Key Points
-
Hox, ParaHox and NK genes are related ANTP-class homeobox genes that are organized into chromosomal clusters in several metazoan lineages. Both Hox and ParaHox clusters are thought to have arisen by duplication and divergence from an ancient ProtoHox cluster that was present in an ancestral animal.
-
Phylogenetic and chromosomal reconstructions indicate that the Hox and ParaHox clusters, as well as other ANTP homeobox classes (Evx, Meox, Gbx, Mnx and En) originated after successive gene duplications and chromosomal breakages from a founder ProtoHox-like gene.
-
A large array of ANTP-class homeobox genes — the homeobox megacluster that includes the Hox, ParaHox and NK clusters — existed early in metazoan evolution. It contained up to 25 homeobox genes, and probably originated before the divergence of cnidarians and bilaterians.
-
An NK cluster of seven homeobox genes existed before the divergence of protostomes and deuterosomes. The organization of the cluster has been maintained largely intact in fruitflies and mosquitoes, but has been split into three in chordates.
-
Hox, ParaHox and NK cluster genes are preferentially expressed in ectodermal, endodermal and mesodermal derivatives, respectively. I speculate that the origin of the three gene clusters was related to the origin, patterning and diversification of the three bilaterian germ layers.
-
The above thesis implies that cnidarians are primitively bilaterian triploblasts; that these gene clusters have been evolving independently in the distinct clades; and that their present-day functions are a composite of ancestral and derived functions.
-
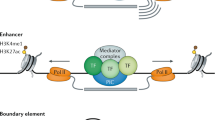
The selective constraint that has favoured the maintenance of homeobox gene clustering is probably the need for close gene linkage; this would enable sequential chromatin decondensation and thereby allow the temporal deployment of gene expression.
-
The linkage constraint is maintained for the Hox and ParaHox clusters in animals that have a slow mode of development, and for the NK cluster in insects. In the lineages for which the constraint has disappeared, the clusters tend to disintegrate by phylogenetic inertia.
Abstract
Once called the 'Rosetta stone' of developmental biology, the homeobox continues to fascinate both evolutionary and developmental biologists. The birth of the homeotic, or Hox, gene cluster, and its subsequent evolution, has been crucial in mediating the major transitions in metazoan body plan. Comparative genomics studies indicate that the more recently discovered ParaHox and NK clusters were linked to the Hox cluster early in evolution, and that together they constituted a 'megacluster' of homeobox genes that conspicuously contributed to body-plan evolution.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
McGinnis, W. & Krumlauf, R. Homeobox genes and axial patterning. Cell 68, 283–302 (1992).
Duboule, D. Temporal collinearity and the phylotypic progression: a basis for the stability of a vertebrate Bauplan and the evolution of morphologies through heterochrony. Development Suppl. 135–142 (1994).
Kmita, M. & Duboule, D. Organizing axes in time and space: 25 years of collinear tinkering. Science 301, 331–333 (2003).
Hurst, L. D., Pá l, C. & Lercher, M. J. The evolutionary dynamics of eukaryotic gene order. Nature Rev. Genet. 5, 299–310 (2004).
Brooke, N. M., Garcia-Fernà ndez, J. & Holland, P. W. H. The ParaHox gene cluster is an evolutionary sister of the Hox gene cluster. Nature 392, 920–922 (1998).
Holland, L. Z. Non-neural ectoderm is really neural: evolution of developmental patterning mechanisms in the non-neural ectoderm of chordates and the problem of sensory cell homologies. J. Exp. Zool. B 304B, 1–20 (2005).
Jagla, K., Bellard, M. & Frasch, M. A cluster of Drosophila homeobox genes involved in mesoderm differentiation programs. Bioessays 23, 125–133 (2001). This paper presents a complete description of linkage information and expression data for the NK cluster genes in D. melanogaster , the organism in which temporal collinearity of the NK cluster genes in the mesoderm was first noticed.
Martínez, P. & Amemiya, C. T. Genomics of the HOX gene cluster. Comp. Biochem. Physiol. B 133, 571–580 (2002).
Prince, E. The Hox paradox: more complex(s) than imagined. Dev. Biol. 249, 1–15 (2002).
Ferrier, D. E. K. & Minguillón, C. Evolution of the Hox/ParaHox gene clusters. Int. J. Dev. Biol. 47, 605–611 (2003).
de Rosa, R. et al. Hox genes in brachiopods and priapulids and protostome evolution. Nature 399, 772–776 (1999).
Powers, T. P. & Amemiya, C. T. Evidence for a Hox14 paralog group in vertebrates. Curr. Biol. 14, R183–R184 (2004).
Minguillón C. et al. No more than 14: the end of the amphioxus Hox cluster. Int. J. Biol. Sci. 1, 19–23 (2005).
Hooman, K., Moghadam, H. K., Ferguson, M. M. & Danzmann, R. G. Organization of Hox clusters in rainbow trout (Oncorhynchus mykiss): a tetraploid model species. J. Mol. Evol. 30 Sep 2005 ( 10.1007/s00239-004-0338-7).
Cook, C. E., Jimé nez, E., Akam, M. & Saló, E. The Hox gene complement of acoel flatworms, a basal bilaterian clade. Evol. Dev. 6, 154–163 (2004).
Baguñà, J. & Riutort, M. The dawn of bilaterian animals. The case of acoelomorph flatworms. Bioessays 26, 1046–1057 (2004).
Finnerty, J. R., Pang, K., Burton, P., Paulson, D. & Martindale, M. Q. Origins of bilateral symmetry: Hox and dpp expression in a sea anemone. Science 304, 1335–1337 (2004).
Ferrier, D. E. K. & Holland, P. W. H. Sipunculan ParaHox genes. Evol. Dev. 3, 263–270 (2001).
Ferrier. D. E. K. & Holland, P. W. H. Ciona intestinalis ParaHox genes: evolution of Hox/ParaHox cluster integrity, developmental mode, and temporal colinearity. Mol. Phylogenet. Evol. 24, 412–417 (2001).
Edvardsen, R. B. et al. Remodelling of the homeobox gene complement in the tunicate Oikopleura dioica. Curr. Biol. 15, R12–R13 (2005).
Finnerty, J. R. & Martindale, M. Q. Ancient origins of axial patterning genes: Hox genes and ParaHox genes in the Cnidaria. Evol. Dev. 1, 16–23 (1999).
Ferrier, D. E. K. & Holland, P. W. H. Ancient origin of the Hox gene cluster. Nature Rev. Genet. 2, 33–38 (2001).
Dush. M. K. & Martin, G. R. Analysis of mouse Evx genes: Evx-1 displays graded expression in the primitive streak. Dev. Biol. 151, 273–287 (1992).
Miller, D. J. & Miles, A. Homeobox genes and the zootype. Nature 365, 215–216 (1993).
Banerjee-Basu, S. & Baxevanis, A. D. Molecular evolution of the homeodomain family of transcription factors. Nucleic Acids Res. 29, 3258–3269 (2001).
Minguillón. C. & Garcia-Fernàndez, J. Genesis and evolution of the Evx and Mox genes and the extended Hox and ParaHox gene clusters. Genome Biol. 4, R12 (2003).
Garcia-Fernàndez, J. Hox, ParaHox, ProtoHox: facts and guesses. Heredity 94, 145–152 (2005).
Seo, H. C. et al. (2004). Hox cluster disintegration with persistent anteroposterior order of expression in Oikopleura dioica. Nature 431, 67–71 (2004). This paper describes the Hox complement of the Urochordate Oikopleura dioica , an animal that has a compact genome, and shows that the Hox cluster is dispersed across the genome. Although no Hox gene is linked to any other (in that they are at least 250 kb apart), there is still evidence for some residual spatial collinearity of expression.
Matsui, T., Hirai, M., Hirano, M. & Kurosawa, Y. The HOX complex neighbored by EVX gene, as well as two other homoebox-containing genes, the GBX-class and the EN-class, are located on the same chormosomes 2 and 7. FEBS Lett. 336, 107–110 (1993).
Pollard, S. & Holland, P. W. H. Evidence for 14 homeobox gene clusters in human genome ancestry. Curr. Biol. 10, 1059–1062 (2000).
Castro, L. F. & Holland, P. W. H. Chromosomal mapping of ANTP class homeobox genes in amphioxus: piecing together ancestral genomes. Evol. Dev. 5, 459–465 (2003). The authors analyse the linkage of Hox, ParaHox and NK genes in the cephalochordate amphioxus, and confirm that the pre-duplicative amphioxus genome contains the three main blocks of genes, which are similar in organization to the ancestral chordate condition that has been deduced from the human genome.
Kim, Y. & Nirenberg, M. Drosophila NK-homeobox genes. Proc. Natl Acad. Sci. USA 86, 7716–7720 (1989).
Jagla, K., et al. ladybird, a tandem of homeobox genes that maintain late wingless expression in terminal and dorsal epidermis of the Drosophila embryo. Development 124, 91–100 (1997).
Luke, G. N. et al. Dispersal of NK homeobox gene clusters in amphioxus and humans. Proc. Natl Acad. Sci. USA 100, 5292–5295 (2003). This study shows that the composition of the NK gene cluster in the pre-duplicative amphioxus genome is broken at the same positions as in the vertebrate clusters. The authors conclude that the constraints to maintain linkage in chordates have been relaxed, whereas a strong functional constraint must be present in D. melanogaster.
Gauchat, D. et al. Evolution of Antp-class genes and differential expression of Hydra Hox/ParaHox genes in anterior patterning. Proc. Natl Acad. Sci. USA 97, 4493–4498 (2000).
Manuel, M. & LeParco, Y. Homeobox gene diversification in the calcareous sponge Sycon raphanus. Mol. Phyl. Evol. 17, 97–107 (2000).
Coutinho, C., Fonseca, R. N., Mansure, J. J. & Borojevic, R. Early steps in the evolution of multicellularity: deep structural and functional homologies among homeobox genes in sponges and higher metazoans. Mech. Dev. 120, 429–440 (2003).
Hill, A., Tetrault, J. & Hill, M. Isolation and expression analysis of a poriferan Antp-class Bar-/Bsh-like homeobox gene. Dev. Genes. Evol. 214, 515–523 (2004). This article includes a recent update on and the phylogenetic analyses of all known ANTP-class homeobox genes in sponges.
Lowe, C. J. et al. Anteroposterior patterning in hemichordates and the origins of the chordate nervous system. Cell 113, 853–865 (2003).
Freund, J. N, Domon-Dell, C., Kedinger, M. & Duluc, I. The Cdx-1 and Cdx-2 homeobox genes in the intestine. Biochem. Cell Biol. 76, 957–969 (1998).
Moreno, E. & Morata, G. Caudal is the Hox gene that specifies the most posterior Drosophila segment. Nature 26, 873–877 (1999).
Offield, M. F. et al. PDX-1 is required for pancreatic outgrowth and differentiation of the rostral duodenum. Development 122, 983–995 (1996).
Wysocka-Diller, J., Aisemberg, G. O. & Macagmo, E. R. A novel homeobox cluster expressed in repeated structures of the midgut. Dev. Biol. 171, 439–447 (1995).
Weiss, J. B. et al. Dorsoventral patterning in the Drosophila central nervous system: the intermediate neuroblasts defective homeobox gene specifies intermediate column identity. Genes Dev. 12, 3591–3602 (1998)
Holland, P. W. H. Beyond the Hox: how widespread is homeobox gene clustering? J. Anat. 199, 13–23 (2001).
Rosanas-Urgell, A., Marfany, G. & Garcia-Fernàndez, J. Pdx1-related homeodomain transcription factors are distinctly expressed in adult pancreatic islets. Mol. Cell. Endocrinol. 237, 59–66 (2005).
Grapin-Botton, A. & Melton, D. A. Endoderm development: from patterning to organogenesis. Trends Genet. 16, 124–130 (2000).
Schafer, K., Neuhaus, P., Kruse, J. & Braun, T. The homeobox gene Lbx1 specifies a subpopulation of cardiac neural crest necessary for normal heart development. Circ. Res. 10, 73–80 (2003).
Reecy, J. M. et al. Chicken Nkx-2.8: a novel homeobox gene expressed in early heart progenitor cells and pharyngeal pouch-2 and-3 endoderm. Dev. Biol. 188, 295–311 (1997).
Pera, E. M. & Kessel, M. Demarcation of ventral territories by the homeobox gene NKX2.1 during early chick development. Dev. Genes Evol. 208, 168–171 (1998).
Martindale. M. Q., Pang, K. & Finnerty. J. R. Investigating the origins of triploblasty: 'mesodermal' expression in a diplobastic animal, the sea anemone Nematostella vectensis (phylum, Cnidaria; class, Anthozoa). Development 131, 2463–2474 (2004). Based on expression data from anthozoan embryos, the authors argue in favour of a primitively diploblastic origin of cnidarians, and discuss the possibility that cnidarians are simplified triploblasts.
Seipel, K. & Schmid, V. Evolution of striated muscle: jellyfish and the origin of triploblasty. Dev. Biol. 282, 14–26 (2005). Based on data from jellyfish the authors argue in favour of a triplobastic origin of cnidarians.
Stephenson, T. A. The British Sea Anemones Vol. 1 (Ray Society, London, 1928).
Hyman, L. H. The Invertebrates. Protozoa Through Ctenophra (McGraw-Hill, New York (1940).
Hayward, D. C. et al. Localized expression of a dpp/BMP2/4 ortholog in a coral embryo. Proc. Natl Acad. Sci. USA 99, 8106–8111 (2002).
Finnerty, J. R. The origins of axial patterning in the metazoa: how old is bilateral symmetry? Int. J. Dev. Biol. 47, 523–529 (2003).
Muller, W. E. et al. Bauplan of urmetazoa: basis for genetic complexity of metazoa. Int. Rev. Cytol. 235, 53–92 (2004).
Maldonado, M. Choanoflagellates, choanocytes, and animal multicellularity. Inv. Biol. 123, 1–22 (2004). This data-rich paper summarizes the embryology of sponges. The author provocatively suggests that higher metazoans are derived from the complex larval stages of sponges, and that some sort of bilaterality is already present in sponge embryogenesis.
Jakob, W. et al. The Trox-2 Hox/ParaHox gene of Trichoplax (Placozoa) marks an epithelial boundary. Dev. Genes Evol. 214, 170–175 (2004).
King, N., Hittinger, C. T. & Carroll, S. B. Evolution of key cell signaling and adhesion protein families predates animal origins. Science 301, 361–363 (2003).
Brooke, M. N. & Holland, P. W. H. The evolution of multicellularity and early animal genomes. Curr. Opin. Genet. Dev. 13, 599–603 (2003).
Finnerty, J. R., Paulson, D., Burton, P., Pang, K. & Martindale, M. Q. Early evolution of a homeobox gene: the parahox gene Gsx in the Cnidaria and the Bilateria. Evol. Dev. 4, 331–345 (2003).
Oliver, B. & Mistelli, T. A non-random walk through the genome. Genome Biol. 6, 214 (2005).
Lewis, E. B. A gene complex controlling segmentation in Drosophila. Nature 276, 565–570 (1978).
Spitz, F., Gonzalez, F. & Duboule, D. A global control region defines a chromosomal regulatory landscape containing the HoxD cluster. Cell 113, 405–417 (2003).
Zákany, J., Kmita, M. & Duboule, D. A dual role for Hox genes in limb anterior-posterior assymetry. Science 304, 1669–1672 (2004). Elegantly engineered mice led Duboule's team to identify a further global regulator of the Hox cluster that is involved in the embryonic development of the limb, and to propose that there is more than one class of collinearity.
Chambeyron, S. & Bickmore, W. A. Chromatin decondensation and nuclear reorganization of the HoxB locus upon induction of transcription. Genes Dev. 10, 1119–1130 (2004).
Chambeyron, S., Da Silva, N. R., Lawson, K. A. & Bikmore, W. A. Nuclear re-organisation of the HoxB complex during mouse embryonic development. Development 132, 2215–2223 (2005).
Ikuta, T., Yoshida, N., Satoh, N. & Saiga, H. Ciona intestinalis Hox gene cluster: its dispersed structure and residual colinear expression in development. Proc. Natl Acad. Sci. USA 101, 15118–15123 (2004).
Arenas-Mena, C., Cameron, A. R. & Davidson, E. H. Spatial expression of the Hox complex in the indirect development of a sea urchin. Proc. Natl Acad. Sci. USA 95, 13062–13067 (1998).
Cameron, R. A. et al. Unusual gene order and organization of the sea urchin Hox cluster. J. Exp. Zool. Part B 22 Aug 2005 (10.1002/jez.b.21070).
Lewis, E. B., Pfeiffer, B. D., Mathog, D. R. & Celniker, S. E. Evolution of the homeobox complex in the Diptera. Curr. Biol. 13, R587–R588 (2003).
Negre, B., et al. Conservation of regulatory sequences and gene expression patterns in the disintegrating Drosophila Hox gene complex. Genome Res. 15, 692–700 (2005). The breakage point in distinct Drosophila species indicates that the Hox cluster in insects is disintegrating by phylogenetic inertia, as selective pressure for temporal collinearity is absent. The comparision of full Hox cluster sequences indicates that the breakages do not disrupt individual enhancers, which are generally located at the 5′ end of each Hox gene.
Yasukochi, Y. et al. Organization of the Hox gene cluster of the silkworm, Bombyx mori: a split of the Hox cluster in a non-Drosophila insect. Dev. Genes Evol. 21, 606–614 (2004).
Aboobaker, A. A. & Blaxter, M. L. Hox gene loss during dynamic evolution of the nematode cluster. Curr. Biol. 13, 37–40 (2003).
Spitz, F. & Duboule, D. Reproduction in clusters. Nature 434, 715–716 (2005).
MacLean, J. A., et al. Rhox: a new homeobox gene cluster. Cell 120, 369–382 (2005).
Gilbert S. F. Opening Darwin's black box: teaching evolution through developmental genetics: Nature Rev. Genet. 4, 735–741 (2003).
Holland, N. D. Early cental nervous system evolution: an era of skin brains? Nature Rev. Neurosci. 4, 617–627 (2003).
Luke, G. N. The Amphioxus NK Cluster. Thesis, Univ. Reading, UK (2004).
Acknowledgements
I am very grateful to P. Holland, J. Baguñà, C. Minguillón and the members of the Barcelona Evo-Devo group for passionate discussions. I especially thank G. Luke for allowing me to use and cite his Ph.D. Thesis, and R. Rycroft for checking the English. The author's research is funded by the Ministerio de Educación y Ciencia, Spain, by the Departament d'Universitats, Recerca i Societat de la Informació de la Generalitat de Catalunya (Distinció per la Promoció de la Recerca Universitaria), and by the European Community's 'Neurogenome' Human Potential Programme.
Author information
Authors and Affiliations
Ethics declarations
Competing interests
The author declares no competing financial interests.
Glossary
- BILATERIANS
-
Members of the animal kingdom that have bilateral symmetry — the property of having two similar sides, with definite upper and lower surfaces, and anterior and posterior ends. They include acoelomorphs, protostomes and deuterostomes.
- CNIDARIANS
-
Radially symmetrical animals that have sac-like bodies with only one opening. They include jellyfish, corals, hydra and anemones.
- METAZOAN
-
A multicellular animal.
- MACRO EVOLUTIONARY EVENTS
-
Evolutionary processess that occur above the species level; for example, the origin of phyla and changes in body-plan organization.
- THROUGH-GUT
-
A gut that has two openings: a mouth and an anus.
- TRIPLOBLASTIC
-
An animal that has three primary germ layers: the ectoderm, endoderm and mesoderm. Triploblasts include acoelomorphs, protostomes and deuterostomes.
- CHORDATES
-
The members of a phylum of animals (the Chordata) that is characterized by the possession of a notochord. It includes urochordates (such as the ascidians), cephalochordates (amphioxus) and vertebrates.
- PROTOSTOMES
-
One of the main groups of bilaterally symmetrical animals. The name derives from 'proto' (first) and 'stome' (mouth), because the first opening of the embryo (the blastopore) becomes the definitive mouth.
- DEUTEROSTOMES
-
The second of the two main groups of bilaterally symmetrical animals. The name derives from 'deutero' (second) and 'stome' (mouth), which refers to the origin of the definitive mouth as an opening that is independent from the blastopore of the embryo.
- AMPHIOXUS
-
The invertebrate that is most closely related to vertebrates.
- FLUORESCENCE IN SITU HYBRIDIZATION
-
A technique in which a fluorescently labelled DNA probe is used to detect a particular chromosome or gene with the help of fluorescence microscopy.
- DIPLOBLASTIC
-
An animal that has only two primary germ layers: the ectoderm and the endoderm. Diploblasts include the cnidarians, the ctenophores and — according to some authors — the placozoans and the poriferans.
- CAMBRIAN EXPLOSION
-
The sudden appearance, about 520 million years ago, in the fossil record of many major groups (phyla) of bilaterian animals.
- HEMICHORDATES
-
A phylum of deuterostome marine worms that comprise the enteropneust and pterobranchs.
- PLANULA BLASTOPORE
-
The opening of the planuloid larva. A planula is the swimming larva of cnidarians. Its ciliated epidermis covers either a solid or a hollowed-out mass of endoderm cells. The blastopore will develop into the oral opening of the polyp.
- NEOTENOUS
-
The term neoteny describes the retention of juvenile characteristics in the adult of an animal species.
- PLACOZOANS
-
Simple balloon-like marine animals with a body cavity that is filled with pressurized fluid. They lack most organs and tissues, including a nervous system, They are either the simplest metazoans, or simplified forms of more complex animals. Only a single species (Trichoplax adhaerens) comprises the phylum Placozoa.
- CHOANOCYTES
-
Flagellated cells that line the body cavity of a sponge and that are characterized by a collar of cytoplasm surrounding the flagellum. They are also called collar cells.
Rights and permissions
About this article
Cite this article
Garcia-Fernàndez, J. The genesis and evolution of homeobox gene clusters. Nat Rev Genet 6, 881–892 (2005). https://doi.org/10.1038/nrg1723
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrg1723
This article is cited by
-
The compact genome of the sponge Oopsacas minuta (Hexactinellida) is lacking key metazoan core genes
BMC Biology (2023)
-
Identification of HOXB9 to predict prognosis of endometrial cancer based on comprehensive bioinformatics analysis
European Journal of Medical Research (2023)
-
Homeobox B9 Promotes Colon Cancer Progression by Targeting SRSF3
Digestive Diseases and Sciences (2023)
-
Phylogenetic analysis of forkhead transcription factors in the Panarthropoda
Development Genes and Evolution (2022)
-
Homeobox genes and the specification of neuronal identity
Nature Reviews Neuroscience (2021)