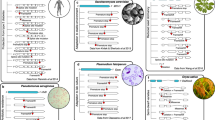
Key Points
-
The divergence of new genes and proteins occurs through mutations that modulate protein function. The effects of these mutations are pleiotropic, thus imposing trade-offs between selection pressures for the existing function and the newly evolving one and among the protein's activity, stability and dosage.
-
Various compensatory and buffering mechanisms, such as gene duplication, upregulation of expression, stabilizing mutations and chaperone folding assistance, can alleviate these trade-offs and so facilitate functional divergence.
-
Despite buffering effects, the fitness distribution of mutations at the protein level, and for whole organisms, is such that most of the mutations are either neutral or deleterious. This results in the rapid and irreversible non-functionalization of proteins that accumulate mutations under no selection.
-
The distribution of fitness effects of mutations for whole organisms is comparable, and possibly even more deleterious, than that of protein mutations.
-
Duplication underlies the divergence of new genes and proteins. Duplication is almost as frequent as point mutations and is a common mechanism for resolving the trade-off conflicts that arise owing to parallel selection pressures. These pressures may regard the existing and the new function and maintenance of the protein's structural stability.
-
Duplication, and the emergence of new genes and proteins, may occur at different stages of the divergence process. The selection pressures that act on the gene and its duplicate may differ, giving rise to different mechanisms of divergence. These mechanisms are described under three schematic models — Ohno's model, the 'divergence before duplication' (DPD) model and the sub-functionalization model.
-
In Ohno's model of divergence, duplication is a neutral event. The duplicated copy of the protein drifts under no selection until a new function becomes under selection. The downside of this model is that under no selection, non-functionalization of the drifting protein is inevitable. Its advantage is that divergence is independent of trade-offs between the new and existing functions.
-
The DPD model is based on a 'generalist' intermediate that confers a selectable degree of both the new and existing functions. Duplication occurs after the acquisition of a new function, and occurs under positive selection to increase protein dosage and/or alleviate trade-offs that make the acquisition of new function depend on loss of the existing one.
-
The sub-functionalization model combines elements of the DPD model and Ohno's model. Duplication is initially a neutral event, but once mutations that partially reduce protein activity or dosage appear, both copies must remain functional. Duplication therefore enables a larger genetic variability to accumulate, thereby facilitating the emergence of new functions.
-
The DPD and sub-functionalization models are both based on mutations with adaptive potential initially accumulating as neutral. As such, they are related to the notions of hidden or apparently neutral variation and of neutral networks.
Abstract
The divergence of new genes and proteins occurs through mutations that modulate protein function. However, mutations are pleiotropic and can have different effects on organismal fitness depending on the environment, as well as opposite effects on protein function and dosage. We review the pleiotropic effects of mutations. We discuss how they affect the evolution of gene and protein function, and how these complex mutational effects dictate the likelihood and mechanism of gene duplication and divergence. We propose several factors that can affect the divergence of new protein functions, including mutational trade-offs and hidden, or apparently neutral, variation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Conant, G. C. & Wolfe, K. H. Turning a hobby into a job: how duplicated genes find new functions. Nature Rev. Genet. 9, 938–950 (2008).
Innan, H. & Kondrashov, F. The evolution of gene duplications: classifying and distinguishing between models. Nature Rev. Genet. 11, 97–108 (2010).
DePristo, M. A., Weinreich, D. M. & Hartl, D. L. Missense meanderings in sequence space: a biophysical view of protein evolution. Nature Rev. Genet. 6, 678–687 (2005).
Pal, C., Papp, B. & Lercher, M. J. An integrated view of protein evolution. Nature Rev. Genet. 7, 337–348 (2006).
Dean, A. M. & Thornton, J. W. Mechanistic approaches to the study of evolution: the functional synthesis. Nature Rev. Genet. 8, 675–688 (2007).
Tokuriki, N. & Tawfik, D. S. Stability effects of mutations and protein evolvability. Curr. Opin. Struct. Biol. 19, 596–604 (2009).
Eyre-Walker, A. & Keightley, P. D. The distribution of fitness effects of new mutations. Nature Rev. Genet. 8, 610–618 (2007).
Camps, M., Herman, A., Loh, E. & Loeb, L. A. Genetic constraints on protein evolution. Crit. Rev. Biochem. Mol. Biol. 42, 313–326 (2007).
Bloom, J. D. et al. Thermodynamic prediction of protein neutrality. Proc. Natl Acad. Sci. USA 102, 606–611 (2005).
Bershtein, S., Segal, M., Bekerman, R., Tokuriki, N. & Tawfik, D. S. Robustness–epistasis link shapes the fitness landscape of a randomly drifting protein. Nature 444, 929–932 (2006).
Bershtein, S. & Tawfik, D. S. Ohno's model revisited: measuring the frequency of potentially adaptive mutations under various mutational drifts. Mol. Biol. Evol. 25, 2311–2318 (2008).
Hecky, J. & Muller, K. M. Structural perturbation and compensation by directed evolution at physiological temperature leads to thermostabilization of β-lactamase. Biochemistry 44, 12640–12654 (2005).
Yue, P. & Moult, J. Identification and analysis of deleterious human SNPs. J. Mol. Biol. 356, 1263–1274 (2006).
Tokuriki, N., Oldfield, C. J., Uversky, V. N., Berezovsky, I. N. & Tawfik, D. S. Do viral proteins possess unique biophysical features? Trends Biochem. Sci. 34, 53–59 (2009).
Wang, X., Minasov, G. & Shoichet, B. K. Evolution of an antibiotic resistance enzyme constrained by stability and activity trade-offs. J. Mol. Biol. 320, 85–95 (2002).
Tokuriki, N., Stricher, F., Serrano, L. & Tawfik, D. S. How protein stability and new functions trade off. PLoS Comput. Biol. 4, e1000002 (2008).
Levin, K. B. et al. Following evolutionary paths to protein–protein interactions with high affinity and selectivity. Nature Struct. Mol. Biol. 16, 1049–1055 (2009).
Lindner, A. B., Madden, R., Demarez, A., Stewart, E. J. & Taddei, F. Asymmetric segregation of protein aggregates is associated with cellular aging and rejuvenation. Proc. Natl Acad. Sci. USA 105, 3076–3081 (2008).
McLoughlin, S. Y. & Copley, S. D. A compromise required by gene sharing enables survival: implications for evolution of new enzyme activities. Proc. Natl Acad. Sci. USA 105, 13497–13502 (2008).
Vick, J. E., Schmidt, D. M. & Gerlt, J. A. Evolutionary potential of (β/α)8-barrels: in vitro enhancement of a 'new' reaction in the enolase superfamily. Biochemistry 44, 11722–11729 (2005).
Khersonsky, O. & Tawfik, D. S. Enzyme promiscuity: a mechanistic and evolurtionary perpective. Ann. Rev. Biochem. 79, 471–505 (2010).
Aharoni, A. et al. The 'evolvability' of promiscuous protein functions. Nature Genet. 37, 73–76 (2005).
Tokuriki, N. & Tawfik, D. S. Protein dynamism and evolvability. Science 324, 203–207 (2009).
Scannell, D. R. & Wolfe, K. H. A burst of protein sequence evolution and a prolonged period of asymmetric evolution follow gene duplication in yeast. Genome Res. 18, 137–147 (2008).
Kaessmann, H. Genetics. More than just a copy. Science 325, 958–959 (2009).
Parker, H. G. et al. An expressed fgf4 retrogene is associated with breed-defining chondrodysplasia in domestic dogs. Science 325, 995–998 (2009).
Andersson, D. I. & Hughes, D. Gene amplification and adaptive evolution in bacteria. Annu. Rev. Genet. 43, 167–195 (2009).
Schimke, R. T. Gene amplification in cultured cells. J. Biol. Chem. 263, 5989–5992 (1988).
Papp, B., Pal, C. & Hurst, L. D. Metabolic network analysis of the causes and evolution of enzyme dispensability in yeast. Nature 429, 661–664 (2004).
Perry, G. H. et al. Diet and the evolution of human amylase gene copy number variation. Nature Genet. 39, 1256–1260 (2007).
Fablet, M., Bueno, M., Potrzebowski, L. & Kaessmann, H. Evolutionary origin and functions of retrogene introns. Mol. Biol. Evol. 26, 2147–2156 (2009).
Jablonka, E. & Lamb, M. J. Epigenetic Inheritance and Evolution: The Lamarckian Dimension (Oxford Univ. Press, Oxford, UK, 1995).
Steele, E. J., Lindley, R. A. & Blanden, R. V. Lamarck's Signature: How Retrogenes Are Changing Darwin's Natural Selection Paradigm, (Allen & Unwin; Perseus Books, Australia, 1988).
Chen, G. K. et al. Preferential expression of a mutant allele of the amplified MDR1 (ABCB1) gene in drug-resistant variants of a human sarcoma. Genes Chromosomes Cancer 34, 372–383 (2002).
Qian, W. & Zhang, J. Gene dosage and gene duplicability. Genetics 179, 2319–2324 (2008).
Goldsmith, M. & Tawfik, D. S. Potential role of phenotypic mutations in the evolution of protein expression and stability. Proc. Natl Acad. Sci. USA 106, 6197–6202 (2009).
Siu, L. K., Ho, P. L., Yuen, K. Y., Wong, S. S. & Chau, P. Y. Transferable hyperproduction of TEM-1 β-lactamase in Shigella flexneri due to a point mutation in the pribnow box. Antimicrob. Agents Chemother. 41, 468–470 (1997).
Hall, B. G. Evolution of a regulated operon in the laboratory. Genetics 101, 335–344 (1982).
Hall, B. G. The EBG system of E. coli: origin and evolution of a novel β-galactosidase for the metabolism of lactose. Genetica 118, 143–156 (2003).
Stoebel, D. M., Dean, A. M. & Dykhuizen, D. E. The cost of expression of Escherichia coli lac operon proteins is in the process, not in the products. Genetics 178, 1653–1660 (2008).
Wagner, A. Energy constraints on the evolution of gene expression. Mol. Biol. Evol. 22, 1365–1374 (2005).
Vavouri, T., Semple, J. I., Garcia-Verdugo, R. & Lehner, B. Intrinsic protein disorder and interaction promiscuity are widely associated with dosage sensitivity. Cell 138, 198–208 (2009).
Veitia, R. A. Gene dosage balance: deletions, duplications and dominance. Trends Genet. 21, 33–35 (2005).
Drummond, D. A., Bloom, J. D., Adami, C., Wilke, C. O. & Arnold, F. H. Why highly expressed proteins evolve slowly. Proc. Natl Acad. Sci. USA 102, 14338–14343 (2005).
Fares, M. A., Ruiz- González, M. X., Moya, A., Elena, S. F. & Barrio, E. Endosymbiotic bacteria: GroEL buffers against deleterious mutations. Nature 417, 398 (2002).
Rutherford, S., Hirate, Y. & Swalla, B. J. The Hsp90 capacitor, developmental remodeling, and evolution: the robustness of gene networks and the curious evolvability of metamorphosis. Crit. Rev. Biochem. Mol. Biol. 42, 355–372 (2007).
Cowen, L. E. & Lindquist, S. Hsp90 potentiates the rapid evolution of new traits: drug resistance in diverse fungi. Science 309, 2185–2189 (2005).
Parent, K. N., Ranaghan, M. J. & Teschke, C. M. A second-site suppressor of a folding defect functions via interactions with a chaperone network to improve folding and assembly in vivo. Mol. Microbiol. 54, 1036–1050 (2004).
Tokuriki, N. & Tawfik, D. S. Chaperonin overexpression promotes genetic variation and enzyme evolution. Nature 459, 668–673 (2009).
Zhang, L. & Watson, L. T. Analysis of the fitness effect of compensatory mutations. HFSP J. 3, 47–54 (2009).
Bershtein, S., Goldin, K. & Tawfik, D. S. Intense neutral drifts yield robust and evolvable consensus proteins. J. Mol. Biol. 379, 1029–1044 (2008).
Hecky, J., Mason, J. M., Arndt, K. M. & Muller, K. M. A general method of terminal truncation, evolution, and re-elongation to generate enzymes of enhanced stability. Methods Mol. Biol. 352, 275–304 (2007).
Kather, I., Jakob, R. P., Dobbek, H. & Schmid, F. X. Increased folding stability of TEM-1 β-lactamase by in vitro selection. J. Mol. Biol. 383, 238–251 (2008).
Marciano, D. C. et al. Genetic and structural characterization of an L201P global suppressor substitution in TEM-1 β-lactamase. J. Mol. Biol. 384, 151–164 (2008).
Kimura, M. The role of compensatory neutral mutations in molecular evolution. J. Genet. 64, 7–19 (1985).
Bloom, J. D., Labthavikul, S. T., Otey, C. R. & Arnold, F. H. Protein stability promotes evolvability. Proc. Natl Acad. Sci. USA 103, 5869–5874 (2006).
McIntosh, B. E., Hogenesch, J. B. & Bradfield, C. A. Mammalian Per-Arnt-Sim proteins in environmental adaptation. Annu. Rev. Physiol. 72, 625–645 (2010).
Lynch, M. Genomics. Gene duplication and evolution. Science 297, 945–947 (2002).
Beckmann, J. S., Estivill, X. & Antonarakis, S. E. Copy number variants and genetic traits: closer to the resolution of phenotypic to genotypic variability. Nature Rev. Genet. 8, 639–646 (2007).
Hastings, P. J., Lupski, J. R., Rosenberg, S. M. & Ira, G. Mechanisms of change in gene copy number. Nature Rev. Genet. 10, 551–564 (2009).
Liao, B. Y. & Zhang, J. Null mutations in human and mouse orthologs frequently result in different phenotypes. Proc. Natl Acad. Sci. USA 105, 6987–6992 (2008).
Ohno, S. Evolution by Gene Duplication (Allen & Unwin; Springer, New York, 1970).
Kimura, M. & Ota, T. On some principles governing molecular evolution. Proc. Natl Acad. Sci. USA 71, 2848–2852 (1974).
Zhang, J. Evolution by gene duplication: an update. Trends Ecol. Evol. 18, 292–298 (2003).
Hughes, A. L. Adaptive evolution after gene duplication. Trends Genet. 18, 433–434 (2002).
Lynch, M. & Katju, V. The altered evolutionary trajectories of gene duplicates. Trends Genet. 20, 544–549 (2004).
Kondrashov, F. A. & Koonin, E. V. A common framework for understanding the origin of genetic dominance and evolutionary fates of gene duplications. Trends Genet. 20, 287–290 (2004).
Bergthorsson, U., Andersson, D. I. & Roth, J. R. Ohno's dilemma: evolution of new genes under continuous selection. Proc. Natl Acad. Sci. USA 104, 17004–17009 (2007).
Kondrashov, F. A. In search of the limits of evolution. Nature Genet. 37, 9–10 (2005).
Boehr, D. D., Nussinov, R. & Wright, P. E. The role of dynamic conformational ensembles in biomolecular recognition. Nature Chem. Biol. 5, 789–796 (2009).
Piatigorsky, J. et al. Gene sharing by D-crystallin and argininosuccinate lyase. Proc. Natl Acad. Sci. USA 85, 3479–3483 (1988).
Piatigorsky, J. Gene Sharing and Evolution: The Diversity of Protein Functions, (Harvard Univ. Press, Cambridge, Massachusetts, USA; London, UK, 2007).
Lee, Y. N., Nechushtan, H., Figov, N. & Razin, E. The function of lysyl-tRNA synthetase and Ap4A as signaling regulators of MITF activity in FceRI-activated mast cells. Immunity 20, 145–151 (2004).
Sedlak, T. W. & Snyder, S. H. Messenger molecules and cell death: therapeutic implications. JAMA 295, 81–89 (2006).
Rosenberg, H. F. RNase A ribonucleases and host defense: an evolving story. J. Leukoc. Biol. 83, 1079–1087 (2008).
Jensen, R. A. Enzyme recruitment in evolution of new function. Annu. Rev. Microbiol. 30, 409–425 (1974).
O'Brien, P. J. & Herschlag, D. Catalytic promiscuity and the evolution of new enzymatic activities. Chem. Biol. 6, R91–R105 (1999).
Palmer, D. R. et al. Unexpected divergence of enzyme function and sequence: 'N-acylamino acid racemase' is o-succinylbenzoate synthase. Biochemistry 38, 4252–4258 (1999).
James, L. C. & Tawfik, D. S. Catalytic and binding poly-reactivities shared by two unrelated proteins: the potential role of promiscuity in enzyme evolution. Protein Sci. 10, 2600–2607 (2001).
Afriat, L., Roodveldt, C., Manco, G. & Tawfik, D. S. The latent promiscuity of newly identified microbial lactonases is linked to a recently diverged phosphotriesterase. Biochemistry 45, 13677–13686 (2006).
Copley, S. D. Evolution of efficient pathways for degradation of anthropogenic chemicals. Nature Chem. Biol. 5, 559–566 (2009).
Copley, S. D. Comprehensive Natural Products II: Chemistry and Biology (eds Mander, L. & Liu, H.-W.) (Elsevier, Oxford, 2010).
Hughes, A. L. The evolution of functionally novel proteins after gene duplication. Proc. Biol. Sci. 256, 119–124 (1994).
Barkman, T. & Zhang, J. Evidence for escape from adaptive conflict? Nature 462, e1; discussion e2–e3 (2009).
Des Marais, D. L. & Rausher, M. D. Escape from adaptive conflict after duplication in an anthocyanin pathway gene. Nature 454, 762–765 (2008).
Lynch, M. & Force, A. The probability of duplicate gene preservation by subfunctionalization. Genetics 154, 459–473 (2000).
Dykhuizen, D. & Hartl, D. L. Selective neutrality of 6PGD allozymes in E. coli and the effects of genetic background. Genetics 96, 801–817 (1980).
Force, A. et al. Preservation of duplicate genes by complementary, degenerative mutations. Genetics 151, 1531–1545 (1999).
Nei, M. The new mutation theory of phenotypic evolution. Proc. Natl Acad. Sci. USA 104, 12235–12242 (2007).
Wagner, A. Robustness and Evolvability in Living Systems (Princeton Univ. Press, Princeton, USA, 2005).
Schuster, P. & Fontana, W. Chance and necessity in evolution: lessons from RNA. Physica D 133, 427–452 (1999).
Wroe, R., Chan, H. S. & Bornberg-Bauer, E. A structural model of latent evolutionary potentials underlying neutral networks in proteins. HFSP J. 1, 79–87 (2007).
Klassen, J. L. Pathway evolution by horizontal transfer and positive selection is accommodated by relaxed negative selection upon upstream pathway genes in purple bacterial carotenoid biosynthesis. J. Bacteriol. 191, 7500–7508 (2009).
Wloch, D. M., Szafraniec, K., Borts, R. H. & Korona, R. Direct estimate of the mutation rate and the distribution of fitness effects in the yeast Saccharomyces cerevisiae. Genetics 159, 441–452 (2001).
Kivisaar, M. Degradation of nitroaromatic compounds: a model to study evolution of metabolic pathways. Mol. Microbiol. 74, 777–781 (2009).
Wackett, L. P. Questioning our perceptions about evolution of biodegradative enzymes. Curr. Opin. Microbiol. 12, 244–251 (2009).
Newcomb, R. D., Gleeson, D. M., Yong, C. G., Russell, R. J. & Oakeshott, J. G. Multiple mutations and gene duplications conferring organophosphorus insecticide resistance have been selected at the Rop-1 locus of the sheep blowfly, Lucilia cuprina. J. Mol. Evol. 60, 207–220 (2005).
Patzoldt, W. L., Hager, A. G., McCormick, J. S. & Tranel, P. J. A codon deletion confers resistance to herbicides inhibiting protoporphyrinogen oxidase. Proc. Natl Acad. Sci. USA 103, 12329–12334 (2006).
O'Maille, P. E. et al. Quantitative exploration of the catalytic landscape separating divergent plant sesquiterpene synthases. Nature Chem. Biol. 4, 617–623 (2008).
Lozovsky, E. R. et al. Stepwise acquisition of pyrimethamine resistance in the malaria parasite. Proc. Natl Acad. Sci. USA 106, 12025–12030 (2009).
Poelwijk, F. J., Kiviet, D. J., Weinreich, D. M. & Tans, S. J. Empirical fitness landscapes reveal accessible evolutionary paths. Nature 445, 383–386 (2007).
Kondrashov, A. S., Sunyaev, S. & Kondrashov, F. A. Dobzhansky–Muller incompatibilities in protein evolution. Proc. Natl Acad. Sci. USA 99, 14878–14883 (2002).
Weinreich, D. M., Delaney, N. F., Depristo, M. A. & Hartl, D. L. Darwinian evolution can follow only very few mutational paths to fitter proteins. Science 312, 111–114 (2006).
Acknowledgements
D.S.T. is the incumbent of the Nella and Leon Benoziyo Professorial Chair. Financial support from the Meil de Botton Aynsley and the EU network BioModularH2 are gratefully acknowledged. We are very grateful to S. Bershtein, N. Tokuriki, F. Kondrashov and J. G. Zhang for their insightful comments regarding this manuscript and to A. Eyre-Walker for providing the data for the figure in Box 1.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Glossary
- Protein mutations
-
Missense mutations that occur in encoded open reading frames.
- Trade-offs
-
Gains of a new activity or property at the expense of other activities or properties.
- Protein stability
-
The capacity of a protein to adopt its native, functional structure. Stability also correlates with cellular protein levels.
- Sub-functionalization
-
Degenerate mutations that result in a gene and its duplicated copy sharing the burden of one function.
- Negative epistasis
-
The combined effect of mutations being more deleterious than expected from their individual effects.
- Protein fitness
-
Levels of physiological function exerted by a given protein variant under a certain selection pressure.
- Non-functionalization
-
The complete inactivation of a gene or protein by highly deleterious mutations.
- Neo-functionalization
-
The divergence of a duplicated gene or protein to execute a new function.
- ΔΔG
-
The stability difference for a protein variant versus its wild-type reference (ΔΔG > 0 indicates lower stability).
- Disordered domains
-
Protein domains with a high degree of random coil and loop regions and a low degree of highly ordered secondary structure.
- Apparently neutral mutations
-
Mutations that have no significant or observable fitness effect under a given environment.
- New-function mutations
-
Mutations that mediate changes in protein activity, typically by increasing a weak, latent promiscuous function.
- New–existing function trade-offs
-
The acquisition of a new function through mutations that undermine the existing function.
- Chaperones
-
Proteins that mediate the correct folding and assembly of other proteins.
- Specialists
-
Genes or proteins that exert one specific function with high proficiency.
- Generalist
-
A gene or protein that exerts multiple functions, typically one primary function and additional secondary or promiscuous functions.
- New-function–stability trade-offs
-
Mutations that increase the new, evolving function but reduce protein stability and protein dosage.
- Productive variation
-
Genetic variation that does not compromise fitness in the dwelling environment but holds the potential for adaptation to new environments.
Rights and permissions
About this article
Cite this article
Soskine, M., Tawfik, D. Mutational effects and the evolution of new protein functions. Nat Rev Genet 11, 572–582 (2010). https://doi.org/10.1038/nrg2808
Issue Date:
DOI: https://doi.org/10.1038/nrg2808
This article is cited by
-
Modelling genetic stability in engineered cell populations
Nature Communications (2023)
-
Random with Respect to Fitness or External Selection? An Important but Often Overlooked Distinction
Acta Biotheoretica (2023)
-
Low protein expression enhances phenotypic evolvability by intensifying selection on folding stability
Nature Ecology & Evolution (2022)
-
In silico evolution of nucleic acid-binding proteins from a nonfunctional scaffold
Nature Chemical Biology (2022)
-
Multi-omics network model reveals key genes associated with p-coumaric acid stress response in an industrial yeast strain
Scientific Reports (2022)