Key Points
-
The human genome harbours a dizzying array of regulatory sequences, such as enhancers with a priori unpredictable, promiscuous and context-dependent behaviour.
-
More than 90% of disease-associated variants lie in non-coding DNA, accumulating in presumptive enhancers. Therefore, a picture is emerging in which diseases and traits are often the consequence of erroneous wiring of regulatory circuitry between enhancers and their target genes.
-
Enhancers communicate with distant target genes through chromatin loops. Such transcription regulatory loops are formed within topologically associated domains (TADs), the structural and functional units of the chromosomes.
-
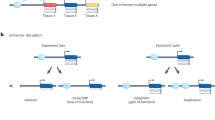
Disrupting enhancer–promoter loops, as well as TAD boundaries, can lead to gene dysregulation and disease, the latter due to previously insulated genes and enhancers being released to form new regulatory chromatin loops.
-
Technological advances and the availability of increasingly detailed genomic contact maps generated by Hi-C are enabling the elucidation of the physical contacts between genes and non-coding sequences in different cell types, which greatly facilitates the assigning of function to disease-associated genetic variation.
-
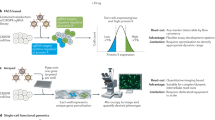
We present a roadmap — from disease-associated genetic variants to molecular mechanisms causing disease — that proposes to integrate epigenetic data with detailed 3D genome information and haplotype-resolved expression analysis to help move results descriptive genetic association studies towards the discovery of the molecular mechanisms underlying disease.
Abstract
Genetic variation associated with disease often appears in non-coding parts of the genome. Understanding the mechanisms by which this phenomenon leads to disease is necessary to translate results from genetic association studies to the clinic. Assigning function to this type of variation is notoriously difficult because the human genome harbours a complex regulatory landscape with a dizzying array of transcriptional regulatory sequences, such as enhancers that have unpredictable, promiscuous and context-dependent behaviour. In this Review, we discuss how technological advances have provided increasingly detailed information on genome folding; for example, genome folding forms loops that bring enhancers and target genes into close proximity. We also now know that enhancers function within topologically associated domains, which are structural and functional units of chromosomes. Studying disease-associated mutations and chromosomal rearrangements in the context of the 3D genome will enable the identification of dysregulated target genes and aid the progression from descriptive genetic association results to discovering molecular mechanisms underlying disease.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Pauling, L. et al. Sickle cell anemia a molecular disease. Science 110, 543–548 (1949).
Ingram, V. M. Gene mutations in human haemoglobin: the chemical difference between normal and sickle cell haemoglobin. Nature 180, 326–328 (1957).
Fritsch, E. F., Lawn, R. M. & Maniatis, T. Characterisation of deletions which affect the expression of fetal globin genes in man. Nature 279, 598–603 (1979).
Van der Ploeg, L. H. et al. γ-Β-Thalassaemia studies showing that deletion of the γ- and δ-genes influences β-globin gene expression in man. Nature 283, 637–642 (1980).
Banerji, J., Rusconi, S. & Schaffner, W. Expression of a β-globin gene is enhanced by remote SV40 DNA sequences. Cell 27, 299–308 (1981).
Benoist, C. & Chambon, P. In vivo sequence requirements of the SV40 early promotor region. Nature 290, 304–310 (1981).
Banerji, J., Olson, L. & Schaffner, W. A lymphocyte-specific cellular enhancer is located downstream of the joining region in immunoglobulin heavy chain genes. Cell 33, 729–740 (1983).
Gillies, S. D., Morrison, S. L., Oi, V. T. & Tonegawa, S. A tissue-specific transcription enhancer element is located in the major intron of a rearranged immunoglobulin heavy chain gene. Cell 33, 717–728 (1983).
Mercola, M., Wang, X. F., Olsen, J. & Calame, K. Transcriptional enhancer elements in the mouse immunoglobulin heavy chain locus. Science 221, 663–665 (1983).
Lander, E. S. et al. Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001).
Venter, J. C. et al. The sequence of the human genome. Science 291, 1304–1351 (2001).
Maurano, M. T. et al. Systematic localization of common disease-associated variation in regulatory DNA. Science 337, 1190–1195 (2012).
Schaub, M. A., Boyle, A. P., Kundaje, A., Batzoglou, S. & Snyder, M. Linking disease associations with regulatory information in the human genome. Genome Res. 22, 1748–1759 (2012). References 12 and 13 reveal that most of the disease-associated variations are located in regulatory sequences.
Ingram, V. M. Sickle-cell anemia hemoglobin: the molecular biology of the first “molecular disease” — the crucial importance of serendipity. Genetics 167, 1–7 (2004).
Welter, D. et al. The NHGRI GWAS Catalog, a curated resource of SNP-trait associations. Nucleic Acids Res. 42, D1001–D1006 (2014).
Roadmap Epigenomics Consortium et al. Integrative analysis of 111 reference human epigenomes. Nature 518, 317–330 (2015).
Thurman, R. E. et al. The accessible chromatin landscape of the human genome. Nature 489, 75–82 (2012).
Shen, Y. et al. A map of the cis-regulatory sequences in the mouse genome. Nature 488, 116–120 (2012).
Abecasis, G. R. et al. An integrated map of genetic variation from 1,092 human genomes. Nature 491, 56–65 (2012).
Dekker, J., Rippe, K., Dekker, M. & Kleckner, N. Capturing chromosome conformation. Science 295, 1306–1311 (2002).
Fromm, M. & Berg, P. Simian virus 40 early- and late-region promoter functions are enhanced by the 72-base-pair repeat inserted at distant locations and inverted orientations. Mol. Cell. Biol. 3, 991–999 (1983).
Spitz, F. & Furlong, E. E. Transcription factors: from enhancer binding to developmental control. Nat. Rev. Genet. 13, 613–626 (2012).
Shlyueva, D., Stampfel, G. & Stark, A. Transcriptional enhancers: from properties to genome-wide predictions. Nat. Rev. Genet. 15, 272–286 (2014).
Heinz, S., Romanoski, C. E., Benner, C. & Glass, C. K. The selection and function of cell type-specific enhancers. Nat. Rev. Mol. Cell Biol. 16, 144–154 (2015).
Whyte, W. A. et al. Master transcription factors and mediator establish super-enhancers at key cell identity genes. Cell 153, 307–319 (2013).
Allen, B. L. & Taatjes, D. J. The Mediator complex: a central integrator of transcription. Nat. Rev. Mol. Cell Biol. 16, 155–166 (2015).
Grosveld, F., van Assendelft, G. B., Greaves, D. R. & Kollias, G. Position-independent, high-level expression of the human beta-globin gene in transgenic mice. Cell 51, 975–985 (1987).
Hnisz, D. et al. Super-enhancers in the control of cell identity and disease. Cell 155, 934–947 (2013).
Kellis, M. et al. Defining functional DNA elements in the human genome. Proc. Natl Acad. Sci. USA 111, 6131–6138 (2014).
Shin, H. Y. et al. Hierarchy within the mammary STAT5-driven Wap super-enhancer. Nat. Genet. 48, 904–911 (2016).
Hay, D. et al. Genetic dissection of the α-globin super-enhancer in vivo. Nat. Genet. 48, 895–903 (2016).
Hnisz, D. et al. Convergence of developmental and oncogenic signaling pathways at transcriptional super-enhancers. Mol. Cell 58, 362–370 (2015).
Bender, M. A. et al. Targeted deletion of 5′HS1 and 5′HS4 of the β-globin locus control region reveals additive activity of the DNaseI hypersensitive sites. Blood 98, 2022–2027 (2001).
Kieffer-Kwon, K. R. et al. Interactome maps of mouse gene regulatory domains reveal basic principles of transcriptional regulation. Cell 155, 1507–1520 (2013).
Lettice, L. A. et al. A long-range Shh enhancer regulates expression in the developing limb and fin and is associated with preaxial polydactyly. Hum. Mol. Genet. 12, 1725–1735 (2003).
Jeong, Y., El-Jaick, K., Roessler, E., Muenke, M. & Epstein, D. J. A functional screen for sonic hedgehog regulatory elements across a 1 Mb interval identifies long-range ventral forebrain enhancers. Development 133, 761–772 (2006).
Sagai, T. et al. A cluster of three long-range enhancers directs regional Shh expression in the epithelial linings. Development 136, 1665–1674 (2009).
Uslu, V. V. et al. Long-range enhancers regulating Myc expression are required for normal facial morphogenesis. Nat. Genet. 46, 753–758 (2014).
Marinic, M., Aktas, T., Ruf, S. & Spitz, F. An integrated holo-enhancer unit defines tissue and gene specificity of the Fgf8 regulatory landscape. Dev. Cell 24, 530–542 (2013).
Montavon, T. et al. A regulatory archipelago controls Hox genes transcription in digits. Cell 147, 1132–1145 (2011).
Andrey, G. et al. A switch between topological domains underlies HoxD genes collinearity in mouse limbs. Science 340, 1234167 (2013).
Khan, M., Vaes, E. & Mombaerts, P. Regulation of the probability of mouse odorant receptor gene choice. Cell 147, 907–921 (2011).
Tasic, B. et al. Promoter choice determines splice site selection in protocadherin α and γ pre-mRNA splicing. Mol. Cell 10, 21–33 (2002).
Lower, K. M. et al. Adventitious changes in long-range gene expression caused by polymorphic structural variation and promoter competition. Proc. Natl Acad. Sci. USA 106, 21771–21776 (2009).
Dillon, N., Trimborn, T., Strouboulis, J., Fraser, P. & Grosveld, F. The effect of distance on long-range chromatin interactions. Mol. Cell 1, 131–139 (1997).
De Gobbi, M. et al. A regulatory SNP causes a human genetic disease by creating a new transcriptional promoter. Science 312, 1215–1217 (2006).
Wijgerde, M., Grosveld, F. & Fraser, P. Transcription complex stability and chromatin dynamics in vivo. Nature 377, 209–213 (1995).
Zabidi, M. A. et al. Enhancer-core-promoter specificity separates developmental and housekeeping gene regulation. Nature 518, 556–559 (2015).
Arnold, C. D. et al. Genome-wide quantitative enhancer activity maps identified by STARR-seq. Science 339, 1074–1077 (2013).
Rada-Iglesias, A. et al. A unique chromatin signature uncovers early developmental enhancers in humans. Nature 470, 279–283 (2011).
Sagai, T., Hosoya, M., Mizushina, Y., Tamura, M. & Shiroishi, T. Elimination of a long-range cis-regulatory module causes complete loss of limb-specific Shh expression and truncation of the mouse limb. Development 132, 797–803 (2005).
Nobrega, M. A., Zhu, Y., Plajzer-Frick, I., Afzal, V. & Rubin, E. M. Megabase deletions of gene deserts result in viable mice. Nature 431, 988–993 (2004).
Sturtevant, A. H. The effects of unequal crossing over at the bar locus in Drosophila. Genetics 10, 117–147 (1925).
Ruf, S. et al. Large-scale analysis of the regulatory architecture of the mouse genome with a transposon-associated sensor. Nat. Genet. 43, 379–386 (2011). This study comprehensively illustrates the importance of genomic context for enhancer function.
Dekker, J., Marti-Renom, M. A. & Mirny, L. A. Exploring the three-dimensional organization of genomes: interpreting chromatin interaction data. Nat. Rev. Genet. 14, 390–403 (2013).
Denker, A. & de Laat, W. The second decade of 3C technologies: detailed insights into nuclear organization. Genes Dev. 30, 1357–1382 (2016).
Dixon, J. R. et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485, 376–380 (2012).
Nora, E. P. et al. Spatial partitioning of the regulatory landscape of the X-inactivation centre. Nature 485, 381–385 (2012). References 57 and 58 demonstrate the presence of TADs in the mammalian genome.
Rao, S. S. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014).
Bouwman, B. A. & de Laat, W. Getting the genome in shape: the formation of loops, domains and compartments. Genome Biol. 16, 154 (2015).
Ptashne, M. Gene regulation by proteins acting nearby and at a distance. Nature 322, 697–701 (1986).
Tolhuis, B., Palstra, R. J., Splinter, E., Grosveld, F. & de Laat, W. Looping and interaction between hypersensitive sites in the active β-globin locus. Mol. Cell 10, 1453–1465 (2002).
Dixon, J. R. et al. Chromatin architecture reorganization during stem cell differentiation. Nature 518, 331–336 (2015).
Deng, W. et al. Controlling long-range genomic interactions at a native locus by targeted tethering of a looping factor. Cell 149, 1233–1244 (2012). This study provides direct evidence that chromatin loops between enhancers and promoters underlie transcription.
Deng, W. et al. Reactivation of developmentally silenced globin genes by forced chromatin looping. Cell 158, 849–860 (2014).
Phillips-Cremins, J. E. et al. Architectural protein subclasses shape 3D organization of genomes during lineage commitment. Cell 153, 1281–1295 (2013).
Palstra, R. J. et al. The β-globin nuclear compartment in development and erythroid differentiation. Nat. Genet. 35, 190–194 (2003).
Krijger, P. H. et al. Cell-of-origin-specific 3D genome structure acquired during somatic cell reprogramming. Cell Stem Cell 18, 597–610 (2016).
Beagan, J. A. et al. Local genome topology can exhibit an incompletely rewired 3D-folding state during somatic cell reprogramming. Cell Stem Cell 18, 611–624 (2016).
Li, G. et al. Extensive promoter-centered chromatin interactions provide a topological basis for transcription regulation. Cell 148, 84–98 (2012).
Williamson, I., Lettice, L. A., Hill, R. E. & Bickmore, W. A. Shh and ZRS enhancer co-localisation is specific to the zone of polarizing activity. Development 143, 2994–3001 (2016).
de Laat, W. & Duboule, D. Topology of mammalian developmental enhancers and their regulatory landscapes. Nature 502, 499–506 (2013).
Bickmore, W. A. & van Steensel, B. Genome architecture: domain organization of interphase chromosomes. Cell 152, 1270–1284 (2013).
Pope, B. D. et al. Topologically associating domains are stable units of replication-timing regulation. Nature 515, 402–405 (2014).
Symmons, O. et al. Functional and topological characteristics of mammalian regulatory domains. Genome Res. 24, 390–400 (2014).
Waszak, S. M. et al. Population variation and genetic control of modular chromatin architecture in humans. Cell 162, 1039–1050 (2015).
Grubert, F. et al. Genetic control of chromatin states in humans involves local and distal chromosomal interactions. Cell 162, 1051–1065 (2015).
Denker, A. & de Laat, W. A. Long-distance chromatin affair. Cell 162, 942–943 (2015).
Heath, H. et al. CTCF regulates cell cycle progression of αβ T cells in the thymus. EMBO J. 27, 2839–2850 (2008).
Chen, H., Tian, Y., Shu, W., Bo, X. & Wang, S. Comprehensive identification and annotation of cell type-specific and ubiquitous CTCF-binding sites in the human genome. PLoS ONE 7, e41374 (2012).
Ong, C. T. & Corces, V. G. CTCF: an architectural protein bridging genome topology and function. Nat. Rev. Genetics 15, 234–246 (2014).
Splinter, E. et al. CTCF mediates long-range chromatin looping and local histone modification in the beta-globin locus. Genes Dev. 20, 2349–2354 (2006).
Handoko, L. et al. CTCF-mediated functional chromatin interactome in pluripotent cells. Nat. Genet. 43, 630–638 (2011).
Yusufzai, T. M., Tagami, H., Nakatani, Y. & Felsenfeld, G. CTCF tethers an insulator to subnuclear sites, suggesting shared insulator mechanisms across species. Mol. Cell 13, 291–298 (2004).
Hadjur, S. et al. Cohesins form chromosomal cis-interactions at the developmentally regulated IFNG locus. Nature 460, 410–413 (2009).
Kagey, M. H. et al. Mediator and cohesin connect gene expression and chromatin architecture. Nature 467, 430–435 (2010).
Parelho, V. et al. Cohesins functionally associate with CTCF on mammalian chromosome arms. Cell 132, 422–433 (2008).
Wendt, K. S. et al. Cohesin mediates transcriptional insulation by CCCTC-binding factor. Nature 451, 796–801 (2008).
Watrin, E., Kaiser, F. J. & Wendt, K. S. Gene regulation and chromatin organization: relevance of cohesin mutations to human disease. Curr. Opin. Genet. Dev. 37, 59–66 (2016).
Vietri Rudan, M. et al. Comparative Hi-C reveals that CTCF underlies evolution of chromosomal domain architecture. Cell Rep. 10, 1297–1309 (2015).
de Wit, E. et al. CTCF binding polarity determines chromatin looping. Mol. Cell 60, 676–684 (2015).
Guo, Y. et al. CRISPR inversion of CTCF sites alters genome topology and enhancer/promoter function. Cell 162, 900–910 (2015).
Sanborn, A. L. et al. Chromatin extrusion explains key features of loop and domain formation in wild-type and engineered genomes. Proc. Natl Acad. Sci. USA 112, E6456–E6465 (2015). References 59, 91–93 show that TADs are formed between convergent CTCF sites.
Gomez-Marin, C. et al. Evolutionary comparison reveals that diverging CTCF sites are signatures of ancestral topological associating domains borders. Proc. Natl Acad. Sci. USA 112, 7542–7547 (2015).
Alipour, E. & Marko, J. F. Self-organization of domain structures by DNA-loop-extruding enzymes. Nucleic Acids Res. 40, 11202–11212 (2012).
Fudenberg, G. et al. Formation of chromosomal domains by loop extrusion. Cell Rep. 15, 2038–2049 (2016).
Merkenschlager, M. & Nora, E. P. CTCF and cohesin in genome folding and transcriptional gene regulation. Annu. Rev. Genomics Hum. Genet. 31, 17–43 (2016).
Narendra, V. et al. CTCF establishes discrete functional chromatin domains at the Hox clusters during differentiation. Science 347, 1017–1021 (2015).
Dowen, J. M. et al. Control of cell identity genes occurs in insulated neighborhoods in mammalian chromosomes. Cell 159, 374–387 (2014).
Ji, X. et al. 3D chromosome regulatory landscape of human pluripotent cells. Cell Stem Cell 18, 262–275 (2016).
Gibcus, J. H. & Dekker, J. The hierarchy of the 3D genome. Mol. Cell 49, 773–782 (2013).
Kind, J. & van Steensel, B. Genome-nuclear lamina interactions and gene regulation. Curr. Opin. Cell Biol. 22, 320–325 (2010).
de Wit, E. et al. The pluripotent genome in three dimensions is shaped around pluripotency factors. Nature 501, 227–231 (2013).
Bantignies, F. et al. Polycomb-dependent regulatory contacts between distant Hox loci in Drosophila. Cell 144, 214–226 (2011).
Denholtz, M. et al. Long-range chromatin contacts in embryonic stem cells reveal a role for pluripotency factors and polycomb proteins in genome organization. Cell Stem Cell 13, 602–616 (2013).
Wijchers, P. J. et al. Cause and consequence of tethering a SubTAD to different nuclear compartments. Mol. Cell 61, 461–473 (2016).
Krijger, P. H. & de Laat, W. Identical cells with different 3D genomes; cause and consequences? Curr. Opin. Genet. Dev. 23, 191–196 (2013).
Nagano, T. et al. Single-cell Hi-C reveals cell-to-cell variability in chromosome structure. Nature 502, 59–64 (2013).
Dekker, J. & Mirny, L. The 3D genome as moderator of chromosomal communication. Cell 164, 1110–1121 (2016).
Therizols, P. et al. Chromatin decondensation is sufficient to alter nuclear organization in embryonic stem cells. Science 346, 1238–1242 (2014).
1000 Genomes Project Consortium et al. A global reference for human genetic variation. Nature 526, 68–74 (2015).
Tak, Y. G. & Farnham, P. J. Making sense of GWAS: using epigenomics and genome engineering to understand the functional relevance of SNPs in non-coding regions of the human genome. Epigenetics Chromatin 8, 57 (2015).
Kleinjan, D. J. & Coutinho, P. Cis-ruption mechanisms: disruption of cis-regulatory control as a cause of human genetic disease. Brief. Funct. Genomics Proteomic 8, 317–332 (2009).
Benko, S. et al. Highly conserved non-coding elements on either side of SOX9 associated with Pierre Robin sequence. Nat. Genet. 41, 359–364 (2009).
Fukami, M. et al. Complex genomic rearrangement in the SOX9 5′ region in a patient with Pierre Robin sequence and hypoplastic left scapula. Am. J. Med. Genet. A 158A, 1529–1534, (2012).
Taub, R. et al. Translocation of the c-myc gene into the immunoglobulin heavy chain locus in human Burkitt lymphoma and murine plasmacytoma cells. Proc. Natl Acad. Sci. USA 79, 7837–7841 (1982).
Dalla-Favera, R. et al. Human c-myc onc gene is located on the region of chromosome 8 that is translocated in Burkitt lymphoma cells. Proc. Natl Acad. Sci. USA 79, 7824–7827 (1982).
Northcott, P. A. et al. Enhancer hijacking activates GFI1 family oncogenes in medulloblastoma. Nature 511, 428–434 (2014).
Musunuru, K. et al. From noncoding variant to phenotype via SORT1 at the 1p13 cholesterol locus. Nature 466, 714–719 (2010).
Oldridge, D. A. et al. Genetic predisposition to neuroblastoma mediated by a LMO1 super-enhancer polymorphism. Nature 528, 418–421 (2015).
Mansour, M. R. et al. Oncogene regulation. An oncogenic super-enhancer formed through somatic mutation of a noncoding intergenic element. Science 346, 1373–1377 (2014).
Wieczorek, D. et al. A specific mutation in the distant sonic hedgehog (SHH) cis-regulator (ZRS) causes Werner mesomelic syndrome (WMS) while complete ZRS duplications underlie Haas type polysyndactyly and preaxial polydactyly (PPD) with or without triphalangeal thumb. Hum. Mutat. 31, 81–89 (2010).
Lettice, L. A. et al. Opposing functions of the ETS factor family define Shh spatial expression in limb buds and underlie polydactyly. Dev. Cell 22, 459–467 (2012).
Zhang, X. et al. Identification of focally amplified lineage-specific super-enhancers in human epithelial cancers. Nat. Genet. 48, 176–182 (2016).
van de Werken, H. J. et al. Robust 4C-seq data analysis to screen for regulatory DNA interactions. Nat. Methods 9, 969–972 (2012).
Gavrilov, A. A., Chetverina, H. V., Chermnykh, E. S., Razin, S. V. & Chetverin, A. B. Quantitative analysis of genomic element interactions by molecular colony technique. Nucleic Acids Res. 42, e36 (2014).
Jiang, T. et al. Identification of multi-loci hubs from 4C-seq demonstrates the functional importance of simultaneous interactions. Nucleic Acids Res. http://dx.doi.org/10.1093/nar/gkw568 (2016).
Fuss, S. H., Omura, M. & Mombaerts, P. Local and cis effects of the H element on expression of odorant receptor genes in mouse. Cell 130, 373–384 (2007).
Montavon, T., Thevenet, L. & Duboule, D. Impact of copy number variations (CNVs) on long-range gene regulation at the HoxD locus. Proc. Natl Acad. Sci. USA 109, 20204–20211 (2012).
Lupianez, D. G., Spielmann, M. & Mundlos, S. Breaking TADs: how alterations of chromatin domains result in disease. Trends Genet. 32, 225–237 (2016).
Kandoth, C. et al. Mutational landscape and significance across 12 major cancer types. Nature 502, 333–339 (2013).
Filippova, G. N. et al. A widely expressed transcription factor with multiple DNA sequence specificity, CTCF, is localized at chromosome segment 16q22.1 within one of the smallest regions of overlap for common deletions in breast and prostate cancers. Genes Chromosomes Cancer 22, 26–36 (1998).
Kon, A. et al. Recurrent mutations in multiple components of the cohesin complex in myeloid neoplasms. Nat. Genet. 45, 1232–1237 (2013).
Cancer Genome Atlas Research Network. Genomic and epigenomic landscapes of adult de novo acute myeloid leukemia. N. Engl. J. Med. 368, 2059–2074 (2013).
Kemp, C. J. et al. CTCF haploinsufficiency destabilizes DNA methylation and predisposes to cancer. Cell Rep. 7, 1020–1029 (2014).
Mullenders, J. et al. Cohesin loss alters adult hematopoietic stem cell homeostasis, leading to myeloproliferative neoplasms. J. Exp. Med. 212, 1833–1850 (2015).
Viny, A. D. et al. Dose-dependent role of the cohesin complex in normal and malignant hematopoiesis. J. Exp. Med. 212, 1819–1832 (2015).
Corces, M. R. & Corces, V. G. The three-dimensional cancer genome. Curr. Opin. Genet. Dev. 36, 1–7 (2016).
Katainen, R. et al. CTCF/cohesin-binding sites are frequently mutated in cancer. Nat. Genet. 47, 818–821 (2015).
Hnisz, D. et al. Activation of proto-oncogenes by disruption of chromosome neighborhoods. Science 351, 1454–1458 (2016).
Bell, A. C. & Felsenfeld, G. Methylation of a CTCF-dependent boundary controls imprinted expression of the Igf2 gene. Nature 405, 482–485 (2000).
Wang, H. et al. Widespread plasticity in CTCF occupancy linked to DNA methylation. Genome Res. 22, 1680–1688 (2012).
Maurano, M. T. et al. Large-scale identification of sequence variants influencing human transcription factor occupancy in vivo. Nat. Genet. 47, 1393–1401 (2015).
Flavahan, W. A. et al. Insulator dysfunction and oncogene activation in IDH mutant gliomas. Nature 529, 110–114 (2016).
Visser, M., Kayser, M. & Palstra, R. J. HERC2 rs12913832 modulates human pigmentation by attenuating chromatin-loop formation between a long-range enhancer and the OCA2 promoter. Genome Res. 22, 446–455 (2012).
Groschel, S. et al. A single oncogenic enhancer rearrangement causes concomitant EVI1 and GATA2 deregulation in leukemia. Cell 157, 369–381 (2014).
Lupianez, D. G. et al. Disruptions of topological chromatin domains cause pathogenic rewiring of gene-enhancer interactions. Cell 161, 1012–1025 (2015). References 146 and 147 illustrate how chromatin contact profiles help uncover the functional consequences of chromosomal rearrangements and in the discovery of molecular mechanisms underlying disease.
Fischer, J. et al. Inactivation of the Fto gene protects from obesity. Nature 458, 894–898 (2009).
Smemo, S. et al. Obesity-associated variants within FTO form long-range functional connections with IRX3. Nature 507, 371–375 (2014).
Claussnitzer, M. et al. FTO obesity variant circuitry and adipocyte browning in humans. N. Engl. J. Med. 373, 895–907 (2015). References 149 and 150 illustrate how the implementation of both epigenetic and 3D genome maps can greatly assist in the identification of the target gene of disease-associated variants.
de Vree, P. J. et al. Targeted sequencing by proximity ligation for comprehensive variant detection and local haplotyping. Nat. Biotechnol. 32, 1019–1025 (2014).
Selvaraj, S., Schmitt, A. D., Dixon, J. R. & Ren, B. Complete haplotype phasing of the MHC and KIR loci with targeted HaploSeq. BMC Genomics 16, 900 (2015).
Splinter, E. et al. The inactive X chromosome adopts a unique three-dimensional conformation that is dependent on Xist RNA. Genes Dev. 25, 1371–1383 (2011).
Simonis, M. et al. Nuclear organization of active and inactive chromatin domains uncovered by chromosome conformation capture-on-chip (4C). Nat. Genet. 38, 1348–1354 (2006).
Hughes, J. R. et al. Analysis of hundreds of cis-regulatory landscapes at high resolution in a single, high-throughput experiment. Nat. Genet. 46, 205–212 (2014).
Davies, J. O. et al. Multiplexed analysis of chromosome conformation at vastly improved sensitivity. Nat. Methods 13, 74–80 (2016).
Mifsud, B. et al. Mapping long-range promoter contacts in human cells with high-resolution capture Hi-C. Nat. Genet. 47, 598–606 (2015).
Sahlen, P. et al. Genome-wide mapping of promoter-anchored interactions with close to single-enhancer resolution. Genome Biol. 16, 156 (2015).
Dostie, J. et al. Chromosome Conformation Capture Carbon Copy (5C): a massively parallel solution for mapping interactions between genomic elements. Genome Res. 16, 1299–1309 (2006).
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
Fullwood, M. J. et al. An oestrogen-receptor-α-bound human chromatin interactome. Nature 462, 58–64 (2009).
Li, M. J., Wang, L. Y., Xia, Z., Sham, P. C. & Wang, J. GWAS3D: detecting human regulatory variants by integrative analysis of genome-wide associations, chromosome interactions and histone modifications. Nucleic Acids Res. 41, W150–W158 (2013).
Acknowledgements
W.d.L's research is supported by a Netherlands Organisation for Scientific Research-Chemical Sciences (NWO/CW) TOP grant (714.012.002), an NWO-Innovational Research Incentives Scheme (VICI) grant (724.012.003), a European Union grant (2010–259743; Modelling Hepatocellular Carcinoma (MODHEP)) and a grant from The Leducq Foundation (FP058566-01-PR).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Glossary
- Thalassaemias
-
Inherited blood disorders characterized by the abnormal formation of haemoglobin and caused by the reduced production of globin proteins.
- Enhancers
-
Regulatory DNA elements that can upregulate the transcriptional output of a target gene.
- Topologically associated domains
-
(TADs). Boundary-insulated chromosomal segments within which sequences preferentially contact each other.
- Cofactor p300
-
An enhancer-associated protein with histone acetyltransferase activity that recruits transcriptional-activating proteins and can increase gene expression.
- STARR-seq
-
Self-transcribing active regulatory region sequencing. A massively parallel analysis plasmid based method to interrogate the sequence-intrinsic enhancer capacity of genomic segments.
- Housekeeping gene promoter
-
Promoter of a gene that is expressed in all tissues.
- Minimal promoter
-
The minimal portion of the promoter (the core promoter) required to initiate transcription. When integrated in the genome, it depends on enhancers for detectable levels of transcription.
- Enhancer trap assay
-
Assay in which transcriptional output of a genomic integrated reporter gene under the control of a minimal promoter is used to identify regulatory elements in a given genomic context.
- Structural variations
-
Changes in the linear order of multiple base pairs as a consequence of a chromosomal rearrangement (for example, translocation, inversion, deletion, insertion or duplication).
- Nuclear compartments
-
Nuclear sub-volumes in which chromosomal regions with similar chromatin composition spatially aggregate.
- Haplotype
-
A genomic region that is inherited from one parent that has a sequence composition with genetic variations such as single-nucleotide polymorphisms (SNPs) that can be different from those found in the corresponding genomic region inherited by the other parent.
- Chromocentres
-
Sites in the nuclear interior where heterochromatic pericentromeric regions of different chromosomes cluster.
- Polycomb
-
A family of proteins that remodel chromatin to repress transcription.
- SNPs
-
Single-nucleotide polymorphisms causing genetic variation.
- Indels
-
Short DNA sequence insertions or deletions.
- Copy number variations
-
Changes in the number of copies of a given chromosomal segment in a cell caused by their duplication or deletion.
- Drivers
-
Risk-associated variants that cause disease.
- Passengers
-
Risk-associated variants that do not cause disease.
- Linkage disequilibrium
-
Alleles of SNPs are in linkage disequilibrium when their frequency of association in the population is not random, which indicates that they are physically and/or functionally linked.
- Chromosomal breakpoints
-
The sites where chromosomes were incidentally broken and, as consequence of erroneous DNA repair, now flank a chromosomal rearrangement.
- Haploinsufficient
-
A phenotypic state observed despite a diploid organism still carrying one functional gene.
Rights and permissions
About this article
Cite this article
Krijger, P., de Laat, W. Regulation of disease-associated gene expression in the 3D genome. Nat Rev Mol Cell Biol 17, 771–782 (2016). https://doi.org/10.1038/nrm.2016.138
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrm.2016.138
This article is cited by
-
3D Enhancer–promoter networks provide predictive features for gene expression and coregulation in early embryonic lineages
Nature Structural & Molecular Biology (2024)
-
Hi-Tag: a simple and efficient method for identifying protein-mediated long-range chromatin interactions with low cell numbers
Science China Life Sciences (2024)
-
Epiphany: predicting Hi-C contact maps from 1D epigenomic signals
Genome Biology (2023)
-
Expression of mTOR in normal and pathological conditions
Molecular Cancer (2023)
-
Regulation of chromatin organization during animal regeneration
Cell Regeneration (2023)