Key Points
-
The maternal microbiota itself affects the development of the fetus and the neonate.
-
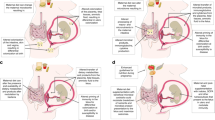
Metabolic exposure of the fetus in utero depends on maternal nutrition and xenobiotic exposure, both of which are modulated by microbial metabolism.
-
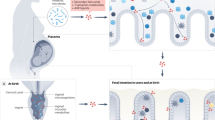
Molecules that originate from the intestinal microorganisms of a mother reach her offspring via the placenta during fetal development, and through maternal milk during the postnatal period.
-
Maternal antibodies amplify microbial molecular transfer, both in utero and during lactation.
-
Signalling from maternal microbial molecules shapes the development and function of the immune system in early life.
Abstract
The mucosal surfaces of mammals are densely colonized with microorganisms that are commonly referred to as the commensal microbiota. It is believed that the fetus in utero is sterile and that colonization with microorganisms starts only after birth. Nevertheless, the unborn fetus is exposed to a multitude of metabolites that originate from the commensal microbiota of the mother that reach systemic sites of the maternal body. The intestinal microbiota is strongly personalized and influenced by environmental factors, including nutrition. Members of the maternal microbiota can metabolize dietary components, pharmaceuticals and toxins, which can subsequently be passed to the developing fetus or the breast-feeding neonate. In this Review, we discuss the complex interplay between nutrition, the maternal microbiota and ingested chemicals, and summarize their effects on immunity in the offspring.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Blackburn, D. G. Reconstructing the evolution of viviparity and placentation. J. Theor. Biol. 192, 183–190 (1998).
Renfree, M. B., Hore, T. A., Shaw, G., Graves, J. A. & Pask, A. J. Evolution of genomic imprinting: insights from marsupials and monotremes. Annu. Rev. Genomics Hum. Genet. 10, 241–262 (2009).
Millen, J. W. & Woollam, D. H. Maternal nutrition in relation to abnormal foetal development. Proc. Nutr. Soc. 19, 1–5 (1960).
Pacha, J. Development of intestinal transport function in mammals. Physiol. Rev. 80, 1633–1667 (2000).
Torow, N. & Hornef, M. W. The neonatal window of opportunity: setting the stage for life-long host–microbial interaction and immune homeostasis. J. Immunol. 198, 557–563 (2017). This article elegantly summarizes literature on the importance of the time period soon after birth in shaping an individual's immune system, its indigenous commensal microbiota and its disease susceptibility later in life.
Buklijas, T. Food, growth and time: Elsie Widdowson's and Robert McCance's research into prenatal and early postnatal growth. Stud. Hist. Philos. Biol. Biomed. Sci. 47, 267–277 (2014).
Walton, A. & Hammond, J. The maternal effects on growth and conformation in Shire Horse–Shetland Pony crosses. Proc. R. Soc. B 125, 311–335 (1938).
Ross, M. G. & Beall, M. H. Adult sequelae of intrauterine growth restriction. Semin. Perinatol. 32, 213–218 (2008).
Fall, C. H. Evidence for the intra-uterine programming of adiposity in later life. Ann. Hum. Biol. 38, 410–428 (2011).
Ravelli, A. C. et al. Glucose tolerance in adults after prenatal exposure to famine. Lancet 351, 173–177 (1998).
Dennison, E. M. et al. Birthweight, vitamin D receptor genotype and the programming of osteoporosis. Paediatr. Perinat. Epidemiol. 15, 211–219 (2001).
Thompson, C., Syddall, H., Rodin, I., Osmond, C. & Barker, D. J. Birth weight and the risk of depressive disorder in late life. Br. J. Psychiatry 179, 450–455 (2001).
Bateson, P. et al. Developmental plasticity and human health. Nature 430, 419–421 (2004).
Ho, S. M., Tang, W. Y., Belmonte de Frausto, J. & Prins, G. S. Developmental exposure to estradiol and bisphenol A increases susceptibility to prostate carcinogenesis and epigenetically regulates phosphodiesterase type 4 variant 4. Cancer Res. 66, 5624–5632 (2006).
Zinkernagel, R. M. Maternal antibodies, childhood infections, and autoimmune diseases. N. Engl. J. Med. 345, 1331–1335 (2001).
Kearney, J. F., Patel, P., Stefanov, E. K. & King, R. G. Natural antibody repertoires: development and functional role in inhibiting allergic airway disease. Annu. Rev. Immunol. 33, 475–504 (2015).
Newbold, R. R. Lessons learned from perinatal exposure to diethylstilbestrol. Toxicol. Appl. Pharmacol. 199, 142–150 (2004).
Breton, J. et al. Gut microbiota limits heavy metals burden caused by chronic oral exposure. Toxicol. Lett. 222, 132–138 (2013).
Wilson, I. D. & Nicholson, J. K. Gut microbiome interactions with drug metabolism, efficacy, and toxicity. Transl Res. 179, 204–222 (2017).
Palmer, A. C. Nutritionally mediated programming of the developing immune system. Adv. Nutr. 2, 377–395 (2011).
Chen, X., Welner, R. S. & Kincade, P. W. A possible contribution of retinoids to regulation of fetal B lymphopoiesis. Eur. J. Immunol. 39, 2515–2524 (2009).
van de Pavert, S. A. et al. Maternal retinoids control type 3 innate lymphoid cells and set the offspring immunity. Nature 508, 123–127 (2014).
Fraker, P. J. & King, L. E. Reprogramming of the immune system during zinc deficiency. Annu. Rev. Nutr. 24, 277–298 (2004).
Benediktsson, R., Calder, A. A., Edwards, C. R. & Seckl, J. R. Placental 11β-hydroxysteroid dehydrogenase: a key regulator of fetal glucocorticoid exposure. Clin. Endocrinol. (Oxf.) 46, 161–166 (1997).
Backhed, F. et al. The gut microbiota as an environmental factor that regulates fat storage. Proc. Natl Acad. Sci. USA 101, 15718–15723 (2004). This is a landmark paper that reports the pervasive metabolic consequences of microbial metabolism and molecular exchange for the host.
Holmes, E., Li, J. V., Athanasiou, T., Ashrafian, H. & Nicholson, J. K. Understanding the role of gut microbiome–host metabolic signal disruption in health and disease. Trends Microbiol. 19, 349–359 (2011).
Mandal, S. et al. Fat and vitamin intakes during pregnancy have stronger relations with a pro-inflammatory maternal microbiota than does carbohydrate intake. Microbiome 4, 55 (2016).
Clark, A. & Mach, N. Role of vitamin D in the hygiene hypothesis: the interplay between vitamin D, vitamin D receptors, gut microbiota, and immune response. Front. Immunol. 7, 627 (2016).
Holmes, E., Li, J. V., Marchesi, J. R. & Nicholson, J. K. Gut microbiota composition and activity in relation to host metabolic phenotype and disease risk. Cell Metab. 16, 559–564 (2012).
Ma, J. et al. High-fat maternal diet during pregnancy persistently alters the offspring microbiome in a primate model. Nat. Commun. 5, 3889 (2014).
Roediger, W. E. Role of anaerobic bacteria in the metabolic welfare of the colonic mucosa in man. Gut 21, 793–798 (1980).
Furusawa, Y. et al. Commensal microbe-derived butyrate induces the differentiation of colonic regulatory T cells. Nature 504, 446–450 (2013).
Arpaia, N. et al. Metabolites produced by commensal bacteria promote peripheral regulatory T-cell generation. Nature 504, 451–455 (2013). References 31–33 show clear effects of microbial metabolites on host immune development and function.
Smith, P. M. et al. The microbial metabolites, short-chain fatty acids, regulate colonic Treg cell homeostasis. Science 341, 569–573 (2013).
LeBlanc, J. G. et al. Bacteria as vitamin suppliers to their host: a gut microbiota perspective. Curr. Opin. Biotechnol. 24, 160–168 (2013).
Schulman, S. G. Fundamentals of interaction of ionizing radiations with chemical, biochemical, and pharmaceutical systems. J. Pharm. Sci. 62, 1745–1757 (1973).
Albert, M. J., Mathan, V. I. & Baker, S. J. Vitamin B12 synthesis by human small intestinal bacteria. Nature 283, 781–782 (1980).
Camilo, E. et al. Folate synthesized by bacteria in the human upper small intestine is assimilated by the host. Gastroenterology 110, 991–998 (1996).
Collado, M. C., Isolauri, E., Laitinen, K. & Salminen, S. Distinct composition of gut microbiota during pregnancy in overweight and normal-weight women. Am. J. Clin. Nutr. 88, 894–899 (2008).
Koren, O. et al. Host remodeling of the gut microbiome and metabolic changes during pregnancy. Cell 150, 470–480 (2012).
Rossant, J. & Cross, J. C. Placental development: lessons from mouse mutants. Nat. Rev. Genet. 2, 538–548 (2001).
Sulik, K. K. Genesis of alcohol-induced craniofacial dysmorphism. Exp. Biol. Med. (Maywood) 230, 366–375 (2005).
Hilbert, J. M., Ning, J., Symchowicz, S. & Zampaglione, N. Placental transfer of quazepam in mice. Drug Metab. Dispos. 14, 310–312 (1986).
Coan, P. M. et al. Adaptations in placental nutrient transfer capacity to meet fetal growth demands depend on placental size in mice. J. Physiol. 586, 4567–4576 (2008).
Constancia, M. et al. Placental-specific IGF-II is a major modulator of placental and fetal growth. Nature 417, 945–948 (2002).
Li, L. et al. Regulation of maternal behavior and offspring growth by paternally expressed Peg3. Science 284, 330–333 (1999).
Heijmans, B. T. et al. Persistent epigenetic differences associated with prenatal exposure to famine in humans. Proc. Natl Acad. Sci. USA 105, 17046–17049 (2008).
Eggert, H., Kurtz, J. & Diddens-de Buhr, M. F. Different effects of paternal trans-generational immune priming on survival and immunity in step and genetic offspring. Proc. Biol. Sci. 281, 20142089 (2014).
Koch, M. A. et al. Maternal IgG and IgA antibodies dampen mucosal T helper cell responses in early life. Cell 165, 827–841 (2016). This study demonstrates the presence of maternal microbiota-specific IgG and IgA in the maternal milk, and shows that they are transferred to the offspring to establish functional mutualism with its incoming microbiota.
Grindstaff, J. L., Brodie, E. D. 3rd & Ketterson, E. D. Immune function across generations: integrating mechanism and evolutionary process in maternal antibody transmission. Proc. Biol. Sci. 270, 2309–2319 (2003).
Gautreaux, M. D., Deitch, E. A. & Berg, R. D. T lymphocytes in host defense against bacterial translocation from the gastrointestinal tract. Infect. Immun. 62, 2874–2884 (1994).
O'Boyle, C. J. et al. Microbiology of bacterial translocation in humans. Gut 42, 29–35 (1998).
Perez, P. F. et al. Bacterial imprinting of the neonatal immune system: lessons from maternal cells? Pediatrics 119, e724–e732 (2007).
Fardini, Y., Chung, P., Dumm, R., Joshi, N. & Han, Y. W. Transmission of diverse oral bacteria to murine placenta: evidence for the oral microbiome as a potential source of intrauterine infection. Infect. Immun. 78, 1789–1796 (2010).
Yajima, M. et al. Bacterial translocation in neonatal rats: the relation between intestinal flora, translocated bacteria, and influence of milk. J. Pediatr. Gastroenterol. Nutr. 33, 592–601 (2001).
Aagaard, K. et al. The placenta harbors a unique microbiome. Sci. Transl Med. 6, 237ra65 (2014). This study demonstrates the presence of non-pathogenic commensal bacteria in human placental samples.
Goldenberg, R. L., Hauth, J. C. & Andrews, W. W. Intrauterine infection and preterm delivery. N. Engl. J. Med. 342, 1500–1507 (2000).
Smith, K., McCoy, K. D. & Macpherson, A. J. Use of axenic animals in studying the adaptation of mammals to their commensal intestinal microbiota. Semin. Immunol. 19, 59–69 (2007).
Hapfelmeier, S. et al. Reversible microbial colonization of germ-free mice reveals the dynamics of IgA immune responses. Science 328, 1705–1709 (2010).
Gomez de Aguero, M. et al. The maternal microbiota drives early postnatal innate immune development. Science 351, 1296–1302 (2016). This paper describes the mechanism by which maternal gestational colonization shapes the composition and function of the early life immune system.
Jacobowitz Israel, E., Patel, V. K., Taylor, S. F., Marshak-Rothstein, A. & Simister, N. E. Requirement for a β2-microglobulin-associated Fc receptor for acquisition of maternal IgG by fetal and neonatal mice. J. Immunol. 154, 6246–6251 (1995).
Malek, A., Sager, R., Kuhn, P., Nicolaides, K. H. & Schneider, H. Evolution of maternofetal transport of immunoglobulins during human pregnancy. Am. J. Reprod. Immunol. 36, 248–255 (1996).
Neu, J. Gastrointestinal development and meeting the nutritional needs of premature infants. Am. J. Clin. Nutr. 85, 629S–634S (2007).
Siggers, R. H., Siggers, J., Thymann, T., Boye, M. & Sangild, P. T. Nutritional modulation of the gut microbiota and immune system in preterm neonates susceptible to necrotizing enterocolitis. J. Nutr. Biochem. 22, 511–521 (2011).
Kramer, D. R. & Cebra, J. J. Early appearance of “natural” mucosal IgA responses and germinal centers in suckling mice developing in the absence of maternal antibodies. J. Immunol. 154, 2051–2062 (1995).
Harris, N. L. et al. Mechanisms of neonatal mucosal antibody protection. J. Immunol. 177, 6256–6262 (2006).
Rogier, E. W. et al. Secretory antibodies in breast milk promote long-term intestinal homeostasis by regulating the gut microbiota and host gene expression. Proc. Natl Acad. Sci. USA 111, 3074–3079 (2014). This study provides clear evidence that maternal antibodies in breast milk have a protective role in the offspring.
Kau, A. L. et al. Functional characterization of IgA-targeted bacterial taxa from undernourished Malawian children that produce diet-dependent enteropathy. Sci. Transl Med. 7, 276ra24 (2015). This is a key paper showing the interactions between human infant microbiotas and their immune systems that determine environmental intestinal enteropathy, which is a severe and generally underappreciated problem in the developing world that affects the long-term development and health of affected individuals.
Blanton, L. V. et al. Gut bacteria that prevent growth impairments transmitted by microbiota from malnourished children. Science 351, aad3311 (2016).
Waterland, R. A. & Jirtle, R. L. Transposable elements: targets for early nutritional effects on epigenetic gene regulation. Mol. Cell. Biol. 23, 5293–5300 (2003).
Burris, H. H. et al. Offspring DNA methylation of the aryl-hydrocarbon receptor repressor gene is associated with maternal BMI, gestational age, and birth weight. Epigenetics 10, 913–921 (2015).
McGill, J. et al. Fetal exposure to ethanol has long-term effects on the severity of influenza virus infections. J. Immunol. 182, 7803–7808 (2009).
Lee, G. R., Kim, S. T., Spilianakis, C. G., Fields, P. E. & Flavell, R. A. T helper cell differentiation: regulation by cis elements and epigenetics. Immunity 24, 369–379 (2006).
Schuyler, R. P. et al. Distinct trends of DNA methylation patterning in the innate and adaptive immune systems. Cell Rep. 17, 2101–2111 (2016).
Lipka, D. B. et al. Identification of DNA methylation changes at cis-regulatory elements during early steps of HSC differentiation using tagmentation-based whole genome bisulfite sequencing. Cell Cycle 13, 3476–3487 (2014).
Cabezas-Wallscheid, N. et al. Identification of regulatory networks in HSCs and their immediate progeny via integrated proteome, transcriptome, and DNA methylome analysis. Cell Stem Cell 15, 507–522 (2014).
Moles, L. et al. Bacterial diversity in meconium of preterm neonates and evolution of their fecal microbiota during the first month of life. PLoS ONE 8, e66986 (2013).
Arrieta, M. C., Stiemsma, L. T., Amenyogbe, N., Brown, E. M. & Finlay, B. The intestinal microbiome in early life: health and disease. Front. Immunol. 5, 427 (2014).
Backhed, F. et al. Dynamics and stabilization of the human gut microbiome during the first year of life. Cell Host Microbe 17, 690–703 (2015).
Acknowledgements
The authors thank colleagues in their laboratory and U. Sauer, W. Hardt and C. Mueller for helpful discussions on the topic. The work of A.J.M. is funded by grants from the Swiss National Science Foundation (SNSF 310030B_160262 and SNSF Sinergia CRSII3_154414) and the Swiss SystemsX programme (GutX)); M.G.A. holds an Ambizione Grant of the Swiss National Science Foundation (PZ00P3_168012); and S.C.G.-V. is supported by a European Molecular Biology Organization Long-Term Fellowship (ALTF 841–2013).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Glossary
- Thalidomide
-
A drug that was prescribed primarily as a sedative or hypnotic. It was used to treat nausea and to cure morning sickness in pregnant women until it was discovered that it caused an absence of limbs in the offspring at birth.
- Hypothalamic–pituitary–adrenal axis
-
(HPA axis). One of the major neuroendocrine systems that controls, among other things, reactions to stress, the immune system, digestion and emotions. It consists of a complex set of feedforward and feedback mechanisms between the hypothalamus, the pituitary gland and the adrenal cortex. Neuroendocrine neurons in the hypothalamus produce corticotropin-releasing factor, which acts on the anterior pituitary gland to induce the production of adrenocorticotropic hormone (ACTH). ACTH induces the adrenal gland to release glucocorticoids, such as cortisol.
- Coprophagia
-
The eating of faeces, which is a normal behaviour in many animals.
- Haemochorial placenta
-
A type of placenta present in humans and some rodents, in which maternal blood is in direct contact with the chorion.
- Allantois
-
A bag-like structure that forms part of the developing conceptus and that has a role in nutrition and excretion.
Rights and permissions
About this article
Cite this article
Macpherson, A., de Agüero, M. & Ganal-Vonarburg, S. How nutrition and the maternal microbiota shape the neonatal immune system. Nat Rev Immunol 17, 508–517 (2017). https://doi.org/10.1038/nri.2017.58
Published:
Issue Date:
DOI: https://doi.org/10.1038/nri.2017.58
This article is cited by
-
Determinants of microbial colonization in the premature gut
Molecular Medicine (2023)
-
Konjac flour-mediated gut microbiota alleviates insulin resistance and improves placental angiogenesis of obese sows
AMB Express (2023)
-
Maternal effects drive intestinal development beginning in the embryonic period on the basis of maternal immune and microbial transfer in chickens
Microbiome (2023)
-
Early-life prophylactic antibiotic treatment disturbs the stability of the gut microbiota and increases susceptibility to H9N2 AIV in chicks
Microbiome (2023)
-
Dietary supplementation of laminarin improves the reproductive performance of sows and the growth of suckling piglets
Journal of Animal Science and Biotechnology (2023)