Key Points
-
In genetic analysis, there are often competing explanations for the same data. Sophisticated mathematical models have been developed that can encapsulate these problems in terms of parameters that need to be inferred.
-
Bayesian statistical methods are well suited to help pick out the most reasonable parameter values — as well as to choose between entire models — and they provide a framework for including background information to help with this.
-
The goal of Bayesian analysis is to compute the probability distribution of parameter values and model specifications given the data. This is called the posterior distribution.
-
It is only with the development of high-speed computing over the past ten years that the potential of Bayesian methods has been realized. As a consequence, statistical analysis in genetics has undergone a dramatic shift.
-
A computational method that has had most influence is Markov chain Monte Carlo, which allows parameter values to be drawn from the posterior distribution.
-
Example areas that have used Bayesian methods include: population genetics, detecting the effects of selection, sequence analysis, SNP discovery, haplotype identification, analysis of gene expression, association mapping and linkage-disequilibrium mapping.
-
A review of applications in these areas demonstrates the following advantages of Bayesian methods over other approaches: use of background information; the ability to include uncertainty in all parameter values; ease in making inferences about some parameters irrespective of the values of others; and lack of ad hoc calculations and approximations that are often associated with alternative statistical methods.
-
There are still computational difficulties with Bayesian approaches. Further improvements are needed both in testing the accuracy of the computation involved and also in model checking.
Abstract
Bayesian statistics allow scientists to easily incorporate prior knowledge into their data analysis. Nonetheless, the sheer amount of computational power that is required for Bayesian statistical analyses has previously limited their use in genetics. These computational constraints have now largely been overcome and the underlying advantages of Bayesian approaches are putting them at the forefront of genetic data analysis in an increasing number of areas.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Shoemaker, J. S., Painter, I. S. & Weir, B. S. Bayesian statistics in genetics: a guide for the uninitiated. Trends Genet. 15, 354–358 (1999).
Gelman, A., Carlin, J. B., Stern, H. S. & Rubin, D. B. Bayesian Data Analysis (Chapman and Hall, London, 1995).
Cavalli-Sforza, L. L. & Edwards, A. W. F. Phylogenetic analysis: models and estimation procedures. Evolution 32, 550–570 (1967).
Ewens, W. J. The sampling theory of selectively neutral alleles. Theor. Popul. Biol. 3, 87–112 (1972). The first use of a sampling distribution in population genetics. This paper anticipates modern approaches, such as the coalescent theory, that model the sampling distribution of chromosomes.
Kingman, J. F. C. The coalescent. Stochastic Process. Appl. 13, 235–248 (1982).
Hudson, R. R. Properties of a neutral allele model with intragenic recombination. Theor. Popul. Biol. 23, 183–201 (1983).
Felsenstein, J. Estimating effective population size from samples of sequences: inefficiency of pairwise and segregating sites as compared to phylogenetic estimates. Genet. Res. 59, 139–147 (1992).
Griffiths, R. C. & Tavaré, S. Ancestral inference in population genetics. Statistical Sci. 9, 307–319 (1994).
Markovtsova, L., Marjoram, P. & Tavaré, S. The effect of rate variation on ancestral inference in the coalescent. Genetics 156, 1427–1436 (2000).
Tavaré, S., Balding, D. J., Griffiths, R. C. & Donnelly, P. Inferring coalescence times from DNA sequence data. Genetics 145, 505–518 (1997).
Wilson, I. J. & Balding, D. J. Genealogical inference from microsatellite data. Genetics 150, 499–510 (1998). An early paper that uses MCMC to carry out a fully Bayesian analysis of population-genetic data.
Beerli, P. & Felsenstein, J. Maximum likelihood estimation of a migration matrix and effective population sizes in n subpopulations by using a coalescent approach. Proc. Natl Acad. Sci. USA 98, 4563–4568 (2001).
Nielsen, R. & Wakeley, J. Distinguishing migration from isolation: a Markov chain Monte Carlo approach. Genetics 158, 885–896 (2001).
Wakeley, J., Nielsen, R., Liu-Cordero, S. N. & Ardlie, K. The discovery of single-nucleotide polymorphisms and inferences about human demographic history. Am. J. Hum. Genet. 69, 1332–1347 (2001).
Storz, J. F., Beaumont, M. A. & Alberts, S. C. Genetic evidence for long-term population decline in a savannah-dwelling primate: inferences from a hierarchical Bayesian model. Mol. Biol. Evol. 19, 1981–1990 (2002).
Rannala, B. & Yang, Z. Bayes estimation of species divergence times and ancestral population sizes using DNA sequences from multiple loci. Genetics 164, 1645–1656 (2003).
Marjoram, P., Molitor, J., Plagnol, V. & Tavaré, S. Markov chain Monte Carlo without likelihoods. Proc. Natl Acad. Sci. USA 100, 15324–15328 (2003).
Beaumont, M. A., Zhang, W. & Balding, D. J. Approximate Bayesian computation in population genetics. Genetics 162, 2025–2035 (2002).
Wilson, I. J., Weale, M. E. & Balding, D. J. Inferences from DNA data: population histories, evolutionary processes and forensic match probabilities. J. Roy. Stat. Soc. A Sta. 166, 155–188 (2003).
Cavalli-Sforza, L. L., Menozzi, P. & Piazza, A. The History and Geography of Human Genes (Princeton Univ. Press, Princeton, 1994).
Devlin, B. & Roeder, K. Genomic control for association studies. Biometrics 55, 997–1004 (1999).
Pritchard, J. K. & Rosenberg, N. A. Use of unlinked genetic markers to detect population stratification in association studies. Am. J. Hum. Genet. 65, 220–228 (1999).
Pritchard, J. K., Stephens, M., Rosenberg, N. A. & Donnelly, P. Association mapping in structured populations. Am. J. Hum. Genet. 67, 170–181 (2000).
Pritchard, J. K. & Donnelly, P. Case–control studies of association in structured or admixed populations. Theor. Popul. Biol. 60, 227–237 (2001).
Davies, N., Villablanca, F. X. & Roderick, G. K. Bioinvasions of the medfly Ceratitis capitata: source estimation using DNA sequences at multiple intron loci. Genetics 153, 351–360 (1999).
Bonizzoni, M. et al. Microsatellite analysis of medfly bioinfestations in California. Mol. Ecol. 10, 2515–2524 (2001).
Pritchard, J. K., Stephens, M. & Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 155, 945–959 (2000). An influential paper in the development of Bayesian methods to study cryptic population structure. The program described in it, Structure, has been widely used in molecular ecology.
Dawson, K. J. & Belkhir, K. A Bayesian approach to the identification of panmictic populations and the assignment of individuals. Genet. Res. 78, 59–77 (2001).
Wright, S. Evolution and the Genetics of Populations: The Theory of Gene Frequencies (Chicago Univ. Press, Chicago, 1969).
Corander, J., Waldmann, P. & Sillanpaa, M. J. Bayesian analysis of genetic differentiation between populations. Genetics 163, 367–374 (2003).
Wilson, G. A. & Rannala, B. Bayesian inference of recent migration rates using multilocus genotypes. Genetics 163, 1177–1191 (2003).
Bamshad, M. & Wooding, S. P. Signatures of natural selection in the human genome. Nature Rev. Genet. 4, 99–111 (2003).
Storz, J. F. & Beaumont, M. A. Testing for genetic evidence of population expansion and contraction: an empirical analysis of microsatellite DNA variation using a hierarchical Bayesian model. Evolution 56, 154–166 (2002).
Beaumont, M. A. & Balding, D. J. Identifying adaptive genetic divergence among populations from genome scans. Mol. Ecol. (in the press).
Bustamante, C. D., Nielsen, R. & Hartl, D. L. Maximum likelihood and Bayesian methods for estimating the distribution of selective effects among classes of mutations using DNA polymorphism data. Theor. Popul. Biol. 63, 91–103 (2003).
Nielsen, R. Statistical tests of selective neutrality in the age of genomics. Heredity 86, 641–647 (2001).
Nielsen, R. & Yang, Z. Likelihood models for detecting positively selected amino acid sites and applications to the HIV-1 envelope gene. Genetics 148, 929–936 (1998). The first formal statistical method for inferring site-specific selection on DNA codons.
Holder, M. & Lewis, P. O. Phylogeny estimation: traditional and Bayesian approaches. Nature Rev. Genet. 4, 275–284 (2003). Reviews the many recent applications of Bayesian inference in phylogeny estimation.
Durbin, R., Eddy, S., Krogh, A. & Mitchison, G. Biological Sequence Analysis, (Cambridge Univ. Press, Cambridge, 1998).
Lawrence, C. E. et al. Detecting subtle sequence signals: a Gibbs sampling strategy for multiple alignment. Science 262, 208–214 (1993). The methods and models used in this paper have led to the development of a large number of Bayesian methods for the analyses of sequence data by some of the authors and their groups.
Churchill, G. A. Stochastic models for heterogeneous DNA sequences. Bull. Math. Biol. 51, 79–94 (1989). One of the earliest papers to use a hidden Markov model to analyse DNA sequence data.
Borodovsky, M., McIninch, J. Genmark: parallel gene recognition for both DNA strands. Comput. Chem. 17, 123–133 (1993).
Liu, J. S., Neuwald, A. F. & Lawrence, C. E. Bayesian models for multiple local sequence alignment and Gibbs sampling strategies. J. Am. Stat. Ass. 90, 1156–1170 (1995).
Webb, B. M., Liu, J. S. & Lawrence, C. E. BALSA: Bayesian algorithm for local sequence alignment. Nucleic Acids Res. 30, 1268–1277 (2002).
Thompson, W., Rouchka, E. C., Lawrence, C. E. Gibbs recursive sampler: finding transcription factor binding sites. Nucleic Acids Res. 31, 3580–3585 (2003).
Liu, J. S. & Lawrence, C. E. Bayesian inference on biopolymer models. Bioinformatics 15, 38–52 (1999).
Liu, J. S. & Logvinenko, T. in Handbook of Statistical Genetics (eds Balding, D. J., Bishop, M. & Cannings, C.) 66–93 (John Wiley and Sons, Chichester, 2003).
Churchill, G. A. & Lazareva, B. Bayesian restoration of a hidden Markov chain with aplications to DNA sequencing. J. Comput. Biol. 6, 261–277 (1999).
Human Genome Sequencing Consortium. Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001).
Venter, J. C. et al. The sequence of the human genome. Science 291, 1304–1351 (2001).
Polanski, A. & Kimmel, M. New explicit expressions for relative frequencies of single-nucleotide polymorphisms with application to statistical inference on population growth. Genetics 165, 427–436 (2003).
Zhu, Y. L. et al. Single-nucleotide polymorphisms in soybean. Genetics 163, 1123–1134 (2003).
Marth, G. T. et al. A general approach to single-nucleotide polymorphism discovery. Nature Genet. 23, 452–456 (1999).
Irizarry, K. et al. Genome-wide analysis of single-nucleotide polymorphisms in human expressed sequences. Nature Genet. 26, 233–236 (2000).
Ott, J. Analysis of Human Genetic Linkage (Johns Hopkins, Baltimore, 1999).
Long, J. C., Williams, R. C. & Urbanek, M. An E-M algorithm and testing strategy for multiple-locus haplotypes. Am. J. Hum. Genet. 56, 799–810 (1995).
Excoffier, L. & Slatkin, M. Maximum-likelihood estimation of molecular haplotype frequencies in a diploid population. Mol. Biol. Evol. 12, 921–927 (1995).
Niu, T., Qin, Z. S., Xu, X. & Liu, J. S. Bayesian haplotype inference for multiple linked single-nucleotide polymorphisms. Am. J. Hum. Genet. 70, 157–169 (2002).
Stephens, M., Smith, N. J. & Donnelly, P. A new statistical method for haplotype reconstruction from population data. Am. J. Hum. Genet. 68, 978–989 (2001).
Dempster, A. P., Laird, N. M. & Rubin, D. B. Maximum likelihood from incomplete data via the EM algorithm. J. Roy. Statist. Soc. B39, 1–38 (1977).
Slatkin, M. & Excoffier, L. Testing for linkage disequilibrium in genotypic data using the Expectation-Maximization algorithm. Heredity 76, 377–383 (1996).
Butte, A. The use and analysis of microarray data. Nature Rev. Genet. 1, 951–960 (2002).
Huber, W., von Heydebreck, A. & Vingron, M. in Handbook of Statistical Genetics (eds Balding, D. J., Bishop, M. & Cannings, C.) 162–187 (John Wiley and Sons, Chichester, 2003).
Baldi, P. & Long, A. D. A Bayesian framework for the analysis of microarray expression data: regularized t-test and statistical inferences of gene changes. Bioinformatics 17, 509–519 (2001).
Storey, J. D. & Tibshirani, R. Statistical significance for genomewide studies. Proc. Natl Acad. Sci. USA 100, 9440–9445 (2003).
Ibrahim, J. G., Chen, M. H. & Gray, R. J. Bayesian models for gene expression with DNA microarray data. J. Am. Stat. Ass. 97, 88–99 (2002).
Ishwaran, H. & Rao, J. S. Detecting differentially expressed genes in microarrays using Bayesian model selection. J. Am. Stat. Ass. 98, 438–455 (2003).
Lee, K. E., Sha, N., Dougherty, E. R., Vannucci, M. & Mallick, B. K. Gene selection: a Bayesian variable selection approach. Bioinformatics 19, 90–97 (2003).
Zhang, M. Q. Large-scale gene expression data analysis: a new challenge to computational biologists. Genome Res. 9, 681–688 (2003).
Heard, N. A., Holmes, C. C. & Stephens, D. A. A quantitative study of gene regulation involved in the immune response of anopheline mosquitoes: an application of Bayesian hierarchical clustering of curves. Department of Statistics, Imperial College, London [online], <http://stats.ma.ic.ac.uk/~ccholmes/malaria_clustering.pdf> (2003).
Dove, A. Mapping project moves forward despite controversy. Nature Med. 12, 1337 (2002).
Rannala, B. Finding genes influencing susceptibility to complex diseases in the post-genome era. Am. J. Pharmacogenomics 1, 203–221 (2001).
Sham, P. Statistics in Human Genetics, (Oxford Univ. Press, New York, 1998).
Jorde, L. B. Linkage disequilibrium and the search for complex disease genes. Genome Res. 10, 1435–1444 (2000).
Spielman, R. S., McGinnis, R. E. & Ewens, W. J. Transmission test for linkage disequilibrium: the insulin gene region and insulin-dependent diabetes mellitus (IDDM). Am. J. Hum. Genet. 52, 506–516 (1993). The first application of a family-based association test. The transmission disequilibrium test has been highly influential and spawned many related approaches.
Denham, M. C. & Whittaker, J. C. A Bayesian approach to disease gene location using allelic association. Biostatistics 4, 399–409 (2003).
Sham, P. C. & Curtis, D. An extended transmission/disequilibrium test (TDT) for multi-allele marker loci. Ann. Hum. Genet. 59, 323–336 (1995).
Paetkau, D., Calvert, W., Stirling, I. & Strobeck, C. Microsatellite analysis of population-structure in Canadian polar bears. Mol. Ecol. 4, 347–354 (1995).
Rannala, B. & Mountain, J. L. Detecting immigration by using multilocus genotypes. Proc. Natl Acad. Sci. USA 94, 9197–9201 (1997).
Sillanpaa, M. J., Kilpikari, R., Ripatti, S., Onkamo, P. & Uimari, P. Bayesian association mapping for quantitative traits in a mixture of two populations. Genet. Epidemiol. 21 (Suppl. 1), S692–S699 (2001).
Hoggart, C. J. et al. Control of confounding of genetic associations in stratified populations. Am. J. Hum. Genet. 72, 1492–1504 (2003).
Bodmer, W. F. Human genetics: the molecular challenge. Cold Spring Harb. Symp. Quant. Biol. 51, 1–13 (1986).
Lander, E. S. & Botstein, D. Mapping complex genetic traits in humans: new methods using a complete RFLP linkage map. Cold Spring Harb. Symp. Quant. Biol. 51, 49–62 (1986).
Dean, M. et al. Approaches to localizing disease genes as applied to cystic fibrosis. Nucleic Acids Res. 18, 345–350 (1990).
Hastbacka, J. et al. Linkage disequilibrium mapping in isolated founder populations: diastrophic dysplasia in Finland. Nature Genet. 2, 204–211 (1992).
Rannala, B. & Slatkin, M. Methods for multipoint disease mapping using linkage disequilibrium. Genet. Epidemiol. 19 (Suppl. 1), S71–S77 (2000). A comprehensive review of the various likelihood approximations used in linkage-disequilibrium gene mapping.
Rannala, B. & Reeve, J. P. High-resolution multipoint linkage-disequilibrium mapping in the context of a human genome sequence. Am. J. Hum. Genet. 69, 159–178 (2001). The first use of the human genome sequence as an informative prior for Bayesian gene mapping.
Morris, A. P., Whittaker, J. C. & Balding, D. J. Fine-scale mapping of disease loci via shattered coalescent modeling of genealogies. Am. J. Hum. Genet. 70, 686–707 (2002).
Rannala, B. & Reeve, J. P. Joint Bayesian estimation of mutation location and age using linkage disequilibrium. Pac. Symp. Biocomput. 526–534 (2003).
Reeve, J. P. & Rannala, B. DMLE+: Bayesian linkage disequilibrium gene mapping. Bioinformatics 18, 894–895 (2002).
Liu, J. S., Sabatti, C., Teng, J., Keats, B. J. & Risch, N. Bayesian analysis of haplotypes for linkage disequilibrium mapping. Genome Res. 11, 1716–1724 (2001).
Liu, J. S. Monte Carlo Methods for Scientific Computing (Springer, New York, 2001).
Pavlovic, V., Garg, A. & Kasif, S. A Bayesian framework for combining gene predictions. Bioinformatics 18, 19–27 (2002).
Jansen, R. et al. A Bayesian networks approach for predicting protein–protein interactions from genomic data. Science 302, 449–453 (2003).
Ross, S. M. Simulation, (Academic, New York, 1997).
Ripley, B. D. Stochastic Simulation (Wiley and Sons, New York, 1987).
Hudson, R. R. Gene genealogies and the coalescent process. Oxford Surveys Evol. Biol. 7, 1–44 (1990).
Metropolis, N. Rosenbluth, A. N., Rosenbluth, M. N., Teller, A. H. & Teller, E. Equations of state calculations by fast computing machine. J. Chem. Phys. 21, 1087–1091 (1953).
Hastings, W. K. Monte Carlo sampling methods using Markov chains and their application. Biometrika 57, 97–109 (1970).
Pritchard, J. K., Seielstad, M. T., Perez-Lezaun, A. & Feldman, M. W. Population growth of human Y chromosomes: a study of Y chromosome microsatellites. Mol. Biol. Evol. 116, 1791–1798 (1999). The first paper to use an ABC approach to infer population-genetic parameters in a complicated demographic model.
Beaumont, M. A. Detecting population expansion and decline using microsatellites. Genetics 153, 2013–2029 (1999).
Drummond, A. J., Nicholls, G. K., Rodrigo, A. G. & Solomon, W. Estimating mutation parameters, population history and genealogy simultaneously from temporally spaced sequence data. Genetics 161, 1307–1320 (2002).
Pybus, O. G., Drummond, A. J., Nakano, T., Robertson, B. H. & Rambaut, A. The epidemiology and iatrogenic transmission of hepatitis C virus in Egypt: a Bayesian coalescent approach. Mol. Biol. Evol. 20, 381–387 (2003).
Beaumont, M. A. Estimation of population growth or decline in genetically monitored populations. Genetics 164, 1139–1160 (2003).
Elston, R. C. & Stewart, J. A general model for the analysis of pedigree data. Human Heredity 21, 523–542 (1971).
Lander, E. S. & Green, P. Construction of multilocus genetic linkage maps in humans. Proc. Natl Acad. Sci. USA 84, 2362–2367 (1987).
Krugylak, L., Daly, M. J. & Lander, E. S. Rapid multipoint linkage analysis of recessive traits in nuclear families, including homozygosity mapping. Am. J. Hum. Gen. 56, 519–527 (1995).
Lange, K. & Sobel, E. A random walk method for computing genetic location scores. Am. J. Hum. Gen. 49, 1320–1334 (1991).
Thompson, E. A. in Computer Science and Statistics: Proceedings of the 23rd Symposium on the Interface (eds Keramidas, E. M. & Kaufman, S. M.) 321–328 (Interface Foundation of North America, Fairfax Station, Virginia, 1991).
Hoeschele, I. in Handbook of Statistical Genetics (ed. Balding, D. J.) 599–644 (John Wiley and Sons, New York, 2001). An extensive review of methods used to map quantitative trait loci in humans and other species.
Acknowledgements
We thank the four anonymous referees for their comments. Work on this paper was supported by grants from the Biotechnology and Biological Sciences Research Council and the Natural Environment Research Council to M.A.B., and by grants from the National Institutes of Health and the Canadian Institute of Health Research to B.R.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
Related links
DATABASES
OMIM
FURTHER INFORMATION
Bayesian population genetics programs and links
Bayesian population genetics programs and links
Bayesian population genetics programs and links
Bayesian sequence analysis web sites
Bayesian sequence analysis web sites
Detecting selection with comparative data, population genetic analysis
Genetic analysis software links (linkage analysis)
Genetic Software Forum (discussion list)
National Center for Biotechnology Information
Glossary
- STATISTICAL INFERENCE
-
The process whereby data are observed and then statements are made about unknown features of the system that gave rise to the data.
- PROBABILISTIC MODEL
-
A model in which the data are modelled as random variables, the probability distribution of which depends on parameter values. Bayesian models are sometimes called fully probabilistic because the parameter values are also treated as random variables.
- LIKELIHOOD
-
The probability of the data fora particular set of parameter values.
- MARKOV CHAIN
-
A model that is suitable for modelling a sequence of random variables, such as nucleotide base pairs in DNA, in which the probability that a variable assumes any specific value depends only on the value of a specified number of most recent variables that precede it. In an nth-order Markov chain, the probability distribution of a variable depends on the n preceding observations.
- MARGINAL LIKELIHOOD
-
Also known as the 'prior predictive distribution'. The probability distribution of the data irrespective of the parameter values.
- RANDOM VARIABLE
-
A quantity that might take any of a range of values (discrete or continuous) that cannot be predicted with certainty but only described probabilistically.
- JOINT PROBABILITY DISTRIBUTION
-
The probability distribution of all combinations of two or more random variables.
- PRIOR [DISTRIBUTION]
-
The probability distribution of parameter values before observing the data.
- CONDITIONAL DISTRIBUTION
-
The distribution of one or more random variables when other random variables of a joint probability distribution are fixed at particular values.
- POSTERIOR DISTRIBUTION
-
The conditional distribution of the parameter given the observed data.
- POINT ESTIMATE
-
A summary of the location of a parameter value. In a Bayesian setting, this is generally the mean, mode or median of the posterior distribution.
- INTERVAL ESTIMATE
-
An estimate of the region in which the true parameter value is believed to be located.
- METHOD OF MOMENTS
-
A method for estimating parameters by using theory to obtain a formula for the expected value of statistics measured from the data as a function of the parameter values to be estimated. The observed values of these statistics are then equated to the expected values. The formula is inverted to obtain an estimate of the parameter.
- FREQUENTIST INFERENCE
-
Statistical inference in which probability is interpreted as the relative frequency of occurrences in an infinite sequence of trials.
- COALESCENT THEORY
-
A theory that describes the genealogy of chromosomes or genes. Under many life-history schemes (discrete generations, overlapping generations, non-random mating, and so on), taking certain limits, the statistical distribution of branch lengths in genealogies follows a simple form. Coalescent theory describes this distribution.
- PARAMETRIC BOOTSTRAPPING
-
The process of repeatedly simulating new data sets with parameters that are inferred from the observed data, and then re-estimating the parameters from these simulated data sets. This process is used to obtain confidence intervals.
- EFFECTIVE POPULATION SIZE
-
(Ne). The size of a random mating population under a simple Fisher–Wright model that has an equivalent rate of inbreeding to that of the observed population, which might have additional complexities such as variable population size or biased sex ratio.
- NON-IDENTIFIABLE [PARAMETERS]
-
One or more model parameters are non-identifiable if different combinations of the parameters generate the same likelihood of the data.
- HIERARCHICAL BAYESIAN MODEL
-
In a standard Bayesian model, the parameters are drawn from prior distributions, the parameters of which are fixed by the modeller. In a hierarchical model, these parameters, usually referred to as 'hyperparameters', are also free to vary and are themselves drawn from priors, often referred to as 'hyperpriors'. This form of modelling is most useful for data that is composed of exchangeable groups, such as genes, for which the possibility is required that the parameters that describe each group might or might not be the same.
- APPROXIMATE BAYESIAN COMPUTATION
-
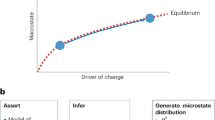
The data are simplified by representation as a set of summary statistics and simulations used to draw samples from the joint distribution of parameters and summary statistics (that is, the distribution shown in figure 1). The posterior distribution is approximated by estimating the conditional distribution of parameters in the vicinity of the summary statistics that are measured from the data (the vertical dotted line in figure 1) avoiding the need to calculate a likelihood function.
- MULTILOCUS GENOTYPES
-
The combinations of alleles that are observed when individuals are simultaneously genotyped at two or more genetic marker loci.
- ASSOCIATION STUDY
-
If two or more variables have joint outcomes that are more frequent than would be expected by chance (if the two variables were independent), they are associated. An association study statistically examines patterns of co-occurrence of variables, such as genetic variants and disease phenotypes, to identify factors (genes) that might contribute to disease risk.
- INBREEDING COEFFICIENT
-
The probability of homozygosity by descent — that is, the probability that a zygote obtains copies of the same ancestral gene from both its parents because they are related.
- COMPARATIVE METHODS
-
Methods for comparing traits across species to identify trends in character evolution that indicate the effects of natural selection.
- EMPIRICAL BAYES PROCEDURE
-
A hierarchical model in which the hyperparameter is not a random variable but is estimated by some other (often classical) means.
- HIDDEN MARKOV MODEL
-
This is an enhancement of a Markov chain model, in which the state of each observation is drawn randomly from a distribution, the parameters of which follow a Markov chain. For example, the parameter might be an indicator for whether a DNA region is coding or non-coding, and the observation is the base at each nucleotide.
- DYNAMIC PROGRAMMING
-
A large class of programmimg algorithms that are based on breaking a large problem down (if possible) into incremental steps so that, at any given stage, optimal solutions are known sub-problems.
- BORROW STRENGTH
-
This is the tendency in a hierarchical Bayesian model for the posterior distributions of parameters among exchangeable units (for example, genes) to become narrower as a result of pooling information across units.
- MODEL SELECTION
-
The process of choosing among different models given their posterior probability.
- PARALOGOUS
-
This refers to sequences that have arisen by duplications within a single genome.
- ELSTON–STEWART ALGORITHM
-
An iterative algorithm for linkage mapping. The algorithm calculates the likelihood of marker genotypes on a pedigree. Calculations on the basis of the algorithm are efficient for relatively large families, but its application is typically limited to a small number of markers.
- LANDER–GREEN–KRUGYLAK ALGORITHM
-
An iterative algorithm that is used for linkage mapping. It iteratively calculates the likelihood across markers on a chromosome, rather than across families, as in the Elston–Stewart algorithm. This allows efficient calculation of pedigree likelihoods for small families with many linked markers.
- FAMILY-BASED ASSOCIATION TESTS
-
A general class of genetic association tests that uses families with one or more affected children as the observations rather than unrelated cases and controls. The analysis treats the allele that is transmitted to (one or more) affected children from each parent as the 'case' and the untransmitted allele is treated as the 'control' to avoid the influence of population subdivision.
- BAYES FACTOR
-
The ratio of the prior probabilities of the null versus the alternative hypotheses over the ratio of the posterior probabilities. This can be interpreted as the relative odds that the hypothesis is true before and after examining the data. If the prior odds are equal, this simplifies to become the likelihood ratio.
- LD MAPPING
-
A procedure for fine-scale localization to a region of a chromosome of a mutation that causes a detectable phenotype (often a disease) by use of linkage disequilibrium between the phenotype that is induced by the mutation and markers that are located near the mutation on the chromosome.
- CONVERGENCE
-
The inexorable tendency for a mathematical function to approach some particular value (or set of values) with increasing n. In the case of Markov chain Monte Carlo, n is the number of simulation replicates and the values that the chain approaches are the posterior probabilities.
Rights and permissions
About this article
Cite this article
Beaumont, M., Rannala, B. The Bayesian revolution in genetics. Nat Rev Genet 5, 251–261 (2004). https://doi.org/10.1038/nrg1318
Issue Date:
DOI: https://doi.org/10.1038/nrg1318
This article is cited by
-
Stronger genetic differentiation among within-population genetic groups than among populations in Scots pine provides new insights into within-population genetic structuring
Scientific Reports (2024)
-
Bayesian estimates for genetic and phenotypic parameters of growth traits in Sahiwal cattle
Tropical Animal Health and Production (2023)
-
Phylogeography reveals complex historical processes and different evolutionarily significant units in Aegla scamosa freshwater crabs
Hydrobiologia (2023)
-
Adaptability and stability of Coffea canephora to dynamic environments using the Bayesian approach
Scientific Reports (2022)
-
Updated knowledge in the estimation of genetics parameters: a Bayesian approach in white oat (Avena sativa L.)
Euphytica (2022)