Key Points
-
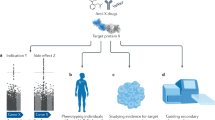
The use of genetic analyses to predict efficacy and safety in drug discovery and development is becoming a pipeline technology. A framework for assisting in key decisions at crucial points in the pharmaceutical pipeline, using prospective pipeline pharmacogenetics, is described.
-
Although genetic and genomic technologies have been applied to target discovery, applications of prospective efficacy pharmacogenetics at the crucial proof-of-concept Phase IIA can assist in decision-making with regard to progressing an asset and can reduce attrition.
-
Prospective confirmation in larger Phase-IIB studies of efficacy predictors that were identified in Phase IIA can assist in the design of larger registration trials, potentially making clinical trial studies smaller, faster and less expensive.
-
Safety pharmacogenetics can allow the rapid identification of potential toxicity-linked human genetic profiles; an example of data obtained during the course of Phase-III clinical trials and retrospective examples are described.
-
A clear distinction needs to be made between prospective pipeline data, which can decrease attrition and diagnostic profiles, which are usually defined in retrospective studies and confirmed before clinical use.
-
Strategies for practical, post-marketing risk-management surveillance are proposed, making use of rapid high-density SNP-genotyping profiles in relatively small numbers of patients who experience adverse events.
Abstract
Pharmacogenetics provides opportunities for informed decision-making along the pharmaceutical pipeline. There is a growing literature of retrospective studies of marketed medicines that describe efficacy or safety on the basis of patient genotypes. These studies emphasize the potential prospective use of genome information to enhance success in finding new medicines. An example of a prospective efficacy pharmacogenetic Phase-IIA proof-of-concept study is described. Inserting a rapidly performed efficacy pharmacogenetic step after initial clinical data are obtained can provide confidence for a commitment to full drug development. The rapid identification of adverse events during and after drug development using genomic mapping tools is also reviewed.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Cardon, L. R. & Bell, J. I. Association study designs for complex diseases. Nature Rev. Genet. 2, 91–99 (2001). An informative review of the application of genetic association methods used to identify genes associated with complex diseases.
Tabor, H. K., Risch, N. J. & Myers, R. M. Candidate-gene approaches for studying complex genetic traits: practical considerations. Nature Rev. Genet. 3, 391–397 (2002).
Weiss, K. M. & Clark, A. G. Linkage disequilibrium and the mapping of complex human traits. Trends Genet. 18, 19–24 (2002). An overview of the applications of LD mapping for diseases, but equally applicable to safety (adverse-event) PGx.
Goldstein, D. B., Tate, S. K. & Sisodiya, S. M. Pharmacogenetics goes genomic. Nature Rev. Genet. 4, 937–947 (2003).
Carlson, C. S. et al. Selecting a maximally informative set of single-nucleotide polymorphisms for association analyses using linkage disequilibrium. Am. J. Hum. Genet. 74, 106–120 (2004).
Dawson, E. et al. A first-generation linkage disequilibrium map of human chromosome 22. Nature 418, 544–548 (2002).
Lai, E. et al. Medical applications of haplotype-based SNP maps: learning to walk before we run. Nature Genet. 32, 353 (2002). Comment on the difference between a strategy of whole-genome mapping, limited to a full panel of small, well-defined LD blocks, versus the use of an initial screening set to determine regions of extended LD for fine mapping (for examples see references 10, 16 and 47).
Roses, A. D. Pharmacogenetics. Hum. Mol. Genet. 10, 2261–2267 (2001).
Roses, A. D. Genome-based pharmacogenetics and the pharmaceutical industry. Nature Rev. Drug Discov. 1, 541–554 (2002).
Xu, C. F. et al. Identification of a pharmacogenetic effect by linkage disequilibrium mapping. Pharmacogenomics J. (in the press). A proof of principle analysis of the application of whole-genome LD to map a locus (loci) associated with an adverse effect of a medicine.
Food and Drug Administration. Innovation or Stagnation: Challenge and Opportunity on the Critical Path to New Medical Products [online], <http://www.fda.gov/oc/initiatives/criticalpath/whitepaper.html> (2004).
Lindsay, M. A. Target discovery. Nature Rev. Drug Discov. 2, 831–838 (2003).
Chanda, S. K. & Caldwell, J. S. Fulfilling the promise: drug discovery in the post-genomic era. Drug Discov. Today 8, 168–174 (2003).
Collins, F. S. Genetics: an explosion of knowledge is transforming clinical practice. Geriatrics 54, 410–417 (1999).
Subramanian, G., Adams, M. D., Venter, J. C. & Broder, S. Implications of the human genome for understanding human biology and medicine. JAMA 286, 2296–2307 (2001).
McCarthy, L. C. et al. Single-nucleotide polymorphism alleles in the insulin receptor gene are associated with typical migraine. Genomics 78, 135–149 (2001). An early example of the use of high density SNP mapping in a region of putative genetic linkage, in this case for the identification of specific SNP variants of the insulin receptor gene that are associated with migraine.
Hakonarson, H., Gulcher, J. R. & Stefansson, K. deCODE genetics, Inc. Pharmacogenomics 4, 209–215 (2003).
Kristjansson, K. et al. Linkage of essential hypertension to chromosome 18q. Hypertension 39, 1044–1049 (2002).
Nathan, D. G. Clinical research: perceptions, reality, and proposed solutions. National Institutes of Health Director's Panel on Clinical Research. JAMA 280, 1427–1431 (1998).
Boston Consulting Group. A Revolution in R&D: How Genomics and Genetics Will Affect Drug Development Costs and Tmes in Parexel's Pharmaceutical R&D Statistical Sourcebook, 2003/2003 (Parexel International, Waltham, USA, 2003).
Harris, S. & Foord, S. M. Transgenic gene knock-outs: functional genomics and therapeutic target selection. Pharmacogenomics 1, 433–443 (2000).
Brenner, S. E. Target selection for structural genomics. Nature Struct. Biol. 7 (Suppl.), 967–969 (2000).
DeFife, K. M. & Wong-Staal, F. Integrated approaches to therapeutic target gene discovery. Curr. Opin. Drug Discov. Dev. 5, 683–689 (2002).
Searls, D. B. Pharmacophylogenomics: genes, evolution and drug targets. Nature Rev. Drug Discov. 2, 613–623 (2003). An analysis of the importance of placing genomic variation in an evolutionary context, taking advantage of phylogenetic relationships and comparisons, to better understand function and in particular selective pressures.
Debouck, C. & Metcalf, B. The impact of genomics on drug discovery. Annu. Rev. Pharmacol. Toxicol. 40, 193–207 (2000).
Drews, J. Drug discovery: a historical perspective. Science 287, 1960–1964 (2000).
Thorisson, G. A. & Stein L. D. The SNP Consortium website: past, present and future. Nucleic Acids Res. 31, 124–127 (2003).
Vaschetto, M., Weissbrod, T., Bodle, D. & Guner, O. Enabling high-throughput discovery. Curr. Opin. Drug Discov. Dev. 6, 377–383 (2003).
Duyk, J. Attrition and translation. Science 302, 603–605 (2003).
Twyman, R. M. & Primrose, S. B. Techniques patents for SNP genotyping. Pharmacogenomics 4, 67–79 (2003).
Renegar, G., Rieser, P. & Manasco, P. Family consent and the pursuit of better medicines through genetics research. J. Contin. Educ. Health Prof. 21, 265–270 (2001).
Horrobin, D. F. Modern biomedical research: an internally self-consistent universe with little contact with medical reality? Nature Rev. Drug Discov. 2, 151–154 (2003).
Roses, A. D. High through-put disease-associated target identification. Drug Discov. Today (in the press). Description of high throughput multi-gene target testing to determine variants that are highly associated with human disease.
Lynch, T. J. et al. Activating mutations in the epidermal growth factor receptor underlying responsiveness of non-small-cell lung cancer to gefitinib. N. Engl. J. Med. 350, 2129–2139 (2004).
Paez, J. G. et al. EGFR mutations in lung cancer: correlation with clinical response to gefitinib therapy. Science 304, 1497–1500 (2004).
Dos Santos, C. et al. A common polymorphism of the growth hormone receptor is associated with increased responsiveness to growth hormone. Nature Genet. 36, 720–724 (2004).
Tantisira, K. G. et al. Corticosteroid pharmacogenetics: association of sequence variants in CRHR1 with improved lung function in asthmatics treated with inhaled corticosteroids. Hum. Mol. Genet. 13, 1353–1359 (2004).
Sesti, G. et al. The Arg972 variant in insulin receptor substrate-1 is associated with an increased risk of secondary failure to sulfonylurea in patients with type 2 diabetes. Diabetes Care 27, 1394–1398 (2004).
Macedo, A., Farre, M. & Banos, J. E. Placebo effect and placebos: what are we talking about? Some conceptual and historical considerations. Eur. J. Clin. Pharmacol. 59, 337–342 (2003).
Goetz, C. G., Janko, K., Blasucci, L. & Jaglin, J. A. Impact of placebo assignment in clinical trials of Parkinson's disease. Mov. Disord. 18, 1146–1149 (2003).
Anonymous. The Fruits of Genomics: Drug Pipelines Face Indigestion Until the New Biology Ripens. (Lehman Brothers, McKinsey and Co., New York, 2001).
Tufts Center for the Study of Drug Development. Backgrounder: How New Drugs Move Through the Development and Approval Process <http://csdd.tufts.edu/NewsEvents?RecentNews.asp?newsid=4> (2001).
Gilbert, J., Henkse, P. & Singh, A. Rebuilding big pharma's business model. In vivo, the business and medicine report. Windhover Information, 21, No. 10, (2003).
Noble, M. E., Endicott, J. A. & Johnson, L. N. Protein kinase inhibitors: insights into drug design from structure. Science 303, 1800–1805 (2004).
Vogel, C. L. & Franco S. X. Clinical experience with trastuzumab (herceptin). Breast J. 9, 452–462 (2003).
Goldstein, D. B., Ahmadi, K. R., Weale, M. E. & Wood N. W. Genome scans and candidate gene approaches in the study of common diseases and variable drug responses. Trends Genet. 19, 615–622 (2003).
Lai, E., Riley, J., Purvis, I. & Roses, A. A 4-Mb high-density single nucleotide polymorphism-based map around human APOE. Genomics 54, 31–38 (1998). An early example of LD mapping of a region of genetic linkage to demonstrate the polymorphisms of SLC12A8, a gene coding for a member of the solute carrier family 12 proteins that are associated with familial psoriasis.
Hewett, D. et al. Identification of a psoriasis susceptibility candidate gene by linkage disequilibrium mapping with a localized single nucleotide polymorphism map. Genomics 79, 305–314 (2002).
deCODE Genetics. deCODE and Merck & Co., Inc. form broad drug development alliance. deCODE News [online], <http://www.decode.com/main/view.jsp?branch=5026&e342RecordID=1901&e342DataStoreID=3917> (2004).
Merck Press Release. Drug development; pharmaceutical company joins drug development alliance. Genomics & Genetics Weekly 38, (2004).
Melzer, D., Detmer, D. & Simmern, R. Pharmacogenetics and public policy: expert views in Europe and North America. Pharmacogenomics 4, 689–691 (2003).
Lee, S. S. Race, distributive justice and the promise of pharmacogenomics: ethical considerations. Am. J. Pharmacogenomics 3, 385–392 (2003).
Hapgood, R. The potential and limitations of personalized medicine in the doctor-patient relationship. Pharmacogenomics 4, 685–687 (2003).
Drazen, J. M. et al. Pharmacogenetic association between ALOEX5 promoter genotype and the response to anti-asthma treatment. Nature Genet. 22, 168–170 (2003).
Wechsler, M. E. & Israel, E. Pharmacogenetics of treatment with leukotriene modifiers. Curr. Opin. Allergy Clin. Immunol. 2, 395–401 (2002).
Spear, B. B., Heath-Chiozzi, M. & Huff, J. Clinical application of pharmacogenetics. Trends Mol. Med. 7, 201–204 (2001).
Roses, A. D. 2025: the practice of neurology: back from the future. Arch. Neurol. 58, 1766–1767 (2001).
Ahmadi, K. R. et al. A single nucleotide polymorphism tagging set for human drug metabolism and transport. Science (in the press). A study and analysis of the use of LD mapping to determine a tagged set of SNPs for 56 human drug metabolizing enzymes.
Griffiths, J. D., Stark, R. J., Ding, J. C. & Cooper, I. A. Vincristine neurotoxicity in Charcot–Marie–Tooth syndrome. Med. J. Aust. 143, 305–306 (1985).
Danoff, T. M. et al. A Gilbert's syndrome UGT1A1 variant confers susceptibility to tranilast-induced hyperbilirubinemia. Pharmacogenomics J. 4, 49–53 (2004).
Gibbs, R. A. et al. The International HapMap Project. Nature 426, 789–796 (2003).
Bowman, C. Classification using SNP profiles. <http://www.luc.ac.be/censtat/RSS2003> (2003).
Norbert, P. W. & Roses A. D. Pharmacogenetics and pharmacogenomics: recent developments, their clinical relevance and some ethical, social, and legal implications. J. Mol. Med. 81, 35–40 (2003).
Lesko, L. J. et al. Pharmacogenetics and pharmacogenomics in drug development and regulatory decision making: report of the first FDA-PWG-PhRMA-DruSafe Workshop. J. Clin. Pharmacol. 43, 342–358 (2003). A view from the FDA, the agency that regulates pharmaceutical licenses in the US, on decision making using pharmacogenetics and pharmacogenomics during drug development, including definition of key terms generically referred to as “biomakers.”
Buchanan, A. et al. Pharmacogenetics: ethical issues and policy options. Kennedy Inst. Ethics J. 12, 1–15 (2002).
Welsh-Bohmer, K. A. et al. Apolipoprotein E genotypes in a neuropathological series from the Consortium to Establish a Registry for Alzheimer's Disease. Ann. Neurol. 42, 319–325 (1997).
Acknowledgements
This research was fully supported from the sales of medicines. I especially thank my successive Presidents of Research and Development, both physician-scientists, James Niedel and Tachi Yamada, for having the foresight and commitment to support and apply strategic genetic research in 1997 when human linkage research was considered as peaked, and since 2000 when medical research investments for the future were still unfashionable for pharmaceutical R&D organizations. The ongoing pharmacogenetics work that is described in this review has been contributed to by hundreds of GlaxoSmithKline (GSK) employees over a seven year period. I also thank the Genetic Executive Teams, who have shared a vision of safer and more effective medicines, the thousands of patients who consented to participate in pharmacogenetic research protocols, Ronald Krall who provided access to all patients participating in GSK drug development, my wife, Ann Saunders, who provided intellectual, emotional and laboratory support, and my daughters — Maija, Stephanie and Joanna — who created a place to live, to work and to love.
Author information
Authors and Affiliations
Ethics declarations
Competing interests
A. D. Roses is a full-time employee of GlaxoSmithKline and currently holds non-paid appointments as a clinical professor of neurology at Duke University Medical Center, and on the Science Board of the US Food and Drug Administration.
Related links
Related links
DATABASES
Entrez
FURTHER INFORMATION
Food and Drug Administration Science Board meeting (22 April 2004)
Food and Drug Administration Science Board presentation (22 April 2004)
Glossary
- PHASE I
-
Initial clinical studies usually involve small numbers of normal human volunteers and small doses to assess safety, metabolism and excretion of a drug or drug combination.
- PHASE IIA
-
After the initial Phase-I studies, these randomized and controlled clinical studies are used to assess efficacy (proof of concept) and to continue to assess a drug or drug combination in a relatively small number of patients (tens of patients). These early studies can be non-comparative or double-blind comparisons.
- PHASE IIB
-
After Phase-IIA studies, these randomized and controlled clinical studies are used to assess the efficacy and dose-ranging of a drug or drug combination in larger groups of patients (hundreds of patients). In some designs, there is a positive interim end point so that Phase IIB can be extended into Phase-III registration trials. Safety continues to be assessed.
- PHASE III
-
Randomized and controlled clinical studies in patients (thousands of patients) designed to evaluate the comparative safety and efficacy of a drug or drug combination. Also includes the principal data used by regulatory agencies to approve or reject a product-licensing application.
- EFFICACY PHARMACOGENETICS
-
The study of genetic variation that underlies variability in the efficacy of drugs for treating disease.
- INTENT TO TREAT
-
Analysis of clinical-trial results that includes all data from patients in the groups to which they were randomized (that is, assigned through random distribution) even if they never received the treatment.
- LINKAGE DISEQUILIBRIUM
-
The condition in which the frequency of a particular haplotype for two loci is significantly different from that expected under random mating. The expected frequency is the product of observed allelic frequencies at each locus.
- TRANILAST
-
The name that was used for a specific anti-restenosis drug while it was being investigated by GlaxoSmithKline.
- HYPERBILIRUBINAEMIA
-
A high level of bilirubin in the blood. This can cause yellowing of the skin (jaundice).
- PERIPHERAL NEUROPATHY
-
A problem in peripheral nerve function (any part of the nervous system except the brain and spinal cord) that might cause pain, numbness, tingling, swelling and muscle weakness in various parts of the body. Neuropathies might be caused by physical injury, infection, toxic substances, disease (for example, cancer, diabetes, kidney failure or malnutrition) or drugs such as anti-cancer drugs.
- CHARCOT–MARIE–TOOTH NEUROPATHY
-
A genetic disease that is characterized by progressive peripheral neuropathy and debilitating muscular weakness, particularly of the limbs. There are multiple inherited mutations of several genes that result in similar phenotypes.
- ABSOLUTE D′
-
For specified alleles at two distinct loci, D′ is the absolute difference between the observed and expected haplotype frequencies, divided by the maximum value that the difference could possibly attain.
- RESTENOSIS
-
A re-narrowing or blockage of an artery at the same site at which treatment, such as an angioplasty or stent procedure, has already taken place.
- GILBERT'S DISEASE
-
A benign syndrome of increased sensitivity to external drugs that causes hyperbilirubinaemia but does not progress to severe liver impairment.
- BAYES FACTOR
-
The ratio of the posterior odds to the prior odds.
- GENETIC LOAD
-
The degree to which a given trait can be attributed to genetic variation by the proportion of cases that carry the allele.
Rights and permissions
About this article
Cite this article
Roses, A. Pharmacogenetics and drug development: the path to safer and more effective drugs. Nat Rev Genet 5, 645–656 (2004). https://doi.org/10.1038/nrg1432
Issue Date:
DOI: https://doi.org/10.1038/nrg1432
This article is cited by
-
Precision medicine for rheumatologists: lessons from the pharmacogenomics of azathioprine
Clinical Rheumatology (2021)
-
A study of hydrophobins-modified menaquinone-7 on osteoblastic cells differentiation
Molecular and Cellular Biochemistry (2021)
-
Adaptive Biomarker Population Selection in Phase III Confirmatory Trials with Time-to-Event Endpoints
Statistics in Biosciences (2018)
-
Using Genetics to Improve Addiction Treatment Outcomes
Current Behavioral Neuroscience Reports (2017)
-
Applying genome-wide gene-based expression quantitative trait locus mapping to study population ancestry and pharmacogenetics
BMC Genomics (2014)