J. Biol. Chem. 278, 15905–15910 (2003).
Oshikawa, K. et al. Preferential interaction of TIP120A with Cul1 that is not modified by NEDD8 and not associated with Skp1. Biochem. Biophys. Res. Commun. 303, 1209–1216 (2003).
Chuang, H. W., Zhang, W. & Gray, W. M. Arabidopsis ETA2, an apparent ortholog of the human cullin-interacting protein CAND1, is required for auxin responses mediated by the SCFTIR1 ubiquitin ligase. Plant Cell 16, 1883–1897 (2004).
Feng, S. et al. Arabidopsis CAND1, an unmodified CUL1-interacting protein, is involved in multiple developmental pathways controlled by ubiquitin/proteasome-mediated protein degradation. Plant Cell 16, 1870–1882 (2004).
Cheng, Y., Dai, X. & Zhao, Y. AtCAND1, a HEAT-repeat protein that participates in auxin signaling in Arabidopsis. Plant Physiol. 135, 1020–1026 (2004).
Davis, M. et al. Pseudosubstrate regulation of the SCFβTrCP ubiquitin ligase by hnRNP-U. Genes Dev. 16, 439–451 (2002).
Kitagawa, K., Skowyra, D., Elledge, S. J., Harper, J. W. & Hieter, P. SGT1 encodes an essential component of the yeast kinetochore assembly pathway and a novel subunit of the SCF ubiquitin ligase complex. Mol. Cell 4, 21–33 (1999).
Takahashi, A., Casais, C., Ichimura, K. & Shirasu, K. HSP90 interacts with RAR1 and SGT1 and is essential for RPS2-mediated disease resistance in Arabidopsis. Proc. Natl Acad. Sci. USA 100, 11777–11782 (2003).
Hubert, D. A. et al. Cytosolic HSP90 associates with and modulates the Arabidopsis RPM1 disease resistance protein. EMBO J. 22, 5679–5689 (2003).
Archambault, V. et al. Targeted proteomic study of the cyclin–cdk module. Mol. Cell 14, 699–711 (2004).
Jager, S., Strayle, J., Heinemeyer, W. & Wolf, D. H. Cic1, an adaptor protein specifically linking the 26S proteasome to its substrate, the SCF component Cdc4. EMBO J. 20, 4423–4431 (2001).
Ohlmeyer, J. T. & Schupbach, T. Encore facilitates SCF-ubiquitin-proteasome-dependent proteolysis during Drosophila oogenesis. Development 130, 6339–6349 (2003).
Liao, E. H., Hung, W., Abrams, B. & Zhen, M. An SCF-like ubiquitin ligase complex that controls presynaptic differentiation. Nature 430, 345–350 (2004).
Staropoli, J. F. et al. Parkin is a component of an SCF-like ubiquitin ligase complex and protects postmitotic neurons from kainate excitotoxicity. Neuron 37, 735–749 (2003).
Dornan, D. et al. The ubiquitin ligase COP1 is a critical negative regulator of p53. Nature 429, 86–92 (2004).
Kim, J. H. et al. SCFhFBH1 can act as helicase and E3 ubiquitin ligase. Nucleic Acids Res. 32, 2287–2297 (2004).
Koepp, D. M. et al. Phosphorylation-dependent ubiquitination of cyclin E by the SCFFbw7 ubiquitin ligase. Science 294, 173–177 (2001).
Strohmaier, H. et al. Human F-box protein hCdc4 targets cyclin E for proteolysis and is mutated in a breast cancer cell line. Nature 413, 316–322 (2001).
Nakayama, K. et al. Targeted disruption of Skp2 results in accumulation of cyclin E and p27Kip1, polyploidy and centrosome overduplication. EMBO J. 19, 2069–2081 (2000).
Singer, J. D., Gurian-West, M., Clurman, B. & Roberts, J. M. Cullin-3 targets cyclin E for ubiquitination and controls S phase in mammalian cells. Genes Dev. 13, 2375–2387 (1999).
Nateri, A. S., Riera-Sans, L., Da Costa, C. & Behrens, A. The ubiquitin ligase SCFFbw7 antagonizes apoptotic JNK signaling. Science 303, 1374–1378 (2004).
Cong, F., Zhang, J., Pao, W., Zhou, P. & Varmus, H. A protein knockdown strategy to study the function of β-catenin in tumorigenesis. BMC Mol. Biol. 4, 10 (2003).
Zhang, J., Zheng, N. & Zhou, P. Exploring the functional complexity of cellular proteins by protein knockout. Proc. Natl Acad. Sci. USA 100, 14127–14132 (2003).
Schneekloth, J. S. Jr. et al. Chemical genetic control of protein levels: selective in vivo targeted degradation. J. Am. Chem. Soc. 126, 3748–3754 (2004).
Jin, J. et al. Systematic analysis and nomenclature of mammalian F-box proteins. Genes Dev. 18, 2573–2580 (2004).
Acknowledgements
We thank C.-T. Chien, M. Estelle, E. Kipreos, M. Pagano and D. Wolf for their help with the online supplementary information S2 (table). We also thank G. Kleiger for preparing figure 2. M.D.P. is a postdoctoral fellow and R.J.D. is an assistant investigator of the Howard Hughes Medical Institute, which supports their work.
Author information
Authors and Affiliations
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
Related links
DATABASES
Flybase
Interpro
Protein Data Bank
Saccharomyces genome database
S. pombe gene database
Swiss-Prot
FURTHER INFORMATION
HGNC-Gene-Family Nomenclature — F-Box gene family
Glossary
- UBIQUITIN LIGASE (E3)
-
A protein or protein complex that facilitates the transfer of ubiquitin from the active-site cysteine of a ubiquitin-conjugating enzyme (E2) to a Lys residue of a substrate. Two main classes have been identified on the basis of the presence of either a HECT domain or a RING-like motif.
- CULLIN
-
A family of proteins that is characterized by the presence of a distinct globular C-terminal domain (cullin-homology domain) and a series of N-terminal repeats of a five-helix bundle (cullin repeats).
- SCF
-
A multisubunit ubiquitin ligase (E3). It consists of SKP1, CUL1 and an F-box protein that confers substrate specificity, as well as a RING protein that is also known as HRT1, RBX1 or ROC1.
- CULLIN–RING LIGASES
-
(CRLs). A superfamily of ubiquitin ligases that is characterized by an enzymatic core that contains a cullin-family member and a RING protein. The core is linked to specific substrates by adaptor proteins (or domains) and various receptor subunits.
- ANAPHASE-PROMOTING COMPLEX/CYCLOSOME
-
(APC/C). A multisubunit ubiquitin ligase that contains a RING subunit (APC11) and a distant cullin homologue (APC2). It has a key role in regulating the eukaryotic cell cycle.
- SKP1
-
(S-phase-kinase-associated protein-1). A 23-kDa protein that functions as an adaptor between CUL1 and F-box proteins. SKP1 was identified as a protein that, together with SKP2, associates with cyclin-A–CDK complexes. It might have other functions, such as binding to centromeres and regulating the assembly of vacuolar ATPases.
- RING-H2 DOMAIN
-
A protein motif that consists of a defined pattern of cysteine and histidine residues (Cys-X(2)-Cys-X(9–39)-Cys-X(1–3)-His-X(2–3)-His-X(2)-Cys-X(4–48)-Cys-X(2)-Cys; where X is any amino acid) and that coordinates two zinc molecules. This motif interacts directly with E2 enzymes. Proteins such as MDM2, which have variations of the basic RING motif, can nonetheless sustain ubiquitylation. In CRL RING proteins, the final Cys is an Asp.
- BTB DOMAIN
-
A domain that was identified in 'Broad-complex, Tramtrack and Bric-a-brac' proteins in D. melanogaster and that is usually located in the N-terminal region of proteins. Originally known as the POZ domain, this motif is found in several virus proteins. Numerous BTB-domain proteins contain a second protein–protein interaction motif, such as zinc-finger and Kelch motifs.
- F-BOX MOTIF
-
Originally identified in cyclin F, this structural motif adopts a fold that is similar to the BTB domain, binds to SKP1 and is found in receptors that assemble into CUL1-containing cullin–RING ligases. Proteins that contain the F-box motif are found in all eukaryotes.
- AUTOUBIQUITYLATION
-
This term is loosely used to refer to the covalent transfer of ubiquitins to a Lys of an E2 or E3 component of the ubiquitin-conjugating machinery. This mechanism might regulate the assembly of some cullin–RING ligases by causing the proteasome-mediated degradation of some substrate receptors when substrate levels are depleted.
- DEGRON
-
A portion of a protein that is necessary and sufficient to confer its degradation by the ubiquitin–proteasome system.
- LYS-48-LINKED POLYUBIQUITIN CHAIN
-
Ubiquitin polymers in which the ε-NH2 group of Lys48 of ubiquitin is linked by an isopeptide bond to the C terminus of the next ubiquitin molecule in the chain. Such a chain can either be unanchored or attached at its proximal (C-terminal) end to the ε-NH2 group of a substrate Lys.
- PROTEASOME
-
A 2-MDa protein complex that degrades ubiquitylated proteins in an ATP-dependent manner.
- ALLOVALENCY
-
A kinetic model that indicates that the affinity observed for some ligand–receptor interactions might increase nonlinearly depending on the polyvalency and flexibility of the ligand. This hypothesis has been proposed for Sic1, as increasing the number of phosphate groups on Sic1 from five to six significantly increases its binding affinity for Cdc4, even though Cdc4 seems to contain only one key phosphopeptide-binding site.
- HECT DOMAIN
-
('homologous to E6-AP C terminus' domain). HECT- and RING-domain-containing proteins represent the two main classes of E3 ubiquitin ligases. In contrast to RING ligases, HECT-domain ligases form an essential thioester intermediate with ubiquitin as it is being transferred from the E2 enzyme to the substrate.
- UBC4/5 FAMILY
-
Two related E2-enzyme families that are structurally distinct from UBC3, the E2 that has been shown, using genetics, to interact with SCF. Although UBC4 and UBC5 ubiquitylate SCF substrates in vitro, it is unclear if they do so in vivo.
- NEDD8
-
A small protein that is greater than 50% identical to ubiquitin and is conjugated as a single molecule to a specific Lys residue in all cullin-family members. ATP, a heterodimeric E1 (ULA1–UBA3) and the E2 UBC12 are required for the covalent attachment of NEDD8.
- 'HIT AND RUN' HYPOTHESIS
-
A mechanistic model for ubiquitin transfer by SCF ubiquitin ligases, which proposes that the dissociation of the E2 enzyme Cdc34 from SCF is required for ubiquitin transfer to the substrate.
- NEDDYLATION/DENEDDYLATION
-
The attachment/removal of NEDD8, in this case, to/from cullin-family members. This cyclical process might regulate the assembly and activity of cullin–RING ligases. It has recently been reported that the RING ligase MDM2 is also regulated by neddylation.
- COP9 SIGNALOSOME
-
(CSN). An eight-subunit complex that was originally identified in plants. It cleaves NEDD8 from cullins and also associates with the deubiquitylating enzyme UBP12.
- 'JAMM' MOTIF
-
A metalloprotease motif (His-X-His-X(10)-Asp) that was originally identified in the CSN5 subunit of the COP9 signalosome and the RPN11 subunit of 26S proteasome. The JAMM motif is thought to be directly involved in the cleavage of NEDD8 from cullins.
- CAND1/TIP120A
-
(cullin-associated and neddylation-dissociated protein-1/TATA-binding-protein interacting protein-120A). A protein that specifically associates with deneddylated cullins to sequester them in an unassembled and inactive state. Putative CAND1 homologues have been identified in most eukaryotic model organisms, except for Saccharomyces cerevisiae.
Rights and permissions
About this article
Cite this article
Petroski, M., Deshaies, R. Function and regulation of cullin–RING ubiquitin ligases. Nat Rev Mol Cell Biol 6, 9–20 (2005). https://doi.org/10.1038/nrm1547
Issue Date:
DOI: https://doi.org/10.1038/nrm1547
This article is cited by
-
Csn5 inhibits autophagy by regulating the ubiquitination of Atg6 and Tor to mediate the pathogenicity of Magnaporthe oryzae
Cell Communication and Signaling (2024)
-
Deciphering the role of neddylation in tumor microenvironment modulation: common outcome of multiple signaling pathways
Biomarker Research (2024)
-
ZSWIM4 regulates embryonic patterning and BMP signaling by promoting nuclear Smad1 degradation
EMBO Reports (2024)
-
CRL2APPBP2-mediated TSPYL2 degradation counteracts human mesenchymal stem cell senescence
Science China Life Sciences (2024)
-
Insights into the diverse mechanisms and effects of variant CUL3-induced familial hyperkalemic hypertension
Cell Communication and Signaling (2023)