Key Points
-
Codon usage varies widely between species, between genes in a genome and between sites in a gene.
-
Explanations for natural variation in codon usage fall into two categories: mutational and selective.
-
Mutational mechanisms are responsible for most codon-usage variation between species; by contrast, selection for translation efficiency accounts for much of the systematic variation across a genome (except in mammals).
-
Translationally efficient codons may increase elongation rate, accuracy or both.
-
Rapid elongation should not be expected to influence protein yield per mRNA molecule for an endogenous gene, but it may be relevant for an overexpressed transgene.
-
The codons that provide efficient translation of an overexpressed transgene may differ from the efficient codons for an endogenous gene.
-
High-throughput measurements of endogenous mRNA levels, protein levels and ribosomal occupancies provide a detailed description of translation processes.
-
Libraries of randomized genes can elucidate design principles for efficient transgene expression, even without uncovering underlying mechanisms.
Abstract
Despite their name, synonymous mutations have significant consequences for cellular processes in all taxa. As a result, an understanding of codon bias is central to fields as diverse as molecular evolution and biotechnology. Although recent advances in sequencing and synthetic biology have helped to resolve longstanding questions about codon bias, they have also uncovered striking patterns that suggest new hypotheses about protein synthesis. Ongoing work to quantify the dynamics of initiation and elongation is as important for understanding natural synonymous variation as it is for designing transgenes in applied contexts.
Similar content being viewed by others
Main
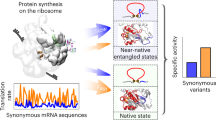
When the inherent redundancy of the genetic code was discovered, scientists were rightly puzzled by the role of synonymous mutations1. The central dogma of molecular biology suggests that synonymous mutations — those that do not alter the encoded amino acid — will have no effect on the resulting protein sequence and, therefore, no effect on cellular function, organismal fitness or evolution. Nonetheless, in most sequenced genomes, synonymous codons are not used in equal frequencies. This phenomenon, termed codon-usage bias (Fig. 1), is now recognized as crucial in shaping gene expression and cellular function through its effects on diverse processes, ranging from RNA processing to protein translation and protein folding. Naturally occurring codon biases are pervasive, and they can be extremely strong. Some species such as Thermus thermophilus avoid certain codons almost entirely. Synonymous mutations are important in applied settings as well — the use of particular codons can increase the expression of a transgene by more than 1,000-fold2.
The relative synonymous codon usage (RSCU)127 is plotted for 50 randomly selected genes from each of nine species. RSCU ranges from 0 (when the codon is absent), through 1 (when there is no bias) to 6 (when a single codon is used in a six-codon family). Methionine, tryptophan and stop codons are omitted. Genes are in rows and codons are in columns, with C- and G-ending codons on the left side of each panel. Note the extensive heterogeneity of codon usage among human genes. Other measures of a gene's codon bias include the codon adaptation index (CAI; the similarity of codon usage to a reference set of highly expressed genes)35, the frequency of 'optimal' codons (FOP)28 and the tRNA adaptation index (tAI; the similarity of codon usage to the relative copy numbers of tRNA genes)128.
We already enjoy a broad array of often conflicting hypotheses for the mechanisms that induce codon-usage biases in nature, and for their effects on protein synthesis and cellular fitness. Until recently, we have been unable to systematically interrogate these hypotheses through large-scale experimentation. As a result, despite decades of interest and substantial progress in understanding codon-usage biases, there is an overabundance of plausible explanatory models whose relative, quantitative contributions are seldom compared.
Advances in synthetic biology, mass spectrometry and sequencing now provide tools for systematically elucidating the molecular and cellular consequences of synonymous nucleotide variation. Such studies have refined our understanding of the relative roles of initiation, elongation, degradation and misfolding in determining protein expression levels of individual genes and the overall fitness of a cell. This information, in turn, is helping researchers to distinguish among the forces that shape naturally occurring patterns of codon usage. Researchers can also leverage high-throughput studies in applied settings that require controlled, heterologous gene expression, for example, to improve design principles for vaccine development and gene therapy.
Here we review the causes, consequences, and practical use of codon-usage biases. Because we already benefit from several outstanding reviews on naturally occurring codon biases3,4,5,6,7, we focus here on those classical hypotheses that remain unresolved and the recent developments arising from high-throughput studies. We begin by summarizing the empirical patterns of codon usage that are observed across species, across genomes and across individual genes. We describe the diverse array of mechanistic hypotheses for the causes of such variation and the sequence signatures that support them. Against this backdrop of hypotheses and sequence analysis, we describe experimental work that relates codon usage to endogenous gene expression and cellular fitness. From this, we turn to experimental studies on heterologous gene expression and their implications both for understanding natural synonymous variation and for engineering new constructs in applied settings.
Mechanistic hypotheses
Significant deviations from uniform codon choice have been observed in species from all taxa, including bacteria, archaea, yeast, fruitflies, worms and mammals. The overall codon usage in a genome can differ dramatically between species, although seldom between closely related species6.
Mutational versus selective hypotheses. Explanations for patterns of codon usage, within or between species, fall into two distinct categories that are associated with two independent forces in molecular evolution: mutation and natural selection3,4,5.
A mutational explanation posits that codon bias arises from the properties of underlying mutational processes — for example, biases in nucleotides that are produced by point mutations8, contextual biases in the point mutation rates or biases in repair. Mutational explanations are neutral because they do not require any fitness advantage or detriment to be associated with alternative synonymous codons. Mutational mechanisms are typically invoked to explain interspecific variation in codon usage, especially among unicellular organisms.
Explanations involving natural selection posit that synonymous mutations somehow influence the fitness of an organism, and they can therefore be promoted or repressed throughout evolution. Selective mechanisms are typically invoked to explain variation in codon usage across a genome or across a gene, although some interspecific variation is also attributable to such mechanisms (see below).
Selective and neutral explanations for codon usage are not mutually exclusive, and both types of mechanisms surely have a role in patterning synonymous variation within and between genomes3,5,9. Below we discuss the patterns of codon usage that have been documented at various levels of biological organization in light of their mutational or selective causes.
Patterns of codon usage
Patterns across species. The strongest single determinant of codon-usage variation across species is genomic GC content. In fact, differences in codon usage between bacterial species can be accurately predicted from the nucleotide content in their non-coding regions3,10. Genomic GC content is itself typically determined by mutational processes that act across the whole genome. As a result, most interspecific variation in codon usage is attributed to mutational mechanisms3,10, although the molecular causes of mutation biases are largely unknown10. Contrary to early expectations, the GC content of bacterial genomes or protein-coding genes is not correlated with optimal growth temperature (although, interestingly, structural RNAs show such a correlation)11.
In those species for which the point mutation rate depends strongly on the sequence context of a nucleotide — for example, in mammals, which experience hypermutable CpG dinucleotides — the mutational model predicts a strong context dependence of codon usage, which has indeed been observed12. Thus, at the genomic scale, neutral processes that do not discriminate among synonymous mutations remain plausible for explaining interspecific variation in codon usage among higher eukaryotes and they are well accepted as the primary determinants of interspecific variation in most other taxa (but see Ref. 13).
Aside from mutation biases, adaptation of codon usage to cellular tRNA abundances can also influence synonymous sequence variation across species (see below), as codon usage and tRNA regulation can co-evolve. Finally, some neutral processes that are responsible for codon bias across taxa are not mutational per se. Even in the absence of selection at synonymous sites, selection at non-synonymous sites can induce differences in nucleotide composition between coding and non-coding regions5,14,15,16.
Patterns across a genome. There is often systematic variation in codon usage among the genes in a genome, usually attributed to selection. In organisms, including Escherichia coli, Saccharomyces cerevisiae, Caenorhabditis elegans, Arabidopsis thaliana, Drosophila melanogaster and possibly also mammals (see below), there is a significant positive correlation between a gene's expression level and the degree of its codon bias, and a negative correlation between expression level and the rate of synonymous substitutions between divergent species9,17,18 — features that are difficult to explain through mutation alone. Although mutational effects could possibly co-vary with expression levels, because transcription can be mutagenic19,20, this effect is unlikely to account for the correlations between codon usage and expression levels that are observed in numerous species5,19,21.
The classic explanation for systematic variation across a genome is selectionist: codon bias is more extreme in highly expressed genes to match a skew in Iso-accepting tRNAs and, thereby, provide a fitness advantage through increased translation efficiency or accuracy of protein synthesis9,17,22,23,24,25,26,27. There is strong evidence for this hypothesis in several species, mostly in the form of broad correspondences between the 'preferred codons' that are used in highly expressed genes and measures of relative tRNA abundances28,29,30,31,32. As a result, translational selection remains the dominant explanation for systematic variation in codon usage among genes, despite the fact that supporting evidence is sometimes incomplete: direct measurements of tRNA abundances are rare in higher eukaryotes; the correspondence of tRNA abundance with tRNA copy number5 is weak in D. melanogaster and humans33; and 30% of bacterial species show no evidence of translational selection34.
There are two possible directions of causality relating an endogenous gene's expression level and the degree of its codon adaptation35 to tRNA abundances. In one view2,36,37 high codon adaptation induces strong protein expression, because rapid and/or accurate elongation increases a given protein's rate of synthesis; in the other view, strong expression selects for high codon adaptation to avoid costs that scale with a gene's expression level. In the biotechnology literature, the former interpretation is de rigueur, whereas in the literature on molecular evolution, the latter interpretation prevails3,4,5. The idea that high codon adaptation induces high protein levels per mRNA molecule does not square well with the notion that initiation is generally rate limiting for endogenous protein production7,9,38,39 (although it may apply to heterologous genes (see below)). When initiation is limiting, the elongation rate should not influence the amount of protein that is produced from a given message7,9 (Fig. 2). Moreover, from an evolutionary perspective, if high protein levels are desirable, it would seem easier to tune a promoter for increased transcription than to select on hundreds of individual synonymous mutations, each of which has only a marginal effect on the overall amount of protein synthesis. Conversely, the use of poorly adapted codons to slow the translation of genes expressed at low levels36 would seem wasteful compared to simply reducing transcription or slowing initiation.
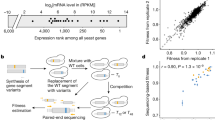
The steady-state rate of protein synthesis and density of ribosomes bound on an mRNA both depend on the rates of initiation and elongation. When elongation is the rate-limiting step in a gene's translation (case A), the message will be covered as densely as possible by ribosomes and faster elongation will tend to increase the rate of protein synthesis. However, most endogenous genes are believed to be initiation limited (cases B, C and D), so that their transcripts are not completely covered by ribosomes. This is evidenced by extensive variability in ribosome densities across endogenous mRNAs67. For two initiation-limited genes with the same initiation rate, the mRNA with faster elongation (afforded by, say, higher codon adaptation to tRNA pools) will have a lower density of translating ribosomes (C versus B) but no greater rate of termination. Thus, when initiation is limiting, high codon adaptation should not be expected to increase the amount of protein that is produced per mRNA molecule (protein amounts are the same in B and C). A lower density of ribosomes can also occur when two initiation-limited genes have the same elongation rate, but one has a slower initiation rate (D versus C). In this case, the amount of protein that is produced will be lower for the mRNA that has the slower initiation rate (D). The extent to which variation in ribosome densities67 arises from variation in initiation versus elongation rates remains to be determined. In all cases shown here, as is true for most endogenous genes, the gene's mRNA does not account for a substantial proportion of total cellular mRNA, so that the rates of initiation and elongation do not substantially alter the pool of free ribosomes (compare with Fig. 4).
Although evolutionary studies generally agree that high expression selects for high codon adaptation in endogenous genes (as opposed to the converse), the precise nature of fitness gains associated with translationally adapted codons remains a topic of active debate (Box 1). Furthermore, even though translational efficiency is energetically beneficial to the cell, efficient translation generally increases the amount of cell-to-cell variation in expression levels40 and this noise is typically deleterious41. Although translational selection has received the most attention, systematic variation in codon usage across a genome can also be caused by neutral processes in certain species; these processes include horizontal gene transfer42, different nucleotide bias in leading and lagging strands of replication in bacteria43 and isochore structure in mammals (Box 2).
Patterns across a gene. Codon usage can vary dramatically even within a single gene. Synonymous mutations at specific sites may experience selection because they disrupt motifs that are recognized by transcriptional or by post-transcriptional regulatory mechanisms, for example, microRNAs. Sites that require ribosomal pausing for proper co-translational protein folding or ubiquitin modification44 may experience selection for poorly adapted codons45 or strong mRNA folding46. Codon choice that promotes proper nucleosome positioning is selectively advantageous in eukaryotes, especially in 5′ regions47. And, finally, in mammals, synonymous mutations near an intron–exon boundary can create spurious splice sites or disrupt splicing control elements4,48, causing disease4. This phenomenon helps to explain the reduced rate of synonymous substitutions and SNP density near splicing control elements49,50. Selection for proper splicing also extends to D. melanogaster, and sequence variation suggests that it is probably an even stronger force than translational selection in shaping codon usage near intron–exon boundaries51.
Although important, the mechanisms of intragenic codon-usage variation described above are typically restricted to specific taxa or special classes of sites. Recent studies have argued for three mechanisms that produce systematic variation in codon usage across the sites in a gene in a diverse range of species.
One of these mechanisms is selection against strong 5′ mRNA structure to facilitate translation initiation. mRNA structure near the 5′ end of a coding region is generally disadvantageous9 as it can inhibit ribosomal initiation52,53 (Fig. 3a). Eyre-Walker and Bulmer proposed selection against mRNA structure to explain a trend towards reduced codon adaptation in the 5′ region of E. coli genes and a corresponding reduced rate of synonymous substitutions across divergent species54. More recently, following similar observations in E. col i55, Gu et al. demonstrated a broad trend in all sequenced prokaryotes and eukaryotes towards reduced mRNA stability near the translation initiation sites of genes, especially for GC-rich genes56. This study relied on computational predictions of mRNA structure in short windows; combined with large-scale experimental studies (see below), this work suggests a systematic role for selection on mRNA structure in shaping codon usage in the first 30–60 nucleotides of genes.
a | Structure in the ribosome binding site (RBS) usually inhibits initiation. However, initiation can occur when the structured element is positioned between the Shine–Dalgarno sequence (SD) and the start codon (AUG)129, or 15 nucleotides downstream of the start codon130,131. b | Synonymous mutations in the region from nucleotide −4 to +37 of a GFP gene alter the predicted folding energies by up to 12 kcal mol−1. A 5′ mRNA folding energy of below −10 kcal mol−1 strongly inhibits GFP expression in Escherichia coli55. c | More than 40% of human genes have predicted 5′ folding energies below the −10 kcal mol−1 threshold and are therefore expected to express poorly in E. coli without modification. AU, arbitrary units. Part b is modified, with permission, from Ref. 55 © (2009) American Association for the Advancement of Science.
Tuller et al.57 recently described a second, systematic trend in the pattern of intragenic codon usage: a 'ramp' of poorly adapted codons in the first 90–150 nucleotides of genes, which had earlier been observed in bacteria, yeast and fruitflies58,59. This pattern has been preserved across divergent species even when tRNA pools (estimated from gene copy numbers) have changed57. A ramp of poorly adapted codons presumably slows elongation at the start of a gene, which may provide several physiological benefits. Slow 5′ elongation is predicted to reduce the frequency of ribosomal traffic jams towards the 3′ end57,60, thus reducing the cost of wasted ribosomes and of spontaneous or collision-induced abortions. Alternatively, a ramp of slow elongation may facilitate recruitment of chaperone proteins to the emergent peptide61. Other explanations, unrelated to elongation rate, are also plausible such as weaker selection for accurate translation near the start of a gene, where missense and nonsense errors would be less costly24,59. The earliest interpretation of unusual 5′ codon usage posited selection to increase the initiation rate9; interestingly, the 5′ region of poorly adapted codons identified by Tuller et al. overlaps significantly with the region in which synonymous codon choice systematically reduces mRNA stability54,55,56,58. It remains unclear which selective mechanisms are primarily responsible for the unusual and nearly universal pattern of 5′ codon usage. Multiple mechanisms may certainly operate in different genes; however, it is unclear why a single gene should experience selection both to increase its rate of ribosomal initiation9 and to reduce the subsequent rate of its early elongation57.
Cannarozzi et al.62 recently exposed a third, novel pattern of intragenic codon usage in eukaryotes: the re-use or autocorrelation of codons across a gene sequence, driven, they argue, to improve elongation efficiency through tRNA recycling. If a recently used tRNA molecule is bound to the ribosome, or if it diffuses slowly compared to ribosomal progression and re-acylation63, then it would be efficient to re-use the same tRNA molecule for subsequent incorporations of the same amino acid. This physical model predicts selection for using the same codon or, more generally, a codon that is read by the same tRNA species, at nearby sites in a gene that encode the same amino acid. Indeed, Cannarozzi et al. observed significant autocorrelation of codons across gene sequences in most eukaryotes, especially in genes that are rapidly upregulated in response to stress. Of course, autocorrelation would also be predicted if all sites in a gene independently experience pressure for biased codon usage, for example, to match the global pool of tRNAs. To control for overall codon usage, Cannarozzi et al. compared the degree of autocorrelation in actual gene sequences to gene sequences that had been reshuffled at random, finding more autocorrelation on average in the unshuffled genes, although only marginally so. More convincingly, they observed that autocorrelation is strongest for iso-accepting codons of rare tRNAs in highly expressed genes, which is predicted by the tRNA-recycling hypothesis but not by a selective pressure that applies at all sites independently.
Measurements of endogenous expression
Recent developments in mass spectrometry and fluorescence microscopy allow large-scale measurements of endogenous protein levels64,65,66. Together with techniques for quantifying ribosomal occupancy67 and measuring elongation dynamics68, these advances provide a spectacularly detailed account of basic cellular processes, with implications for our understanding of codon biases.
Variation in protein/mRNA ratios. Shotgun proteomics have revealed an extensive role for post-transcriptional processes in determining eventual protein levels in bacteria, yeast69, worms70, fruitflies70 and especially mammals65,66. Whereas the imperfect correlations between protein and mRNA levels (R2 ≈ 47–77% in E. col i65,71, 73% in yeast65 and 29% in humans66) may previously have been seen as measurement noise, researchers have since attributed much of the variation in protein/mRNA ratios to sequence-derived characteristics of genes. In a recent study in human cells66, the strongest correlates of steady-state protein levels, controlling for mRNA levels, were coding-sequence length (reflecting the fact that longer transcripts are less stable72 or slower to initiate73), amino acid content (reflecting the variable costs associated with synthesizing different amino acids, or variable rates of protein degradation) and predicted 5′ mRNA structure (reflecting lower initiation rates when the 5′ structure is strong). Importantly, the codon adaptation index35, which correlates strongly with mRNA levels in yeast65 and weakly in humans66, shows little or no significant correlation with the amount of protein per mRNA molecule in either organism65,66; this suggests that codon adaptation does not significantly increase the protein yield from a given message, at least among endogenous genes74,75. It is important to note that steady-state protein levels are influenced by both protein production and protein degradation, so any variation in degradation rates unrelated to codon usage will further reduce the correlation between codon usage and protein/mRNA ratios.
Ribosomal footprints. Ingolia et al. recently devised a clever application of RNA–seq to quantify ribosome-protected RNA fragments in a cell, thereby estimating 'ribosomal footprints' across the transcriptome67. This method has provided rich information about translational regulation and it has uncovered some startling phenomena, such as an abundance of upstream ORFs with non-AUG start codons. The footprint data in yeast show a greater mean density of ribosomes in the first 100–150 codons of genes, suggesting locally slow elongation; this is consistent with the observed presence of poorly adapted codons in the 5′ region58,59. There is also a significant negative correlation, genome-wide, between a transcript's ribosome density and the experimentally measured strength of mRNA structure near its start site76 suggesting that strong 5′ mRNA structure retards translational initiation and reduces the density of translating ribosomes.
Remarkably, on averaging data from all yeast genes, Tuller et al.37 also observed a negative correlation between predicted mRNA folding energy and ribosome density among the first 65 codons, suggesting that strong mRNA structure downstream of the start site retards translational elongation. This observation is surprising, given the helicase activity of translating ribosomes77; however, the correlation between the genome-wide average profiles of mRNA folding and ribosome density does not imply a correlation at the level of individual sites. Ingolia et al. also measured ribosomal footprints under amino acid starvation and found that one-third of yeast genes showed substantially increased or decreased translational efficiency in these conditions compared with controls67. A detailed parsing of the relationship between a gene's amino acid content and translational response to starvation may improve design principles for overexpressed heterologous genes, which often induce starvation78,79 (see below).
Translational efficiency. Notions of translational efficiency differ in the literature on gene expression. Ingolia et al.67 defined the translational efficiency of a gene as the number of bound ribosomes per mRNA molecule; by contrast, Tuller et al.37,57 and others defined efficiency as protein yield per mRNA molecule (that is, the ratio of protein abundance to mRNA abundance). The second definition is more relevant to issues of total protein synthesis, whereas the former definition may be more relevant to ribosomal availability and overall cellular fitness. These two notions of translational efficiency are only weakly correlated for endogenous genes (R2 < 2.5% comparing the data by Ingolia et al.67 to Ref. 80), indicating that the density of ribosomes on a given mRNA molecule does not determine the amount of protein that is produced from it. Similarly, in yeast, a gene's codon adaptation index35 explains less than 3% of the variance in protein abundance per mRNA67. Both of these observations are consistent with the idea that, for most endogenous genes, the initiation is rate limiting for protein production38,39 and therefore determines the amount of protein produced from each message, regardless of ribosome density or codon adaptation7,9 (Fig. 2); however, this logic may not apply to overexpressed heterologous genes, which are described in the following section (Fig. 4).
Unlike most endogenous genes, mRNA from an overexpressed transgene may account for a substantial proportion of total cellular mRNA. In this case, slow elongation (caused by poor codon adaptation to charged tRNA pools, say) can increase the density of bound ribosomes and thereby reduce the pool of available ribosomes in the cell. Such a depletion of available ribosomes will feed back to reduce the initiation rate of subsequent translating ribosomes on the message, thereby reducing the rate of protein synthesis. This is illustrated schematically by comparing overexpressed mRNAs with slow elongation (above) and rapid elongation (below), but identical initiation sequences. Thus, the relationship between codon adaptation and the rate of protein synthesis per mRNA molecule may differ for an overexpressed transgene compared to an endogenous gene (Fig. 2).
Measurements of heterologous expression
Codon bias has a crucial role in heterologous gene expression. However, there is often a disconnection between technological and evolutionary studies of codon bias — a gap that partly reflects genuine differences between endogenous and heterologous situations. In many biotechnological applications, a transgene is massively overexpressed, accounting for up to 30% of the protein mass in cell. As a result, the principles that relate heterologous codon usage to protein levels may differ substantially from the endogenous case.
The idea that initiation generally limits translation may not apply to an overexpressed transgene whose mRNA accounts for a large proportion of total cellular mRNA. In such a case, inefficient use of ribosomes along the overexpressed mRNA may be sufficient to feed back and significantly deplete available ribosomes, thereby reducing initiation rates and retarding further heterologous protein production9 (Fig. 4). Thus, we might expect that the elongation effects of codon usage will influence protein yields per mRNA molecule for overexpressed genes. Nonetheless, we should not necessarily expect that the codons that are adapted to efficient elongation for endogenous genes will correspond to the efficient codons for heterologous genes, because overexpression causes amino acid starvation and concomitant alternations in the abundances of charged tRNAs78,79,81. Indeed, there was no significant correlation between codon adaptation35 and expression levels in two large-scale systematic experiments55,79. In fact, even endogenous genes that are essential during amino acid starvation such as amino acid biosynthetic enzymes preferentially use codons that are poorly adapted to the typical pool of charged tRNAs, but are well adapted to starvation-induced tRNA pools78,79.
Despite the complications described above, the field of codon optimization has traditionally focused on adjusting codon usage to match cellular tRNA abundances in standard conditions, disregarding other dimensions of bias. However, strategies are now changing. Several recent studies advocate for the role of global nucleotide content82,83, local mRNA folding55,84, codon pair bias85, a codon ramp57 or codon correlations62 in optimizing heterologous expression (Table 1).
Effects of codon adaptation on expression levels. Many studies show strong effects of rare codons on heterologous expression. In E. coli, stretches of rare AGA or AGG codons cause ribosome pausing and co-translational cleavage of mRNA86, ribosomal frameshifting87 or amino acid misincorporation88. Consistent with theoretical expectations, codons that are read by rare tRNAs can slow elongation by several fold89. And stretches of AGG codons near the ribosome-binding site (RBS) can reduce protein yields by obstructing translation initiation90. Although such studies are convincing, they usually address the effect of a subset of rare codons, often in long stretches, in E. coli cells; it is not known whether these principles can be applied in general.
Observations such as those above were quickly followed by efforts to adjust the global codon adaptation of transgenes to cellular tRNA abundances. Several approaches have been proposed: 'CAI maximization' replaces all codons by the most preferred codons in the target genome, but this could result in unbalanced charged tRNA pools2; 'codon harmonization'91 puts some non-preferred codons in positions that correspond to predicted protein domain boundaries; and 'codon sampling' adjusts the codon usage to reflect the overall usage in the target genome. In the absence of tRNA abundance estimates, codon frequencies in the target genome are sometimes used. It has also been suggested that codon usage should match the profile of charged tRNAs rather than total tRNAs79,81. The utility of codon adaptation approaches is still unclear, as they have not been systematically compared against each other, and several anecdotal studies argue both for (for example, Ref. 92) and against (for example, Ref. 93) their efficiency.
Codon adaptation algorithms typically optimize many sequence properties at once. This makes it difficult to determine which parameter causes observed differences in expression. In two recent multi-gene studies, between 60% and 70% of genes experienced increased expression upon codon optimization94,95 but whether this was a direct consequence of increased codon adaptation or other sequence properties is unclear. In our study of 154 synonymous variants of GFP, we observed no significant correlation between the codon adaption index35 and expression levels in E. col i55, but a weak positive correlation was later found using nonlinear regressions37,96. In any case, adaptation of codon usage is limited to species with pronounced and well-understood variation in tRNA concentrations, such as bacteria and yeast.
Effects of nucleotide bias on expression levels. Nucleotide biases are pervasive in natural genes and have the potential to alter the interactions of mRNA with DNA, with proteins and with itself, thereby influencing RNA production, degradation and translation rates. Many of these effects are characterized, but this knowledge has yet to find its way into standard codon optimization procedures.
GC-rich mRNAs can form strong secondary structures, and, in bacteria, strong structure near the RBS prohibits initiation53,55,97 (Fig. 3b). As a result, more than 40% of human genes would be expected to express poorly when placed in E. coli without modification (Fig. 3c). Strong structure near the start codon reduces heterologous expression in yeast as well (G.K., unpublished observations), consistent with evolutionary analyses56. No such effect has been described in mammals; on the contrary, high GC content generally increases expression levels in mammalian cells (see below). However, a strong mRNA hairpin in the coding sequence has been reported to interfere with translation in mammalian cells84 and strong hybrids between RNA and DNA (the R-loops) may interfere with transcription98.
GC-poor mRNAs are unlikely to fold strongly, but they often carry other sequence elements that limit expression. For example, low GC content is commonly believed to limit the expression of Plasmodium falciparum genes in E. coli, although the mechanisms are unknown. Such mRNAs may be targets for RNase E, which cleaves AU-rich sequences with low sequence specificity99. The situation is slightly clearer in mammals, in which low GC content (or high A content) has been shown to reduce expression82,83. This effect is common knowledge in virology, as HIV and human papilloma virus (HPV) genes are poorly expressed in human cells unless the gene sequences are optimized100,101. The rate-limiting step in these cases may be transcription or nuclear RNA export82,83, which is consistent with the efficient expression of GC-poor genes in cytoplasmic transcription systems based on the vaccinia virus101.
Little is known about the functional consequences of replication strand-related bias or CTAG avoidance, which are common in prokaryotes. High CpG content was reported to correlate with high expression in mammalian cells102, possibly by altering the distribution of nucleosomes on DNA.
Other effects of synonymous mutations on expression levels. Other examples of synonymous mutations influencing expression have been described as primarily anecdotal observations. In E. coli, overrepresented codon pairs103 were proposed to decrease translation elongation rates104, although this conclusion was later disputed105. In an attempt to produce attenuated strains, Coleman et al.85 partially de-optimized codon pairs in the poliovirus genome and observed a reduction in protein yield of several fold and a reduction in viral infectivity of 1,000-fold in mammalian cells. A version of GFP with autocorrelated codon usage showed 30% lower ribosome density in yeast, suggesting faster elongation, than a version with anticorrelated codon usage62. And a synonymous mutation in the human multidrug-resistance protein 1 (MDR1) gene was proposed to influence mRNA stability106 or protein folding and substrate specificity107. These observations are all intriguing and form important avenues for future systematic studies to determine their molecular bases.
Conclusions
Recent years have begun to see a convergence of experimental work on endogenous and heterologous gene expression, as both types of studies take advantage of high-throughput, quantitative techniques. Heterologous studies using large libraries of random or unbiased synonymous sequence variation55,81,97 are especially important for uncovering and comparing general rules to optimize expression. By contrast, relatively small-scale studies based on preconceived notions of 'optimized' codon usage do not provide sufficient power to distinguish among alternative mechanisms, nor do they allow us to discover any new mechanisms that increase expression. Heterologous studies will be complemented by endogenous measurements of initiation and elongation dynamics, and their effects on protein synthesis as a function of a gene's amino acid content and transcript level.
In the short term, there will be a trade-off between gaining predictive power for transgene optimization and deducing the underlying mechanisms that link codon usage and gene expression. High-dimensional, statistical regressions applied to large libraries of synonymous genes81,96 provide a principled, effective means of increasing heterologous expression. Such techniques are increasingly valuable in applied contexts in which high expression is required — such as viral-delivered gene therapies108,109 — but they do not generally identify molecular mechanisms. Our hope, over the long term, is that cross-fertilization between biotechnological and molecular biological studies will elucidate effective strategies for designing transgenes, as well as the mechanistic principles that underlie their expression.
References
Zuckerkandl, E. & Pauling, L. Molecules as documents of evolutionary history. J. Theor. Biol. 8, 357–366 (1965).
Gustafsson, C., Govindarajan, S. & Minshull, J. Codon bias and heterologous protein expression. Trends Biotechnol. 22, 346–353 (2004).
Hershberg, R. & Petrov, D. A. Selection on codon bias. Annu. Rev. Genet. 42, 287–299 (2008).
Chamary, J. V., Parmley, J. L. & Hurst, L. D. Hearing silence: non-neutral evolution at synonymous sites in mammals. Nature Rev. Genet. 7, 98–108 (2006). An excellent Review of the many surprisingly strong effects of synonymous mutations on splicing.
Duret, L. Evolution of synonymous codon usage in metazoans. Curr. Opin. Genet. Dev. 12, 640–649 (2002).
Sharp, P. M., Averof, M., Lloyd, A. T., Matassi, G. & Peden, J. F. DNA sequence evolution: the sounds of silence. Philos. Trans. R. Soc. Lond. B 349, 241–247 (1995).
Andersson, S. G. & Kurland, C. G. Codon preferences in free-living microorganisms. Microbiol. Rev. 54, 198–210 (1990). A classic, must-read paper that has framed the field of codon-usage adaptation.
Kimura, M. A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J. Mol. Evol. 16, 111–120 (1980).
Bulmer, M. The selection–mutation–drift theory of synonymous codon usage. Genetics 129, 897–907 (1991). A foundational study of both the population genetics that underlie codon bias and the biophysics of translation. This paper emphasizes that, for endogenous genes, elongation speed will not generally influence protein yield per mRNA.
Chen, S. L., Lee, W., Hottes, A. K., Shapiro, L. & McAdams, H. H. Codon usage between genomes is constrained by genome-wide mutational processes. Proc. Natl Acad. Sci. USA 101, 3480–3485 (2004).
Hurst, L. D. & Merchant, A. R. High guanine–cytosine content is not an adaptation to high temperature: a comparative analysis amongst prokaryotes. Proc. R. Soc. Lond. B 268, 493–497 (2001).
Fedorov, A., Saxonov, S. & Gilbert, W. Regularities of context-dependent codon bias in eukaryotic genes. Nucleic Acids Res. 30, 1192–1197 (2002).
Hildebrand, F., Meyer, A. & Eyre-Walker, A. Evidence of selection upon genomic GC-content in bacteria. PLoS Genet. 6, e1001107 (2010).
Morton, B. R. Selection at the amino acid level can influence synonymous codon usage: implications for the study of codon adaptation in plastid genes. Genetics 159, 347–358 (2001).
Cambray, G. & Mazel, D. Synonymous genes explore different evolutionary landscapes. PLoS Genet. 4, e1000256 (2008).
Plotkin, J. B. & Dushoff, J. Codon bias and frequency-dependent selection on the hemagglutinin epitopes of influenza A virus. Proc. Natl Acad. Sci. USA 100, 7152–7157 (2003).
Sharp, P. M. & Li, W. H. The rate of synonymous substitution in enterobacterial genes is inversely related to codon usage bias. Mol. Biol. Evol. 4, 222–230 (1987).
Eyre-Walker, A. & Bulmer, M. Synonymous substitution rates in enterobacteria. Genetics 140, 1407–1412 (1995).
Francino, M. P. & Ochman, H. Deamination as the basis of strand-asymmetric evolution in transcribed Escherichia coli sequences. Mol. Biol. Evol. 18, 1147–1150 (2001).
Majewski, J. Dependence of mutational asymmetry on gene-expression levels in the human genome. Am. J. Hum. Genet. 73, 688–692 (2003).
Duret, L. & Mouchiroud, D. Expression pattern and, surprisingly, gene length shape codon usage in Caenorhabditis, Drosophila, and Arabidopsis. Proc. Natl Acad. Sci. USA 96, 4482–4487 (1999).
Akashi, H. Synonymous codon usage in Drosophila melanogaster: natural selection and translational accuracy. Genetics 136, 927–935 (1994). By quantifying the rates of synonymous substitutions at conserved and non-conserved positions in proteins, Akashi shows that such mutations influence translational accuracy in D. melanogaster.
Akashi, H. & Schaeffer, S. W. Natural selection and the frequency distributions of “silent” DNA polymorphism in Drosophila. Genetics 146, 295–307 (1997).
Eyre-Walker, A. Synonymous codon bias is related to gene length in Escherichia coli: selection for translational accuracy? Mol. Biol. Evol. 13, 864–872 (1996).
Stoletzki, N. & Eyre-Walker, A. Synonymous codon usage in Escherichia coli: selection for translational accuracy. Mol. Biol. Evol. 24, 374–381 (2007).
Drummond, D. A. & Wilke, C. O. Mistranslation-induced protein misfolding as a dominant constraint on coding-sequence evolution. Cell 134, 341–352 (2008).
Zhou, T., Weems, M. & Wilke, C. O. Translationally optimal codons associate with structurally sensitive sites in proteins. Mol. Biol. Evol. 26, 1571–1580 (2009).
Ikemura, T. Correlation between the abundance of Escherichia coli transfer RNAs and the occurrence of the respective codons in its protein genes: a proposal for a synonymous codon choice that is optimal for the E. coli translational system. J. Mol. Biol. 151, 389–409 (1981).
Ikemura, T. Correlation between the abundance of yeast transfer RNAs and the occurrence of the respective codons in protein genes. Differences in synonymous codon choice patterns of yeast and Escherichia coli with reference to the abundance of isoaccepting transfer RNAs. J. Mol. Biol. 158, 573–597 (1982).
Post, L. E. & Nomura, M. Nucleotide sequence of the intercistronic region preceding the gene for RNA polymerase subunit α in Escherichia coli. J. Biol. Chem. 254, 10604–10606 (1979).
Moriyama, E. N. & Powell, J. R. Codon usage bias and tRNA abundance in Drosophila. J. Mol. Evol. 45, 514–523 (1997).
Duret, L. tRNA gene number and codon usage in the C. elegans genome are co-adapted for optimal translation of highly expressed genes. Trends Genet. 16, 287–289 (2000).
Kanaya, S., Yamada, Y., Kinouchi, M., Kudo, Y. & Ikemura, T. Codon usage and tRNA genes in eukaryotes: correlation of codon usage diversity with translation efficiency and with CG-dinucleotide usage as assessed by multivariate analysis. J. Mol. Evol. 53, 290–298 (2001).
Sharp, P. M., Bailes, E., Grocock, R. J., Peden, J. F. & Sockett, R. E. Variation in the strength of selected codon usage bias among bacteria. Nucleic Acids Res. 33, 1141–1153 (2005).
Sharp, P. M. & Li, W. H. The codon Adaptation Index — a measure of directional synonymous codon usage bias, and its potential applications. Nucleic Acids Res. 15, 1281–1295 (1987).
Lavner, Y. & Kotlar, D. Codon bias as a factor in regulating expression via translation rate in the human genome. Gene 345, 127–138 (2005).
Tuller, T., Waldman, Y. Y., Kupiec, M. & Ruppin, E. Translation efficiency is determined by both codon bias and folding energy. Proc. Natl Acad. Sci. USA 107, 3645–3650 (2010).
Bergmann, J. E. & Lodish, H. F. A kinetic model of protein synthesis. Application to hemoglobin synthesis and translational control. J. Biol. Chem. 254, 11927–11937 (1979).
Mathews, M. B., Sonenberg, N. & Hershey, J. W. B. (eds) Translational Control in Biology and Medicine (CHSL Press, New York, 2007).
Ozbudak, E. M., Thattai, M., Kurtser, I., Grossman, A. D. & van Oudenaarden, A. Regulation of noise in the expression of a single gene. Nature Genet. 31, 69–73 (2002).
Fraser, H. B., Hirsh, A. E., Giaever, G., Kumm, J. & Eisen, M. B. Noise minimization in eukaryotic gene expression. PLoS Biol. 2, e137 (2004).
Lawrence, J. G. & Ochman, H. Molecular archaeology of the Escherichia coli genome. Proc. Natl Acad. Sci. USA 95, 9413–9417 (1998).
Karlin, S., Campbell, A. M. & Mrazek, J. Comparative DNA analysis across diverse genomes. Annu. Rev. Genet. 32, 185–225 (1998).
Zhang, F., Saha, S., Shabalina, S. A. & Kashina, A. Differential arginylation of actin isoforms is regulated by coding sequence-dependent degradation. Science 329, 1534–1537 (2010).
Thanaraj, T. A. & Argos, P. Ribosome-mediated translational pause and protein domain organization. Protein Sci. 5, 1594–1612 (1996).
Watts, J. M. et al. Architecture and secondary structure of an entire HIV-1 RNA genome. Nature 460, 711–716 (2009).
Warnecke, T., Batada, N. N. & Hurst, L. D. The impact of the nucleosome code on protein-coding sequence evolution in yeast. PLoS Genet. 4, e1000250 (2008).
Eskesen, S. T., Eskesen, F. N. & Ruvinsky, A. Natural selection affects frequencies of AG and GT dinucleotides at the 5′ and 3′ ends of exons. Genetics 167, 543–550 (2004).
Chamary, J. V. & Hurst, L. D. Biased codon usage near intron-exon junctions: selection on splicing enhancers, splice-site recognition or something else? Trends Genet. 21, 256–259 (2005).
Orban, T. I. & Olah, E. Purifying selection on silent sites — a constraint from splicing regulation? Trends Genet. 17, 252–253 (2001).
Warnecke, T. & Hurst, L. D. Evidence for a trade-off between translational efficiency and splicing regulation in determining synonymous codon usage in Drosophila melanogaster. Mol. Biol. Evol. 24, 2755–2762 (2007).
Bettany, A. J. et al. 5′-secondary structure formation, in contrast to a short string of non-preferred codons, inhibits the translation of the pyruvate kinase mRNA in yeast. Yeast 5, 187–198 (1989).
de Smit, M. H. & van Duin, J. Secondary structure of the ribosome binding site determines translational efficiency: a quantitative analysis. Proc. Natl Acad. Sci. USA 87, 7668–7672 (1990). A thorough study, supported by a convincing theoretical model, of how mRNA folding affects translation initiation.
Eyre-Walker, A. & Bulmer, M. Reduced synonymous substitution rate at the start of enterobacterial genes. Nucleic Acids Res. 21, 4599–4603 (1993).
Kudla, G., Murray, A. W., Tollervey, D. & Plotkin, J. B. Coding-sequence determinants of gene expression in Escherichia coli. Science 324, 255–258 (2009). Uses libraries of synthetic genes to isolate the effects of synonymous mutations on expression. See also references 81 and 97.
Gu, W., Zhou, T. & Wilke, C. O. A universal trend of reduced mRNA stability near the translation-initiation site in prokaryotes and eukaryotes. PLoS Comput. Biol. 6, e1000664 (2010).
Tuller, T. et al. An evolutionarily conserved mechanism for controlling the efficiency of protein translation. Cell 141, 344–354 (2010). Shows that rare codons at the beginning of genes could help prevent 'ribosomal traffic jams'. See references 9, 58 and 59 for an alternative interpretation.
Bulmer, M. Codon usage and intragenic position. J. Theor. Biol. 133, 67–71 (1988).
Qin, H., Wu, W. B., Comeron, J. M., Kreitman, M. & Li, W. H. Intragenic spatial patterns of codon usage bias in prokaryotic and eukaryotic genomes. Genetics 168, 2245–2260 (2004).
Zhang, S., Goldman, E. & Zubay, G. Clustering of low usage codons and ribosome movement. J. Theor. Biol. 170, 339–354 (1994).
Fredrick, K. & Ibba, M. How the sequence of a gene can tune its translation. Cell 141, 227–229 (2010).
Cannarozzi, G. et al. A role for codon order in translation dynamics. Cell 141, 355–367 (2010).
Zouridis, H. & Hatzimanikatis, V. Effects of codon distributions and tRNA competition on protein translation. Biophys. J. 95, 1018–1033 (2008).
Huh, W. K. et al. Global analysis of protein localization in budding yeast. Nature 425, 686–691 (2003).
Lu, P., Vogel, C., Wang, R., Yao, X. & Marcotte, E. M. Absolute protein expression profiling estimates the relative contributions of transcriptional and translational regulation. Nature Biotech. 25, 117–124 (2007).
Vogel, C. et al. Sequence signatures and mRNA concentration can explain two-thirds of protein abundance variation in a human cell line. Mol. Syst. Biol. 6, 400 (2010). References 65 and 66 describe a quantitative proteomics approach that promises new insights into the coding-sequence determinants of protein levels.
Ingolia, N. T., Ghaemmaghami, S., Newman, J. R. & Weissman, J. S. Genome-wide analysis in vivo of translation with nucleotide resolution using ribosome profiling. Science 324, 218–223 (2009). A clever method for mapping the positions of ribosomes on messages with unprecedented accuracy; an essential tool for the study of translation kinetics.
Uemura, S. et al. Real-time tRNA transit on single translating ribosomes at codon resolution. Nature 464, 1012–1017 (2010).
Futcher, B., Latter, G. I., Monardo, P., McLaughlin, C. S. & Garrels, J. I. A sampling of the yeast proteome. Mol. Cell. Biol. 19, 7357–7368 (1999).
Schrimpf, S. P. et al. Comparative functional analysis of the Caenorhabditis elegans and Drosophila melanogaster proteomes. PLoS Biol. 7, e48 (2009).
Taniguchi, Y. et al. Quantifying E. coli proteome and transcriptome with single-molecule sensitivity in single cells. Science 329, 533–538.
Feng, L. & Niu, D. K. Relationship between mRNA stability and length: an old question with a new twist. Biochem. Genet. 45, 131–137 (2007).
Arava, Y., Boas, F. E., Brown, P. O. & Herschlag, D. Dissecting eukaryotic translation and its control by ribosome density mapping. Nucleic Acids Res. 33, 2421–2432 (2005).
Welch, M., Villalobos, A., Gustafsson, C. & Minshull, J. You're one in a googol: optimizing genes for protein expression. J. R. Soc. Interface 6 (Suppl. 4), S467–S476 (2009).
Gouy, M. & Gautier, C. Codon usage in bacteria: correlation with gene expressivity. Nucleic Acids Res. 10, 7055–7074 (1982).
Kertesz, M. et al. Genome-wide measurement of RNA secondary structure in yeast. Nature 467, 103–107 (2010).
Takyar, S., Hickerson, R. P. & Noller, H. F. mRNA helicase activity of the ribosome. Cell 120, 49–58 (2005).
Dittmar, K. A., Sorensen, M. A., Elf, J., Ehrenberg, M. & Pan, T. Selective charging of tRNA isoacceptors induced by amino-acid starvation. EMBO Rep. 6, 151–157 (2005).
Elf, J., Nilsson, D., Tenson, T. & Ehrenberg, M. Selective charging of tRNA isoacceptors explains patterns of codon usage. Science 300, 1718–1722 (2003). A detailed theoretical model of what happens to tRNA in cells under starvation conditions.
Ghaemmaghami, S. et al. Global analysis of protein expression in yeast. Nature 425, 737–741 (2003).
Welch, M. et al. Design parameters to control synthetic gene expression in Escherichia coli. PLoS ONE 4, e7002 (2009).
Kudla, G., Lipinski, L., Caffin, F., Helwak, A. & Zylicz, M. High guanine and cytosine content increases mRNA levels in mammalian cells. PLoS Biol. 4, e180 (2006).
Han, J. S., Szak, S. T. & Boeke, J. D. Transcriptional disruption by the L1 retrotransposon and implications for mammalian transcriptomes. Nature 429, 268–274 (2004).
Nackley, A. G. et al. Human catechol-O-methyltransferase haplotypes modulate protein expression by altering mRNA secondary structure. Science 314, 1930–1933 (2006).
Coleman, J. R. et al. Virus attenuation by genome-scale changes in codon pair bias. Science 320, 1784–1787 (2008).
Hayes, C. S., Bose, B. & Sauer, R. T. Stop codons preceded by rare arginine codons are efficient determinants of SsrA tagging in Escherichia coli. Proc. Natl Acad. Sci. USA 99, 3440–3445 (2002).
Spanjaard, R. A. & van Duin, J. Translation of the sequence AGG-AGG yields 50% ribosomal frameshift. Proc. Natl Acad. Sci. USA 85, 7967–7971 (1988).
Kramer, E. B. & Farabaugh, P. J. The frequency of translational misreading errors in E. coli is largely determined by tRNA competition. RNA 13, 87–96 (2007).
Sorensen, M. A., Kurland, C. G. & Pedersen, S. Codon usage determines translation rate in Escherichia coli. J. Mol. Biol. 207, 365–377 (1989).
Chen, G. F. & Inouye, M. Suppression of the negative effect of minor arginine codons on gene expression; preferential usage of minor codons within the first 25 codons of the Escherichia coli genes. Nucleic Acids Res. 18, 1465–1473 (1990).
Angov, E., Hillier, C. J., Kincaid, R. L. & Lyon, J. A. Heterologous protein expression is enhanced by harmonizing the codon usage frequencies of the target gene with those of the expression host. PLoS ONE 3, e2189 (2008).
Rosenberg, A. H., Goldman, E., Dunn, J. J., Studier, F. W. & Zubay, G. Effects of consecutive AGG codons on translation in Escherichia coli, demonstrated with a versatile codon test system. J. Bacteriol. 175, 716–722 (1993).
Gursky, Y. G. & Beabealashvilli, R. The increase in gene expression induced by introduction of rare codons into the C terminus of the template. Gene 148, 15–21 (1994).
Burgess-Brown, N. A. et al. Codon optimization can improve expression of human genes in Escherichia coli: a multi-gene study. Protein Expr. Purif. 59, 94–102 (2008).
Maertens, B. et al. Gene optimization mechanisms: a multi-gene study reveals a high success rate of full-length human proteins expressed in Escherichia coli. Protein Sci. 19, 1312–1326 (2010).
Supek, F. & Smuc, T. On relevance of codon usage to expression of synthetic and natural genes in Escherichia coli. Genetics 185, 1129–1134 (2010).
Voges, D., Watzele, M., Nemetz, C., Wizemann, S. & Buchberger, B. Analyzing and enhancing mRNA translational efficiency in an Escherichia coli in vitro expression system. Biochem. Biophys. Res. Commun. 318, 601–614 (2004).
El Hage, A., French, S. L., Beyer, A. L. & Tollervey, D. Loss of topoisomerase I leads to R-loop-mediated transcriptional blocks during ribosomal RNA synthesis. Genes Dev. 24, 1546–1558 (2010).
McDowall, K. J., Lin-Chao, S. & Cohen, S. N. A+U content rather than a particular nucleotide order determines the specificity of RNase E cleavage. J. Biol. Chem. 269, 10790–10796 (1994).
Nguyen, K. L. et al. Codon optimization of the HIV-1 vpu and vif genes stabilizes their mRNA and allows for highly efficient Rev-independent expression. Virology 319, 163–175 (2004).
Sokolowski, M., Tan, W., Jellne, M. & Schwartz, S. mRNA instability elements in the human papillomavirus type 16 L2 coding region. J. Virol. 72, 1504–1515 (1998).
Bauer, A. P. et al. The impact of intragenic CpG content on gene expression. Nucleic Acids Res. 38, 3891–3908 (2010).
Gutman, G. A. & Hatfield, G. W. Nonrandom utilization of codon pairs in Escherichia coli. Proc. Natl Acad. Sci. USA 86, 3699–3703 (1989).
Irwin, B., Heck, J. D. & Hatfield, G. W. Codon pair utilization biases influence translational elongation step times. J. Biol. Chem. 270, 22801–22806 (1995).
Cheng, L. & Goldman, E. Absence of effect of varying Thr-Leu codon pairs on protein synthesis in a T7 system. Biochemistry 40, 6102–6106 (2001).
Wang, D., Johnson, A. D., Papp, A. C., Kroetz, D. L. & Sadee, W. Multidrug resistance polypeptide 1 (MDR1, ABCB1) variant 3435C>T affects mRNA stability. Pharmacogenet. Genomics 15, 3693–3704 (2005).
Kimchi-Sarfaty, C. et al. A “silent” polymorphism in the MDR1 gene changes substrate specificity. Science 315, 525–528 (2007).
Foster, H. et al. Codon and mRNA sequence optimization of microdystrophin transgenes improves expression and physiological outcome in dystrophic mdx mice following AAV2/8 gene transfer. Mol. Ther. 16, 1825–1832 (2008).
Arruda, V. R. et al. Peripheral transvenular delivery of adeno-associated viral vectors to skeletal muscle as a novel therapy for hemophilia B. Blood 115, 4678–4688.
Fuglsang, A. The relationship between palindrome avoidance and intragenic codon usage variations: a Monte Carlo study. Biochem. Biophys. Res. Commun. 316, 755–762 (2004).
Drummond, D. A. & Wilke, C. O. The evolutionary consequences of erroneous protein synthesis. Nature Rev. Genet. 10, 715–724 (2009). A readable Review that summarizes the various types of errors that occur in protein synthesis, many of which are directly related to codon usage.
Akashi, H., Kliman, R. M. & Eyre-Walker, A. Mutation pressure, natural selection, and the evolution of base composition in Drosophila. Genetica 102–103, 49–60 (1998).
Marais, G. & Duret, L. Synonymous codon usage, accuracy of translation, and gene length in Caenorhabditis elegans. J. Mol. Evol. 52, 275–280 (2001).
Higgs, P. G. & Ran, W. Coevolution of codon usage and tRNA genes leads to alternative stable states of biased codon usage. Mol. Biol. Evol. 25, 2279–2291 (2008).
Shah, P. & Gilchrist, M. Effect of correlated tRNA abundances on translation errors and evolution of codon usage bias. PLoS Genet. 6, e1001128 (2010).
Bernardi, G. et al. The mosaic genome of warm-blooded vertebrates. Science 228, 953–958 (1985).
Galtier, N., Piganeau, G., Mouchiroud, D. & Duret, L. GC-content evolution in mammalian genomes: the biased gene conversion hypothesis. Genetics 159, 907–911 (2001).
Urrutia, A. O. & Hurst, L. D. The signature of selection mediated by expression on human genes. Genome Res. 13, 2260–2264 (2003).
Comeron, J. M. Selective and mutational patterns associated with gene expression in humans: influences on synonymous composition and intron presence. Genetics 167, 1293–1304 (2004).
Karlin, S. & Mrazek, J. What drives codon choices in human genes? J. Mol. Biol. 262, 459–472 (1996).
Plotkin, J. B., Robins, H. & Levine, A. J. Tissue-specific codon usage and the expression of human genes. Proc. Natl Acad. Sci. USA 101, 12588–12591 (2004).
Dittmar, K. A., Goodenbour, J. M. & Pan, T. Tissue-specific differences in human transfer RNA expression. PLoS Genet. 2, e221 (2006).
Semon, M., Lobry, J. R. & Duret, L. No evidence for tissue-specific adaptation of synonymous codon usage in human. Mol. Biol. Evol. 23, 523–529 (2005).
Chamary, J. V. & Hurst, L. D. Evidence for selection on synonymous mutations affecting stability of mRNA secondary structure in mammals. Genome Biol. 6, R75 (2005).
Seffens, W. & Digby, D. mRNAs have greater negative folding free energies than shuffled or codon choice randomized sequences. Nucleic Acids Res. 27, 1578–1584 (1999).
Duan, J. et al. Synonymous mutations in the human dopamine receptor D2 (DRD2) affect mRNA stability and synthesis of the receptor. Hum. Mol. Genet. 12, 205–216 (2003).
Sharp, P. M., Tuohy, T. M. & Mosurski, K. R. Codon usage in yeast: cluster analysis clearly differentiates highly and lowly expressed genes. Nucleic Acids Res. 14, 5125–5143 (1986).
dos Reis, M., Savva, R. & Wernisch, L. Solving the riddle of codon usage preferences: a test for translational selection. Nucleic Acids Res. 32, 5036–5044 (2004).
Nivinskas, R., Malys, N., Klausa, V., Vaiskunaite, R. & Gineikiene, E. Post-transcriptional control of bacteriophage T4 gene 25 expression: mRNA secondary structure that enhances translational initiation. J. Mol. Biol. 288, 291–304 (1999).
Kozak, M. Downstream secondary structure facilitates recognition of initiator codons by eukaryotic ribosomes. Proc. Natl Acad. Sci. USA 87, 8301–8305 (1990).
Paulus, M., Haslbeck, M. & Watzele, M. RNA stem-loop enhanced expression of previously non-expressible genes. Nucleic Acids Res. 32, e78 (2004).
Zhang, G., Hubalewska, M. & Ignatova, Z. Transient ribosomal attenuation coordinates protein synthesis and co-translational folding. Nature Struct. Mol. Biol. 16, 274–280 (2009).
Nakamura, Y., Gojobori, T. & Ikemura, T. Codon usage tabulated from international DNA sequence databases: status for the year 2000. Nucleic Acids Res. 28, 292 (2000).
Zolotukhin, S., Potter, M., Hauswirth, W. W., Guy, J. & Muzyczka, N. A “humanized” green fluorescent protein cDNA adapted for high-level expression in mammalian cells. J. Virol. 70, 4646–4654 (1996).
Markham, N. R. & Zuker, M. DINAMelt web server for nucleic acid melting prediction. Nucleic Acids Res. 33, W577–W581 (2005).
Lesnik, E. A. et al. Prediction of rho-independent transcriptional terminators in Escherichia coli. Nucleic Acids Res. 29, 3583–3594 (2001).
Bernstein, J. A., Khodursky, A. B., Lin, P. H., Lin-Chao, S. & Cohen, S. N. Global analysis of mRNA decay and abundance in Escherichia coli at single-gene resolution using two-color fluorescent DNA microarrays. Proc. Natl Acad. Sci. USA 99, 9697–9702 (2002).
Lewis, B. P., Shih, I. H., Jones-Rhoades, M. W., Bartel, D. P. & Burge, C. B. Prediction of mammalian microRNA targets. Cell 115, 787–798 (2003).
Jacobs, G. H. et al. Transterm: a database to aid the analysis of regulatory sequences in mRNAs. Nucleic Acids Res. 37, D72–D76 (2009).
Peden, J. F. Analysis of Codon Usage. Thesis, Dept of Genetics, Univ. Nottingham (1999).
Supek, F. & Vlahovicek, K. INCA: synonymous codon usage analysis and clustering by means of self-organizing map. Bioinformatics 20, 2329–2330 (2004).
Pagani, F., Raponi, M. & Baralle, F. E. Synonymous mutations in CFTR exon 12 affect splicing and are not neutral in evolution. Proc. Natl Acad. Sci. USA 102, 6368–6372 (2005).
Burge, C., Campbell, A. M. & Karlin, S. Over- and under-representation of short oligonucleotides in DNA sequences. Proc. Natl Acad. Sci. USA 89, 1358–1362 (1992).
Villalobos, A., Ness, J. E., Gustafsson, C., Minshull, J. & Govindarajan, S. Gene Designer: a synthetic biology tool for constructing artificial DNA segments. BMC Bioinformatics 7, 285 (2006).
Acknowledgements
We thank L. Hurst and C. Wilke for helpful discussions. We apologize to those whose work we were unable to cite because of space constraints. G.K. acknowledges funding from the Wellcome Trust. J.B.P. acknowledges support from the Burroughs Wellcome Fund, the David and Lucile Packard Foundation, the James S. McDonnell Foundation, the Alfred P. Sloan Foundation, the Defense Advanced Research Projects Agency (HR0011-05-1-0057) and the US National Institute of Allergy and Infectious Diseases (2U54AI057168).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
Glossary
- Iso-accepting tRNAs
-
A subset of tRNAs that carry the same amino acid.
- Negative selection
-
A form of natural selection that suppresses alternative genetic variants in favour of the wild type.
- Horizontal gene transfer
-
The transfer of genetic material from one species into another.
- Isochore
-
A large fragment of a chromosome that is characterized by homogeneous GC content.
- Ribosomal pausing
-
A temporary arrest of the ribosome during translation elongation.
- Effective population size
-
The number of individuals in a population that produce viable offspring.
- Biased gene conversion
-
A recombination event in which one variant of genomic sequence is preferentially 'copied and pasted' onto another one.
- Fourfold degenerate sites
-
Positions within the coding sequence of a gene at which all four nucleotides encode the same amino acid.
- Shotgun proteomics
-
Methods of quantifying protein levels in a complex sample, typically using mass spectrometry.
- Codon adaptation index
-
A measure of similarity between the codon usage of a gene and the average codon usage of highly expressed genes in a species.
- RNA–seq
-
Quantitative analysis of RNA in a complex sample by high-throughput sequencing.
- Ribosomal footprint
-
A fragment of mRNA that is protected by ribosomes from nuclease digestion in a ribosomal-profiling experiment.
- Upstream ORFs
-
ORFs that are located 5′ from the primary ORF. They are thought to inhibit translation of the primary ORF.
Rights and permissions
About this article
Cite this article
Plotkin, J., Kudla, G. Synonymous but not the same: the causes and consequences of codon bias. Nat Rev Genet 12, 32–42 (2011). https://doi.org/10.1038/nrg2899
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrg2899
This article is cited by
-
Codon usage characterization and phylogenetic analysis of the mitochondrial genome in Hemerocallis citrina
BMC Genomic Data (2024)
-
Engineering strategies for enhanced heterologous protein production by Saccharomyces cerevisiae
Microbial Cell Factories (2024)
-
Interconnected Codons: Unravelling the Epigenetic Significance of Flanking Sequences in CpG Dyads
Journal of Molecular Evolution (2024)
-
GC heterogeneity reveals sequence-structures evolution of angiosperm ITS2
BMC Plant Biology (2023)
-
The genome of Candidatus phytoplasma ziziphi provides insights into their biological characteristics
BMC Plant Biology (2023)