Abstract
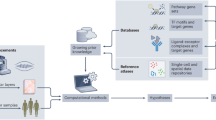
Understanding how the immune system is regulated and responds to pathogens will require whole-system approaches, because the study of single immunological parameters has, so far, been unable to unlock immune-system complexity. Global transcription analysis using microarray technologies provides a new approach to the description of complex biological phenomena. Here, we discuss insights into innate immunity that have been provided by genome-wide approaches and their impact on the interpretation of immune-system complexity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Service, R. F. Complex systems. Exploring the systems of life. Science 284, 80–83 (1999).
Goldenfeld, N. & Kadanoff, L. P. Simple lessons from complexity. Science 284, 87–89 (1999).
Nurse, P. Reductionism. The ends of understanding. Nature 387, 657 (1997).
Whitehead, A. N. Science and the Modern World (Macmillan, New York, 1925).
Keil, D., Luebke, R. W. & Pruett, S. B. Quantifying the relationship between multiple immunological parameters and host resistance: probing the limits of reductionism. J. Immunol. 167, 4543–4552 (2001).
Granucci, F. et al. Inducible IL-2 production by dendritic cells revealed by global gene-expression analysis. Nature Immunol. 2, 882–888 (2001).
Boldrick, J. C. et al. Stereotyped and specific gene-expression programs in human innate immune responses to bacteria. Proc. Natl Acad. Sci. USA 99, 972–977 (2002).
Lockhart, D. J. et al. Expression monitoring by hybridization to high-density oligonucleotide arrays. Nature Biotechnol. 14, 1675–1680 (1996).
Wodicka, L. et al. Genome-wide expression monitoring in Saccharomyces cerevisiae. Nature Biotechnol. 15, 1359–1367 (1997).
DeRisi, J. L., Iyer, V. R. & Brown, P. O. Exploring the metabolic and genetic control of gene expression on a genomic scale. Science 278, 680–686 (1997).
Zhao, N. et al. High-density cDNA filter analysis: a novel approach for large-scale, quantitative analysis of gene expression. Gene 156, 207–213 (1995).
Schena, M., Shalon, D., Davis, R. W. & Brown, P. O. Quantitative monitoring of gene-expression patterns with a complementary DNA microarray. Science 270, 467–470 (1995).
Iyer, V. R. et al. The transcriptional program in the response of human fibroblasts to serum. Science 283, 83–87 (1999).
Glynne, R. J., Ghandour, G. & Goodnow, C. C. Genomic-scale gene-expression analysis of lymphocyte growth, tolerance and malignancy. Curr. Opin. Immunol. 12, 210–214 (2000).
Rogge, L. et al. Transcript imaging of the development of human T helper cells using oligonucleotide arrays. Nature Genet. 25, 96–101 (2000).
Lee, C. K., Klopp, R. G., Weindruch, R. & Prolla, T. A. Gene-expression profile of aging and its retardation by caloric restriction. Science 285, 1390–1393 (1999).
Winzler, C. et al. Maturation stages of mouse dendritic cells in growth-factor-dependent long-term cultures. J. Exp. Med. 185, 317–328 (1997).
Kitano, H. Systems biology: a brief overview. Science 295, 1662–1664 (2002).
Ince, T. A. & Weinberg, R. A. Functional genomics and the breast-cancer problem. Cancer Cell 1, 15–17 (2002).
Dopazo, J. et al. Methods and approaches in the analysis of gene-expression data. J. Immunol. Methods 250, 92–112 (2001).
Staudt, L. M. & Brown, P. O. Genomic views of the immune system. Annu. Rev. Immunol. 18, 829–859 (2000).
Staudt, L. M. Gene-expression physiology and pathophysiology of the immune system. Trends Immunol. 22, 35–40 (2001).
Teague, T. K. et al. Activation changes the spectrum but not the diversity of genes expressed by T cells. Proc. Natl Acad. Sci. USA 96, 12691–12696 (1999).
Fahrer, A. M. et al. A genomic view of immunology. Nature 409, 836–838 (2001).
Manger, I. D. & Relman, D. A. How the host 'sees' pathogens: global gene-expression responses to infection. Curr. Opin. Immunol. 12, 215–218 (2000).
Janeway, C. A. Jr & Medzhitov, R. Innate immune recognition. Annu. Rev. Immunol. 20, 197–216 (2002).
Gordon, S., Clarke, S., Greaves, D. & Doyle, A. Molecular immunobiology of macrophages: recent progress. Curr. Opin. Immunol. 7, 24–33 (1995).
Morrissette, N., Gold, E. & Aderem, A. The macrophage — a cell for all seasons. Trends Cell. Biol. 9, 199–201 (1999).
Laskin, D. L., Weinberger, B. & Laskin, J. D. Functional heterogeneity in liver and lung macrophages. J. Leukocyte Biol. 70, 163–170 (2001).
Wang, Z. M., Liu, C. & Dziarski, R. Chemokines are the main proinflammatory mediators in human monocytes activated by Staphylococcus aureus, peptidoglycan and endotoxin. J. Biol. Chem. 275, 20260–20267 (2000).
Rosenberger, C. M. et al. Salmonella typhimurium infection and lipolysaccharide stimulation induce similar changes in macrophage gene expression. J. Immunol. 164, 5894–5904 (2000).
Ehrt, S. et al. Reprogramming of the macrophage transcriptome in response to interferon-γ and Mycobacterium tuberculosis: signaling role of nitric oxide synthase-2 and phagocyte oxidase. J. Exp. Med. 194, 1123–1139 (2001).
Nau, G. J. et al. Human macrophage activation programs induced by bacterial pathogens. Proc. Natl Acad. Sci. USA 99, 1503–1508 (2002).
Mellman, I. & Steinman, R. M. Dendritic cells: specialized and regulated antigen-processing machines. Cell 106, 255–258 (2001).
Banchereau, J. et al. Immunobiology of dendritic cells. Annu. Rev. Immunol. 18, 767–811 (2000).
Lanzavecchia, A. & Sallusto, F. The instructive role of dendritic cells on T-cell responses: lineages, plasticity and kinetics. Curr. Opin. Immunol. 13, 291–298 (2001).
Langenkamp, A., Messi, M., Lanzavecchia, A. & Sallusto, F. Kinetics of dendritic-cell activation: impact on priming of TH1, TH2 and nonpolarized T cells. Nature Immunol. 1, 311–316 (2000).
Sallusto, F. & Lanzavecchia, A. Efficient presentation of soluble antigen by cultured human dendritic cells is maintained by granulocyte–macrophage colony-stimulating factor plus interleukin-4 and downregulated by tumor-necrosis factor-α. J. Exp. Med. 179, 1109–1118 (1994).
Reid, C. D., Stackpoole, A., Meager, A. & Tikerpae, J. Interactions of tumor necrosis factor with granulocyte–macrophage colony-stimulating factor and other cytokines in the regulation of dendritic-cell growth in vitro from early bipotent CD34+ progenitors in human bone marrow. J. Immunol. 149, 2681–2688 (1992).
Huang, Q. et al. The plasticity of dendritic-cell responses to pathogens and their components. Science 294, 870–875 (2001).
d'Ostiani, C. F. et al. Dendritic cells discriminate between yeast and hyphae of the fungus Candida albicans. Implications for initiation of T helper cell immunity in vitro and in vivo. J. Exp. Med. 191, 1661–1674 (2000).
Borisy, G. G. & Svitkina, T. M. Actin machinery: pushing the envelope. Curr. Opin. Cell Biol. 12, 104–112 (2000).
Movilla, N. & Bustelo, X. N. Biological and regulatory properties of Vav-3, a new member of the Vav family of oncoprotein. Mol. Cell. Biol. 19, 7870–7885 (1999).
Rescigno, M. et al. Bacteria-induced neo-biosynthesis, stabilization and surface expression of functional class I molecules in mouse dendritic cells. Proc. Natl Acad. Sci. USA 95, 5229–5234 (1998).
Hashimoto, S. I. et al. Identification of genes specifically expressed in human activated and mature dendritic cells through serial analysis of gene expression. Blood 96, 2206–2214 (2000).
Rescigno, M. et al. Dendritic-cell survival and maturation are regulated by different signaling pathways. J. Exp. Med. 188, 2175–2180 (1998).
Granucci, F. et al. Transcriptional reprogramming of dendritic cells by differentiation stimuli. Eur. J. Immunol. 31, 2539–2546 (2001).
Andrews, D. M. et al. Infection of dendritic cells by murine cytomegalovirus induces functional paralysis. Nature Immunol. 2, 1077–1084 (2001).
Zitvogel, L. Dendritic and natural killer cells cooperate in the control/switch of innate immunity. J. Exp. Med. 195, F9–F14 (2002).
Csete, M. E. & Doyle, J. C. Reverse engineering of biological complexity. Science 295, 1664–1669 (2002).
Chong, L. & Ray, L. B. Whole-istic biology. Science 295, 1661 (2002).
Gallager, R. & Appenzeller, T. Beyond reductionism. Science 284, 79 (1999).
Dawkins, R. The Selfish Gene (Oxford University Press, New York, 1976).
Noble, D. Modeling the heart — from genes to cells to the whole organ. Science 295, 1678–1682 (2002).
Sorlie, T. et al. Gene-expression patterns of breast carcinomas distinguish tumor subclasses with clinical implications. Proc. Natl Acad. Sci. USA 98, 10869–10874 (2001).
Eisen, M. B., Spellman, P. T., Brown, P. O. & Botstein, D. Cluster analysis and display of genome-wide expression patterns. Proc. Natl Acad. Sci. USA 95, 14863–14868 (1998).
Chu, S. et al. The transcriptional program of sporulation in budding yeast. Science 282, 699–705 (1998).
Tamayo, P. et al. Interpreting patterns of gene expression with self-organizing maps: methods and application to hematopoietic differentiation. Proc. Natl Acad. Sci. USA 96, 2907–2912 (1999).
Butte, A. J. et al. Discovering functional relationships between RNA expression and chemotherapeutic susceptibility using relevance networks. Proc. Natl Acad. Sci. USA 97, 12182–12186 (2000).
Kao, C. M. Functional genomic technologies: creating new paradigms for fundamental and applied biology. Biotechnol. Prog. 15, 304–311 (1999).
Kell, D. B., King, R. D. On the optimization of classes for the assignment of unidentified reading frames in functional genomics programmes: the need for machine learning. Trends Biotechnol. 18, 93–98 (2000).
Shaffer, A. L. et al. Signatures of the immune response. Immunity 15, 375–385 (2001).
Glynne, R. et al. How self-tolerance and the immunosuppressive drug FK506 prevent B-cell mitogenesis. Nature 403, 672–676 (2000).
Crescenzi, M. & Giuliani, A. The main biological determinants of tumor-line taxonomy elucidated by a principal component analysis of microarray data. FEBS Lett. 507, 114–118 (2001).
Raychaudhuri, S., Stuart, J. M. & Altman, R. B. Principal-component analysis to summarize microarray experiments: application to sporulation time series. Pac. Symp. Biocomput. 455–466 (2000).
Acknowledgements
We thank N. Pavelka for the figures. This work was supported by grants from the Italian Association against Cancer (AIRC), the 5th EC Programs (DC strategies and TAGAPO) and MIUR (Ministero dell'Istruzione dell'Università e della Ricerca).
Author information
Authors and Affiliations
Corresponding author
Related links
Related links
DATABASES
Entrez
LocusLink
Rights and permissions
About this article
Cite this article
Ricciardi-Castagnoli, P., Granucci, F. Interpretation of the complexity of innate immune responses by functional genomics. Nat Rev Immunol 2, 881–888 (2002). https://doi.org/10.1038/nri936
Issue Date:
DOI: https://doi.org/10.1038/nri936
This article is cited by
-
Sensing the gut microbiota
Nature Immunology (2020)
-
The Microbiome and Graft Versus Host Disease
Current Stem Cell Reports (2015)
-
A systems-based approach to analyse the host response in murine lung macrophages challenged with respiratory syncytial virus
BMC Genomics (2013)
-
Lymphocytes in obesity-related adipose tissue inflammation
Diabetologia (2012)
-
Systems biology applied to vaccine and immunotherapy development
BMC Systems Biology (2011)