Key Points
-
To protect the integrity of their genome, eukaryotic cells possess evolutionarily conserved surveillance mechanisms that are known as checkpoints, which constantly monitor the status and quality of chromosomal DNA and delay cell-cycle progression in response to replication stress or diverse types of DNA damage.
-
The genetically most vulnerable period of the cell cycle, the DNA-synthesis (S) phase, is protected against various genotoxic stresses by three checkpoints. These are the DNA-damage-induced, replication-independent, intra-S-phase checkpoint (the 'intra-S-phase checkpoint'), which is the main topic of this review; the replication-dependent S-phase checkpoint (the 'replication checkpoint'); and the replication-dependent S–M checkpoint (the 'S–M checkpoint'), which prevents mitotic entry when DNA is incompletely replicated.
-
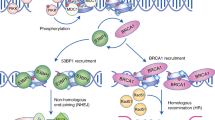
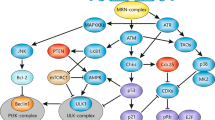
Recent discoveries reveal the distinct molecular mechanisms that activate the two most proximal checkpoint-signalling kinases ataxia-telangiectasia mutated (ATM) and ataxia-telangiectasia and RAD3 related (ATR), respectively. Furthermore, new evidence sheds light on the identity and function of candidate DNA-damage sensors, such as the NBS1–MRE11–RAD50 (MRN) complex (where NBS1 stands for Nijmegen-breakage-syndrome-1 and MRE11 stands for meiotic-recombination protein-11), as well as other components of the checkpoint-signalling cascades including the effectors. In response to DNA double-strand breaks (DSBs), the activated intra-S-phase checkpoint inhibits DNA replication through the combined action of the ATM/ATR–CHK2/CHK1–CDC25A–CDK2 and the ATM–NBS1–FANCD2/SMC1 effector pathways (where CHK stands for checkpoint kinase; CDK stands for cyclin-dependent kinase; FANCD2 stands for Fanconi anaemia complementation group D2; and SMC1 stands for structural maintenance of chromosomes-1).
-
A recently identified class of proximal checkpoint regulators, termed mediators, include p53-binding protein-1 (53BP1), mediator of DNA-damage checkpoint-1 (MDC1) and breast cancer susceptibility protein-1 (BRCA1) for the ATM-mediated responses, and claspin for ATR. They seem to serve as molecular match-makers that are critical for the timely and coordinated activation and maintenance of the checkpoint responses.
-
Live-cell, real-time imaging of early molecular events that are set in motion after formation of DSBs has provided valuable information about the spatio-temporal orchestration of the DSB response. This includes an unexpectedly dynamic behaviour of the checkpoint proteins that are involved in the recognition, processing, signalling and repair of the DNA lesions.
-
Defects in the genes that encode products that participate in the intra-S-phase checkpoint disable the checkpoint response, and thereby cause unscheduled DNA synthesis in irradiated cells (known as radioresistant DNA synthesis; RDS). Such checkpoint defects cause predisposition to a range of life-threatening human diseases including cancer.
Abstract
The precise replication of the genome and the continuous surveillance of its integrity are essential for survival and the avoidance of various diseases. Cells respond to DNA damage by activating a complex network of the so-called checkpoint pathways to delay their cell-cycle progression and repair the defects. In this review we integrate findings on the emerging mechanisms of activation, the signalling pathways and the spatio-temporal organization of the intra-S-phase DNA-damage checkpoint and its impact on the cell-cycle machinery, and discuss its biological significance.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Zhou, B. B. & Elledge, S. J. The DNA damage response: putting checkpoints in perspective. Nature 408, 433–439 (2000).
Nyberg, K. A., Michelson, R. J., Putnam, C. W. & Weinert, T. A. Toward maintaining the genome: DNA damage and replication checkpoints. Annu. Rev. Genet. 36, 617–656 (2002).
Hoeijmakers, J. H. Genome maintenance mechanisms for preventing cancer. Nature 411, 366–374 (2001).
Hartwell, L. H. & Weinert, T. A. Checkpoints: controls that ensure the order of cell cycle events. Science 246, 629–634 (1989).
Shiloh, Y. ATM and related protein kinases: safeguarding genome integrity. Nature Rev. Cancer 3, 155–168 (2003).
Abraham, R. T. Cell cycle checkpoint signaling through the ATM and ATR kinases. Genes Dev. 15, 2177–2196 (2001).
Bartek, J. & Lukas, J. Pathways governing G1/S transition and their response to DNA damage. FEBS Lett. 490, 117–122 (2001).
Lukas, C. et al. DNA damage-activated kinase Chk2 is independent of proliferation or differentiation yet correlates with tissue biology. Cancer Res. 61, 4990–4993 (2001).
Latella, L., Lukas, J., Simone, C., Puri, P. L. & Bartek, J. Differentiation-induced radioresistance in muscle cells. Mol. Cell. Biol. 24, 6350–6361 (2004).
Myung, K., Datta, A. & Kolodner, R. D. Suppression of spontaneous chromosomal rearrangements by S phase checkpoint functions in Saccharomyces cerevisiae. Cell 104, 397–408 (2001). This study, together with other reports from the same laboratory (see references 11 and 106), provides important genetic evidence that in the yeast Saccharomyces cerevisiae , the intra-S-phase checkpoint constitutes an important barrier against gross chromosome rearrangements.
Myung, K., Chen, C. & Kolodner, R. D. Multiple pathways cooperate in the suppression of genome instability in Saccharomyces cerevisiae. Nature 411, 1073–1076 (2001).
West, S. C. Molecular views of recombination proteins and their control. Nature Rev. Mol. Cell Biol. 4, 435–445 (2003).
Painter, R. B. & Young, B. R. Radiosensitivity in ataxia-telangiectasia: a new explanation. Proc. Natl Acad. Sci. USA 77, 7315–7317 (1980).
Merrick, C. J., Jackson, D. & Diffley, J. F. Visualization of altered replication dynamics after DNA damage in human cells. J. Biol. Chem. 279, 20067–20075 (2004).
Costanzo, V. et al. Reconstitution of an ATM-dependent checkpoint that inhibits chromosomal DNA replication following DNA damage. Mol. Cell 6, 649–659 (2000).
Falck, J., Petrini, J. H., Williams, B. R., Lukas, J. & Bartek, J. The DNA damage-dependent intra-S phase checkpoint is regulated by parallel pathways. Nature Genet. 30, 290–294 (2002). Shows that the DSB-induced inhibition of DNA replication is regulated by two, at least partially independent, mechanisms that comprise the ATM–NBS1–SMC1 and ATM–CHK2–CDC25A–CDK2 effector pathways. An analogous 'dual' regulation operates in the intra-S-phase response to DNA interstrand crosslinks (see reference 90).
Shiloh, Y. & Kastan, M. B. ATM: genome stability, neuronal development, and cancer cross paths. Adv. Cancer Res. 83, 209–254 (2001).
Bakkenist, C. J. & Kastan, M. B. DNA damage activates ATM through intermolecular autophosphorylation and dimer dissociation. Nature 421, 499–506 (2003). A landmark study that describes the rapid activation of ATM via intermolecular autophosphorylation on serine 1981. After publication of this paper, the antibody directed against phosphorylated serine 1981 has become a marker of ATM activation in various assays.
Zou, L. & Elledge, S. J. Sensing DNA damage through ATRIP recognition of RPA–ssDNA complexes. Science 300, 1542–1548 (2003). Shows that the RPA-coated single-stranded DNA serves as the specific signal to recruit ATM–ATRIP heterodimers to damaged DNA and/or stalled replication forks. This evolutionarily conserved mechanism is crucial for efficient substrate recognition and checkpoint initiation by ATR.
Lukas, C., Falck, J., Bartkova, J., Bartek, J. & Lukas, J. Distinct spatiotemporal dynamics of mammalian checkpoint regulators induced by DNA damage. Nature Cell Biol. 5, 255–260 (2003). Shows that NBS1 remains concentrated in the damaged nuclear compartments through a dynamic exchange between DSBs and the neighbouring nucleoplasm, whereas CHK2, after a transient 'collision' with DSBs and phosphorylation by ATM, rapidly spreads throughout the nucleus.
Ali, A. et al. Requirement of protein phosphatase 5 in DNA-damage-induced ATM activation. Genes Dev. 18, 249–254 (2004).
D'Amours, D. & Jackson, S. P. The Mre11 complex: at the crossroads of DNA repair and checkpoint signalling. Nature Rev. Mol. Cell Biol. 3, 317–327 (2002).
Petrini, J. H. & Stracker, T. H. The cellular response to DNA double-strand breaks: defining the sensors and mediators. Trends Cell Biol. 13, 458–462 (2003).
Uziel, T. et al. Requirement of the MRN complex for ATM activation by DNA damage. EMBO J. 22, 5612–5621 (2003). The first of a series of studies (see references 25–29) describing the role of the MRN complex in modulating the ATM activity in cells exposed to DSB-generating insults, thereby explaining the clinical resemblance between A-T and A-TLD patients (see also reference 29).
Carson, C. T. et al. The Mre11 complex is required for ATM activation and the G2/M checkpoint. EMBO J. 22, 6610–6620 (2003).
Horejsi, Z. et al. Distinct functional domains of Nbs1 modulate the timing and magnitude of ATM activation after low doses of ionizing radiation. Oncogene 23, 3122–3127 (2004).
Mochan, T. A., Venere, M., DiTullio, R. A. Jr. & Halazonetis, T. D. 53BP1 and NFBD1/MDC1–Nbs1 function in parallel interacting pathways activating ataxia-telangiectasia mutated (ATM) in response to DNA damage. Cancer Res. 63, 8586–8591 (2003).
Costanzo, V., Paull, T., Gottesman, M. & Gautier, J. Mre11 assembles linear DNA fragments into DNA damage signaling complexes. PLoS Biol. 2, E110 (2004).
Lee, J. H. & Paull, T. T. Direct activation of the ATM protein kinase by the Mre11/Rad50/Nbs1 complex. Science 304, 93–96 (2004). An interesting in vitro study that shows multiple direct interactions between the MRN complex and ATM, and provides evidence for a direct stimulation of ATM kinase activity by purified MRN.
Grenon, M., Gilbert, C. & Lowndes, N. F. Checkpoint activation in response to double-strand breaks requires the Mre11/Rad50/Xrs2 complex. Nature Cell Biol. 3, 844–847 (2001).
Nakada, D., Matsumoto, K. & Sugimoto, K. ATM-related Tel1 associates with double-strand breaks through an Xrs2-dependent mechanism. Genes Dev. 17, 1957–1962 (2003).
Zou, L., Cortez, D. & Elledge, S. J. Regulation of ATR substrate selection by Rad17-dependent loading of Rad9 complexes onto chromatin. Genes Dev. 16, 198–208 (2002).
Osborn, A. J., Elledge, S. J. & Zou, L. Checking on the fork: the DNA-replication stress-response pathway. Trends Cell Biol. 12, 509–516 (2002).
Cliby, W. A. et al. Overexpression of a kinase-inactive ATR protein causes sensitivity to DNA-damaging agents and defects in cell cycle checkpoints. EMBO J. 17, 159–169 (1998).
Cortez, D., Guntuku, S., Qin, J. & Elledge, S. J. ATR and ATRIP: partners in checkpoint signaling. Science 294, 1713–1716 (2001).
Sorensen, C. S., Syljuasen, R. G., Lukas, J. & Bartek, J. ATR, Claspin and the Rad9–Rad1–Hus1 complex regulate Chk1 and Cdc25A in the absence of DNA damage. Cell Cycle 3, 941–945 (2004).
Shechter, D., Costanzo, V. & Gautier, J. ATR and ATM regulate the timing of DNA replication origin firing. Nature Cell Biol. 6, 648–655 (2004). This study shows that the initiation of DNA replication in egg extracts from Xenopus laevis requires the ATR and ATM kinases in the absence of DNA damage. Together with references 36, 38 and 78, this study provides evidence that cell-cycle checkpoints operate as an amplification of the physiological processes inherently coupled to structures and/or signals generated by active replicons.
Marheineke, K. & Hyrien, O. Control of replication origin density and firing time in Xenopus egg extracts: role of a caffeine-sensitive, ATR-dependent checkpoint. J. Biol. Chem. 279, 28071–28081 (2004).
Brown, E. J. & Baltimore, D. ATR disruption leads to chromosomal fragmentation and early embryonic lethality. Genes Dev. 14, 397–402 (2000).
de Klein, A. et al. Targeted disruption of the cell-cycle checkpoint gene ATR leads to early embryonic lethality in mice. Curr. Biol. 10, 479–482 (2000).
Liu, Q. et al. Chk1 is an essential kinase that is regulated by Atr and required for the G2/M DNA damage checkpoint. Genes Dev. 14, 1448–1459 (2000).
Takai, H. et al. Aberrant cell cycle checkpoint function and early embryonic death in Chk1−/− mice. Genes Dev. 14, 1439–1447 (2000).
Goldberg, M. et al. MDC1 is required for the intra-S-phase DNA damage checkpoint. Nature 421, 952–956 (2003).
Lou, Z., Minter-Dykhouse, K., Wu, X. & Chen, J. MDC1 is coupled to activated CHK2 in mammalian DNA damage response pathways. Nature 421, 957–961 (2003).
Stewart, G. S., Wang, B., Bignell, C. R., Taylor, A. M. & Elledge, S. J. MDC1 is a mediator of the mammalian DNA damage checkpoint. Nature 421, 961–966 (2003).
Ozaki, T. et al. NFBD1/KIAA0170 is a novel nuclear transcriptional transactivator with BRCT domain. DNA Cell Biol. 19, 475–485 (2000).
Shang, Y. L., Bodero, A. J. & Chen, P. L. NFBD1, a novel nuclear protein with signature motifs of FHA and BRCT, and an internal 41-amino acid repeat sequence, is an early participant in DNA damage response. J. Biol. Chem. 278, 6323–6329 (2003).
Xu, X. & Stern, D. F. NFBD1/KIAA0170 is a chromatin-associated protein involved in DNA damage signaling pathways. J. Biol. Chem. 278, 8795–8803 (2003).
DiTullio, R. A. Jr. et al. 53BP1 functions in an ATM-dependent checkpoint pathway that is constitutively activated in human cancer. Nature Cell Biol. 4, 998–1002 (2002).
Ward, I. M., Minn, K., van Deursen, J. & Chen, J. p53 binding protein 53BP1 is required for DNA damage responses and tumor suppression in mice. Mol. Cell. Biol. 23, 2556–2563 (2003).
Wang, B., Matsuoka, S., Carpenter, P. B. & Elledge, S. J. 53BP1, a mediator of the DNA damage checkpoint. Science 298, 1435–1438 (2002).
Venkitaraman, A. R. Cancer susceptibility and the functions of BRCA1 and BRCA2. Cell 108, 171–182 (2002).
Kitagawa, R., Bakkenist, C. J., McKinnon, P. J. & Kastan, M. B. Phosphorylation of SMC1 is a critical downstream event in the ATM–NBS1–BRCA1 pathway. Genes Dev. 18, 1423–1438 (2004). This study provides further evidence that ATM becomes activated outside DSBs and that its subsequent recruitment to the sites of DNA damage requires NBS1 and BRCA1.
Xu, B., Kim, S. & Kastan, M. B. Involvement of Brca1 in S-phase and G2-phase checkpoints after ionizing irradiation. Mol. Cell. Biol. 21, 3445–3450 (2001).
Kumagai, A. & Dunphy, W. G. Claspin, a novel protein required for the activation of Chk1 during a DNA replication checkpoint response in Xenopus egg extracts. Mol. Cell 6, 839–849 (2000).
Lee, J., Kumagai, A. & Dunphy, W. G. Claspin, a Chk1-regulatory protein, monitors DNA replication on chromatin independently of RPA, ATR, and Rad17. Mol. Cell 11, 329–340 (2003).
Lukas, C. et al. Mdc1 couples DNA double-strand break recognition by Nbs1 with its H2AX-dependent chromatin retention. EMBO J. 23, 2674–2683 (2004). This study describes the real-time recruitment kinetics of NBS1 and MDC1 to the laser-generated DSB sites and uncovers two distinct modes of interaction between NBS1 and the DSB-harbouring nuclear regions.
Rouse, J. & Jackson, S. P. Interfaces between the detection, signaling, and repair of DNA damage. Science 297, 547–551 (2002).
Lou, Z., Chini, C. C., Minter-Dykhouse, K. & Chen, J. Mediator of DNA damage checkpoint protein 1 regulates BRCA1 localization and phosphorylation in DNA damage checkpoint control. J. Biol. Chem. 278, 13599–13602 (2003).
Cerosaletti, K. M. & Concannon, P. Nibrin forkhead-associated domain and breast cancer C-terminal domain are both required for nuclear focus formation and phosphorylation. J. Biol. Chem. 278, 21944–21951 (2003).
Desai-Mehta, A., Cerosaletti, K. M. & Concannon, P. Distinct functional domains of nibrin mediate Mre11 binding, focus formation, and nuclear localization. Mol. Cell. Biol. 21, 2184–2191 (2001).
Manke, I. A., Lowery, D. M., Nguyen, A. & Yaffe, M. B. BRCT repeats as phosphopeptide-binding modules involved in protein targeting. Science 302, 636–639 (2003).
Yu, X., Chini, C. C., He, M., Mer, G. & Chen, J. The BRCT domain is a phospho-protein binding domain. Science 302, 639–642 (2003).
Houtsmuller, A. B. et al. Action of DNA repair endonuclease ERCC1/XPF in living cells. Science 284, 958–961 (1999). A pioneering study that describes the kinetic behaviour of a DNA-damage regulator (ERCC1/XPF) in living mammalian cells.
Rademakers, S. et al. Xeroderma pigmentosum group A protein loads as a separate factor onto DNA lesions. Mol. Cell. Biol. 23, 5755–5767 (2003).
Van Den Boom, V. et al. DNA damage stabilizes interaction of CSB with the transcription elongation machinery. J. Cell Biol. 166, 27–36 (2004).
Essers, J. et al. Nuclear dynamics of RAD52 group homologous recombination proteins in response to DNA damage. EMBO J. 21, 2030–2037 (2002).
Kim, S. T., Xu, B. & Kastan, M. B. Involvement of the cohesin protein, Smc1, in Atm-dependent and independent responses to DNA damage. Genes Dev. 16, 560–570 (2002).
Yazdi, P. T. et al. SMC1 is a downstream effector in the ATM/NBS1 branch of the human S-phase checkpoint. Genes Dev. 16, 571–582 (2002).
Rogakou, E. P., Boon, C., Redon, C. & Bonner, W. M. Megabase chromatin domains involved in DNA double-strand breaks in vivo. J. Cell Biol. 146, 905–916 (1999).
Celeste, A. et al. Histone H2AX phosphorylation is dispensable for the initial recognition of DNA breaks. Nature Cell Biol. 5, 675–679 (2003). Shows that the initial recruitment of DNA repair and signalling proteins to the DSB sites does not require H2AX phosphorylation. It was proposed that although γ-H2AX does not constitute the primary DSB-recognition signal, its function is required to concentrate the DSB regulators in the vicinity of the DNA lesions.
Bartek, J. & Lukas, J. Chk1 and Chk2 kinases in checkpoint control and cancer. Cancer Cell 3, 421–429 (2003).
Donzelli, M. & Draetta, G. F. Regulating mammalian checkpoints through Cdc25 inactivation. EMBO Rep. 4, 671–677 (2003).
Mailand, N. et al. Rapid destruction of human Cdc25A in response to DNA damage. Science 288, 1425–1429 (2000).
Molinari, M., Mercurio, C., Dominguez, J., Goubin, F. & Draetta, G. F. Human Cdc25 A inactivation in response to S phase inhibition and its role in preventing premature mitosis. EMBO Rep. 1, 71–79 (2000).
Falck, J., Mailand, N., Syljuasen, R. G., Bartek, J. & Lukas, J. The ATM–Chk2–Cdc25A checkpoint pathway guards against radioresistant DNA synthesis. Nature 410, 842–847 (2001).
Mailand, N. et al. Regulation of G2/M events by Cdc25A through phosphorylation-dependent modulation of its stability. EMBO J. 21, 5911–5920 (2002).
Sorensen, C. S. et al. Chk1 regulates the S phase checkpoint by coupling the physiological turnover and ionizing radiation-induced accelerated proteolysis of Cdc25A. Cancer Cell 3, 247–258 (2003).
Zhao, H., Watkins, J. L. & Piwnica-Worms, H. Disruption of the checkpoint kinase 1/cell division cycle 25A pathway abrogates ionizing radiation-induced S and G2 checkpoints. Proc. Natl Acad. Sci. USA 99, 14795–14800 (2002).
Donzelli, M. et al. Dual mode of degradation of Cdc25 A phosphatase. EMBO J. 21, 4875–4884 (2002).
Skowyra, D., Craig, K. L., Tyers, M., Elledge, S. J. & Harper, J. W. F-box proteins are receptors that recruit phosphorylated substrates to the SCF ubiquitin-ligase complex. Cell 91, 209–219 (1997).
Kipreos, E. T. & Pagano, M. The F-box protein family. Genome Biol. 1, REVIEWS3002 (2000).
Jackson, P. K. & Eldridge, A. G. The SCF ubiquitin ligase: an extended look. Mol. Cell 9, 923–925 (2002).
Busino, L. et al. Degradation of Cdc25A by β-TrCP during S phase and in response to DNA damage. Nature 426, 87–91 (2003).
Jin, J. et al. SCFβ-TRCP links Chk1 signaling to degradation of the Cdc25A protein phosphatase. Genes Dev. 17, 3062–3074 (2003). References 84 and 85 identified SCFβ-TrCP1/2 as the long-sought ubiquitin ligase for mammalian CDC25A.
Hirao, A. et al. Chk2 is a tumor suppressor that regulates apoptosis in both an ataxia telangiectasia mutated (ATM)-dependent and an ATM-independent manner. Mol. Cell. Biol. 22, 6521–6532 (2002).
Takai, H. et al. Chk2-deficient mice exhibit radioresistance and defective p53-mediated transcription. EMBO J. 21, 5195–5205 (2002).
Nakanishi, K. et al. Interaction of FANCD2 and NBS1 in the DNA damage response. Nature Cell Biol. 4, 913–920 (2002).
Dierov, J., Dierova, R. & Carroll, M. BCR/ABL translocates to the nucleus and disrupts an ATR-dependent intra-S phase checkpoint. Cancer Cell 5, 275–285 (2004).
Pichierri, P. & Rosselli, F. The DNA crosslink-induced S-phase checkpoint depends on ATR-CHK1 and ATR–NBS1–FANCD2 pathways. EMBO J. 23, 1178–1187 (2004).
Fernandez-Capetillo, O., Allis, C. D. & Nussenzweig, A. Phosphorylation of histone H2B at DNA double-strand breaks. J. Exp. Med. 199, 1671–1677 (2004).
Jackson, D. A. & Pombo, A. Replicon clusters are stable units of chromosome structure: evidence that nuclear organization contributes to the efficient activation and propagation of S phase in human cells. J. Cell Biol. 140, 1285–1295 (1998).
Ma, H. et al. Spatial and temporal dynamics of DNA replication sites in mammalian cells. J. Cell Biol. 143, 1415–1425 (1998).
Jessberger, R. The many functions of SMC proteins in chromosome dynamics. Nature Rev. Mol. Cell Biol. 3, 767–778 (2002).
Fernandez-Capetillo, O. et al. DNA damage-induced G2–M checkpoint activation by histone H2AX and 53BP1. Nature Cell Biol. 4, 993–997 (2002).
Zhou, B. B. & Bartek, J. Targeting the checkpoint kinases: chemosensitization versus chemoprotection. Nature Rev. Cancer 4, 216–225 (2004).
O'Driscoll, M., Ruiz-Perez, V. L., Woods, C. G., Jeggo, P. A. & Goodship, J. A. A splicing mutation affecting expression of ataxia-telangiectasia and Rad3-related protein (ATR) results in Seckel syndrome. Nature Genet. 33, 497–501 (2003).
Groth, A. et al. Human Tousled like kinases are targeted by an ATM- and Chk1-dependent DNA damage checkpoint. EMBO J. 22, 1676–1687 (2003).
Krause, D. R. et al. Suppression of Tousled-like kinase activity after DNA damage or replication block requires ATM, NBS1 and Chk1. Oncogene 22, 5927–5937 (2003).
Costanzo, V. et al. An ATR- and Cdc7-dependent DNA damage checkpoint that inhibits initiation of DNA replication. Mol. Cell 11, 203–213 (2003).
Jares, P., Donaldson, A. & Blow, J. J. The Cdc7/Dbf4 protein kinase: target of the S phase checkpoint? EMBO Rep. 1, 319–322 (2000).
Dimitrova, D. S. & Gilbert, D. M. Temporally coordinated assembly and disassembly of replication factories in the absence of DNA synthesis. Nature Cell Biol. 2, 686–694 (2000).
Lopes, M. et al. The DNA replication checkpoint response stabilizes stalled replication forks. Nature 412, 557–561 (2001).
Tercero, J. A. & Diffley, J. F. Regulation of DNA replication fork progression through damaged DNA by the Mec1/Rad53 checkpoint. Nature 412, 553–557 (2001). References 103 and 104 provide evidence that, in addition to the inhibition of late replication origins, the key function of the checkpoint signalling elicited by intra-S-phase DNA damage and/or replication stalling is to protect the integrity of the existing replication forks.
Shimada, K., Pasero, P. & Gasser, S. M. ORC and the intra-S-phase checkpoint: a threshold regulates Rad53p activation in S phase. Genes Dev. 16, 3236–3252 (2002).
Myung, K. & Kolodner, R. D. Suppression of genome instability by redundant S-phase checkpoint pathways in Saccharomyces cerevisiae. Proc. Natl Acad. Sci. USA 99, 4500–4507 (2002).
Brown, E. J. The ATR-independent DNA replication checkpoint. Cell Cycle 2, 188–189 (2003).
Goloudina, A. et al. Regulation of human Cdc25A stability by Serine 75 phosphorylation is not sufficient to activate a S phase checkpoint. Cell Cycle 2, 473–478 (2003).
Hassepass, I., Voit, R. & Hoffmann, I. Phosphorylation at serine 75 is required for UV-mediated degradation of human Cdc25A phosphatase at the S-phase checkpoint. J. Biol. Chem. 278, 29824–29829 (2003).
Galaktionov, K. & Beach, D. Specific activation of cdc25 tyrosine phosphatases by B-type cyclins: evidence for multiple roles of mitotic cyclins. Cell 67, 1181–1194 (1991).
Limoli, C. L. & Ward, J. F. A new method for introducing double-strand breaks into cellular DNA. Radiat. Res. 134, 160–169 (1993).
Brown, K. D. et al. The mismatch repair system is required for S-phase checkpoint activation. Nature Genet. 33, 80–84 (2003).
Cejka, P., Stojic, L., Marra, G. & Jiricny, J. Is mismatch repair really required for ionizing radiation-induced DNA damage signaling? Nature Genet. 36, 432–433 (2004).
Acknowledgements
The authors' research is supported by the Danish Cancer Society and the European Union. The authors apologize to colleagues whose work could only be cited indirectly due to space limitations.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary information S1
Supplementary information S1 (movie) |A real-time monitoring of NBS1 and CHK2 redistribution in response to spatially restricted DSBs. Human osteosarcoma cells (U-2-OS) that express low levels of green fluorescent protein (GFP)-tagged checkpoint kinase-2 (CHK2) and yellow fluorescent protein (YFP)-tagged Nijmegen-breakage-syndrome-1 (NBS1) were mixed and micro-irradiated within the linear subnuclear compartments by a focused laser beam (λ=337 nm). When combined with the presensitization of cells with halogenated thymidine analogues, this treatment results in the generation of DNA double-strand breaks (DSBs) or single-strand breaks specifically in the laser-exposed nuclear areas1. Whereas the NBS1 protein underwent a rapid (on the order of seconds) recruitment to, and accumulation around, the damaged nuclear areas, CHK2 remained highly mobile and disseminated throughout the nucleoplasm. Parallel immunostaining with phospho-specific antibodies confirmed that after a very transient 'collision' with the freshly generated DSBs, CHK2 was distributed throughout the nucleus in its activated (ATM–phosphorylated) form2. Red arrows indicate the laser tracks; the entire movie spans the first 10 minutes after micro-irradiation. (MP4 93 kb)
1. Limoli, C. L. & Ward, J. F. A new method for introducing double-strand breaks into cellular DNA. Radiat. Res. 134, 160–169 (1993). 2. Lukas, C., Falck, J., Bartkova, J., Bartek, J. & Lukas, J. Distinct spatiotemporal dynamics of mammalian checkpoint regulators induced by DNA damage. Nature Cell Biol. 5, 255–260 (2003).
Glossary
- INTRA-S-PHASE CHECKPOINT
-
Replication-independent mechanism that reduces the rate of DNA synthesis in response to DNA double-strand breaks.
- REPLICATION CHECKPOINT
-
A mechanism that blocks S-phase progression in cells that are experiencing stalled replication forks.
- S–M CHECKPOINT
-
A cellular mechanism that prevents mitotic onset in cells with incompletely replicated DNA.
- RADIOMIMETIC DRUG
-
A chemical that has similar effects as ionizing radiation, causing the generation of DNA double-strand breaks.
- REPLICON
-
A unit of chromosomal DNA that is replicated by initiation at a single replication origin.
- REPLICATION ORIGIN
-
A site on chromosomal DNA where replicative DNA synthesis is initiated. Replication origins in human cells are divided into more than 1,000 so-called replication zones, each of which is activated at a characteristic time during S phase.
- RADIORESISTANT DNA SYNTHESIS
-
(RDS). The inability to slow down DNA replication in response to ionizing radiation and/or radiomimetic drugs.
- INTERMOLECULAR AUTOPHOSPHORYLATION
-
The exchange of phosphate groups between two identical protein kinases.
- MRN NUCLEASE COMPLEX
-
A protein complex that comprises the MRE11, RAD50 and NBS1 subunits, and that is involved in the recognition, processing, signalling and repair of DNA double-strand breaks. The nuclease activity is associated with the MRE11 subunit.
- 9-1-1 COMPLEX
-
A heterotrimeric protein complex that comprises the RAD9, RAD1 and HUS1 subunits, and that forms a 'doughnut-shaped' sliding clamp around DNA after damage and/or replication-fork stalling. Its loading onto DNA requires RAD17 (the so-called 'clamp loader' protein).
- REPLICATION PROTEIN A
-
(RPA). A protein that binds and protects single-stranded DNA.
- FHA DOMAIN
-
The forkhead-associated domain is a phospho-threonine-binding motif that was first identified in forkhead transcription factors and, subsequently, in other classes of proteins — including those that are involved in checkpoint signalling.
- BRCT DOMAIN
-
An evolutionarily conserved phospho-serine/threonine-interaction motif that was first identified in the carboxy-terminal part of BRCA1 and, subsequently, in several other checkpoint mediators.
- COHESIN
-
A proteinaceous ring that consists of four subunits (SMC1, SMC3, SCC1, SCC3) and that forms around the replicated DNA strands. Cohesin prevents premature separation of the nascent sister chromatids and some of its components (SMC1) participate in DNA recombination.
- F-BOX PROTEINS
-
A family of proteins that function as the substrate-targeting protein subunits of the SCF (SKP1–CUL1–F-box protein) ubiquitin-ligase complex.
- PHOSPHODEGRON MOTIF
-
A protein domain that, on phosphorylation, serves as a docking site for a specific ubiquitin ligase.
- PSORALEN
-
A compound that sensitizes mammalian cells to ultraviolet radiation.
Rights and permissions
About this article
Cite this article
Bartek, J., Lukas, C. & Lukas, J. Checking on DNA damage in S phase. Nat Rev Mol Cell Biol 5, 792–804 (2004). https://doi.org/10.1038/nrm1493
Issue Date:
DOI: https://doi.org/10.1038/nrm1493
This article is cited by
-
Ubiquitin-specific protease 28: the decipherment of its dual roles in cancer development
Experimental Hematology & Oncology (2023)
-
Vascular and pulmonary effects of ibuprofen on neonatal lung development
Respiratory Research (2023)
-
The oncogenic fusion protein TAZ::CAMTA1 promotes genomic instability and senescence through hypertranscription
Communications Biology (2023)
-
Pyrazole-sulfonamide scaffold featuring dual-tail strategy as apoptosis inducers in colon cancer
Scientific Reports (2023)
-
Immortalization-upregulated protein promotes pancreatic cancer progression by regulating NPM1/FHL1-mediated cell-cycle-checkpoint protein activity
Cell Biology and Toxicology (2023)