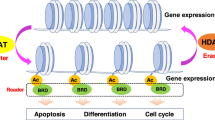
Key Points
-
Epigenetic deregulation can underpin the onset and progression of several human diseases.
-
The expression and/or function of histone deacetylases (HDACs) is often perturbed in cancer, neurological syndromes and immune disorders.
-
HDACs can be effectively targeted using small-molecule chemical compounds, and more selective agents are currently being developed and further tested.
-
Histones are not the only substrates of HDACs, and altered acetylation of diverse cellular proteins may be important in disease aetiology and the response to HDAC inhibitors.
-
HDACs function as the catalytic subunits of large multiprotein complexes, and the molecular and biological consequences of HDAC inhibition need to be assessed in this context.
-
HDAC inhibitors have been approved for the treatment of certain haematological malignancies and are being clinically evaluated alone and in combination with other agents for efficacy in other cancer settings, in neurological diseases and in immune disorders such as autoimmunity.
Abstract
Epigenetic aberrations, which are recognized as key drivers of several human diseases, are often caused by genetic defects that result in functional deregulation of epigenetic proteins, their altered expression and/or their atypical recruitment to certain gene promoters. Importantly, epigenetic changes are reversible, and epigenetic enzymes and regulatory proteins can be targeted using small molecules. This Review discusses the role of altered expression and/or function of one class of epigenetic regulators — histone deacetylases (HDACs) — and their role in cancer, neurological diseases and immune disorders. We highlight the development of small-molecule HDAC inhibitors and their use in the laboratory, in preclinical models and in the clinic.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Berdasco, M. & Esteller, M. Genetic syndromes caused by mutations in epigenetic genes. Hum. Genet. 132, 359–383 (2013).
Baylin, S. B. & Jones, P. A. A decade of exploring the cancer epigenome — biological and translational implications. Nature Rev. Cancer 11, 726–734 (2011).
Berger, S. L. The complex language of chromatin regulation during transcription. Nature 447, 407–412 (2007).
Kouzarides, T. Chromatin modifications and their function. Cell 128, 693–705 (2007).
Dawson, M. A. & Kouzarides, T. Cancer epigenetics: from mechanism to therapy. Cell 150, 12–27 (2012).
Dawson, M. A., Kouzarides, T. & Huntly, B. J. Targeting epigenetic readers in cancer. N. Engl. J. Med. 367, 647–657 (2012). References 5 and 6 are excellent review articles providing a contemporary view of the links between cancer genetics and epigenetics.
Kaminskas, E. et al. Approval summary: azacitidine for treatment of myelodysplastic syndrome subtypes. Clin. Cancer Res. 11, 3604–3608 (2005).
Kaminskas, E., Farrell, A. T., Wang, Y. C., Sridhara, R. & Pazdur, R. FDA drug approval summary: azacitidine (5-azacytidine, Vidaza) for injectable suspension. Oncologist 10, 176–182 (2005).
Daigle, S. R. et al. Potent inhibition of DOT1L as treatment of MLL-fusion leukemia. Blood 122, 1017–1025 (2013).
Yu, W. et al. Catalytic site remodelling of the DOT1L methyltransferase by selective inhibitors. Nature Commun. 3, 1288 (2012).
McCabe, M. T. et al. EZH2 inhibition as a therapeutic strategy for lymphoma with EZH2-activating mutations. Nature 492, 108–112 (2012).
Knutson, S. K. et al. A selective inhibitor of EZH2 blocks H3K27 methylation and kills mutant lymphoma cells. Nature Chem. Biol. 8, 890–896 (2012).
Bonham, K. et al. Effects of a novel arginine methyltransferase inhibitor on T-helper cell cytokine production. FEBS J. 277, 2096–2108 (2010).
Kleinschmidt, M. A., de Graaf, P., van Teeffelen, H. A. & Timmers, H. T. Cell cycle regulation by the PRMT6 arginine methyltransferase through repression of cyclin-dependent kinase inhibitors. PLoS ONE 7, e41446 (2012).
West, A. C. & Johnstone, R. W. New and emerging HDAC inhibitors for cancer treatment. J. Clin. Invest. 124, 30–39 (2014).
Schenk, T. et al. Inhibition of the LSD1 (KDM1A) demethylase reactivates the all-trans-retinoic acid differentiation pathway in acute myeloid leukemia. Nature Med. 18, 605–611 (2012).
Willmann, D. et al. Impairment of prostate cancer cell growth by a selective and reversible lysine-specific demethylase 1 inhibitor. Int. J. Cancer 131, 2704–2709 (2012).
Shi, L., Cui, S., Engel, J. D. & Tanabe, O. Lysine-specific demethylase 1 is a therapeutic target for fetal hemoglobin induction. Nature Med. 19, 291–294 (2013).
Frieling, H. & Bleich, S. Tranylcypromine: new perspectives on an “old” drug. Eur. Arch. Psychiatry Clin. Neurosci. 256, 268–273 (2006).
Tedeschini, E. et al. Efficacy of antidepressants for late-life depression: a meta-analysis and meta-regression of placebo-controlled randomized trials. J. Clin. Psychiatry 72, 1660–1668 (2011).
Filippakopoulos, P. et al. Selective inhibition of BET bromodomains. Nature 468, 1067–1073 (2010).
Nicodeme, E. et al. Suppression of inflammation by a synthetic histone mimic. Nature 468, 1119–1123 (2010).
Delmore, J. E. et al. BET bromodomain inhibition as a therapeutic strategy to target c-Myc. Cell 146, 904–917 (2011).
Mertz, J. A. et al. Targeting MYC dependence in cancer by inhibiting BET bromodomains. Proc. Natl Acad. Sci. USA 108, 16669–16674 (2011).
Dawson, M. A. et al. Inhibition of BET recruitment to chromatin as an effective treatment for MLL-fusion leukaemia. Nature 478, 529–533 (2011).
Zuber, J. et al. RNAi screen identifies Brd4 as a therapeutic target in acute myeloid leukaemia. Nature 478, 524–528 (2011).
Bandukwala, H. S. et al. Selective inhibition of CD4+ T-cell cytokine production and autoimmunity by BET protein and c-Myc inhibitors. Proc. Natl Acad. Sci. USA 109, 14532–14537 (2012).
Choudhary, C. et al. Lysine acetylation targets protein complexes and co-regulates major cellular functions. Science 325, 834–840 (2009). This is an important demonstration of non-histone substrates of HDACs and the biological effects of protein acetylation.
Jaenisch, R. & Bird, A. Epigenetic regulation of gene expression: how the genome integrates intrinsic and environmental signals. Nature Genet. 33, S245–S454 (2003). References 23–29 demonstrate the antitumour activities of novel bromodomain inhibitors.
Rafehi, H. et al. Vascular histone deacetylation by pharmacological HDAC inhibition. Genome Res. http://dx.doi.org/10.1101/gr.168781.113 (2014).
Dudakovic, A. et al. Histone deacetylase inhibition promotes osteoblast maturation by altering the histone H4 epigenome and reduces Akt phosphorylation. J. Biol. Chem. 288, 28783–28791 (2013).
Peart, M. J. et al. Identification and functional significance of genes regulated by structurally different histone deacetylase inhibitors. Proc. Natl Acad. Sci. USA 102, 3697–3702 (2005).
Gray, S. G., Qian, C. N., Furge, K., Guo, X. & Teh, B. T. Microarray profiling of the effects of histone deacetylase inhibitors on gene expression in cancer cell lines. Int. J. Oncol. 24, 773–795 (2004).
Bantscheff, M. et al. Chemoproteomics profiling of HDAC inhibitors reveals selective targeting of HDAC complexes. Nature Biotech. 29, 255–265 (2011). This is a demonstration of the functional role of HDACs in multiprotein complexes and the use of chromoproteomics to demonstrate the specificity of HDAC inhibitors in physiological circumstances.
Delcuve, G. P., Khan, D. H. & Davie, J. R. Roles of histone deacetylases in epigenetic regulation: emerging paradigms from studies with inhibitors. Clin. Epigenet. 4, 5 (2012).
Leder, A. & Leder, P. Butyric acid, a potent inducer of erythroid differentiation in cultured erythroleukemic cells. Cell 5, 319–322 (1975).
Riggs, M. G., Whittaker, R. G., Neumann, J. R. & Ingram, V. M. n-Butyrate causes histone modification in HeLa and Friend erythroleukaemia cells. Nature 268, 462–464 (1977).
Licht, J. D. AML1 and the AML1–ETO fusion protein in the pathogenesis of t(8;21) AML. Oncogene 20, 5660–5679 (2001).
Liu, Y. et al. The tetramer structure of the Nervy homology two domain, NHR2, is critical for AML1/ETO's activity. Cancer Cell 9, 249–260 (2006).
Peterson, L. F. & Zhang, D. E. The 8;21 translocation in leukemogenesis. Oncogene 23, 4255–4262 (2004).
Minucci, S. & Pelicci, P. G. Histone deacetylase inhibitors and the promise of epigenetic (and more) treatments for cancer. Nature Rev. Cancer 6, 38–51 (2006).
Di Croce, L. et al. Methyltransferase recruitment and DNA hypermethylation of target promoters by an oncogenic transcription factor. Science 295, 1079–1082 (2002).
Rego, E. M. et al. Retinoic acid (RA) and As2O3 treatment in transgenic models of acute promyelocytic leukemia (APL) unravel the distinct nature of the leukemogenic process induced by the PML-RARα and PLZF-RARα oncoproteins. Proc. Natl Acad. Sci. USA 97, 10173–10178 (2000).
Licht, J. D. et al. Clinical and molecular characterization of a rare syndrome of acute promyelocytic leukemia associated with translocation (11;17). Blood 85, 1083–1094 (1995).
Halkidou, K. et al. Upregulation and nuclear recruitment of HDAC1 in hormone refractory prostate cancer. Prostate 59, 177–189 (2004).
Zimmermann, S. et al. Reduced body size and decreased intestinal tumor rates in HDAC2-mutant mice. Cancer Res. 67, 9047–9054 (2007).
Bhaskara, S. et al. Hdac3 is essential for the maintenance of chromatin structure and genome stability. Cancer Cell 18, 436–447 (2010). This study indicates a tumour-suppressive role for HDAC3.
Dovey, O. M. et al. Histone deacetylase 1 and 2 are essential for normal T-cell development and genomic stability in mice. Blood 121, 1335–1344 (2013).
Heideman, M. R. et al. Dosage-dependent tumor suppression by histone deacetylases 1 and 2 through regulation of c-Myc collaborating genes and p53 function. Blood 121, 2038–2050 (2013).
Santoro, F. et al. A dual role for Hdac1: oncosuppressor in tumorigenesis, oncogene in tumor maintenance. Blood 121, 3459–3468 (2013). References 48–50 provide experimental evidence supporting the tumour-suppressive functions of HDAC1 and HDAC2.
Guan, J. S. et al. HDAC2 negatively regulates memory formation and synaptic plasticity. Nature 459, 55–60 (2009).
Graff, J. et al. An epigenetic blockade of cognitive functions in the neurodegenerating brain. Nature 483, 222–226 (2012).
Montgomery, R. L., Hsieh, J., Barbosa, A. C., Richardson, J. A. & Olson, E. N. Histone deacetylases 1 and 2 control the progression of neural precursors to neurons during brain development. Proc. Natl Acad. Sci. USA 106, 7876–7881 (2009).
Akhtar, M. W. et al. Histone deacetylases 1 and 2 form a developmental switch that controls excitatory synapse maturation and function. J. Neurosci. 29, 8288–8297 (2009).
Majdzadeh, N., Morrison, B. E. & D'Mello, S. R. Class IIA HDACs in the regulation of neurodegeneration. Front. Biosci. 13, 1072–1082 (2008).
Majdzadeh, N. et al. HDAC4 inhibits cell-cycle progression and protects neurons from cell death. Dev. Neurobiol. 68, 1076–1092 (2008).
Kim, M. S. et al. An essential role for histone deacetylase 4 in synaptic plasticity and memory formation. J. Neurosci. 32, 10879–10886 (2012).
Williams, S. R. et al. Haploinsufficiency of HDAC4 causes brachydactyly mental retardation syndrome, with brachydactyly type E, developmental delays, and behavioral problems. Am. J. Hum. Genet. 87, 219–228 (2010).
Fukada, M. et al. Loss of deacetylation activity of Hdac6 affects emotional behavior in mice. PLoS ONE 7, e30924 (2012).
Lee, V. M., Goedert, M. & Trojanowski, J. Q. Neurodegenerative tauopathies. Annu. Rev. Neurosci. 24, 1121–1159 (2001).
Min, S. W. et al. Acetylation of tau inhibits its degradation and contributes to tauopathy. Neuron 67, 953–966 (2010).
Cohen, T. J. et al. The acetylation of tau inhibits its function and promotes pathological tau aggregation. Nature Commun. 2, 252 (2011).
Irwin, D. J. et al. Acetylated tau neuropathology in sporadic and hereditary tauopathies. Am. J. Pathol. 183, 344–351 (2013).
Selenica, M. L. et al. Histone deacetylase 6 inhibition improves memory and reduces total tau levels in a mouse model of tau deposition. Alzheimers Res. Ther. 6, 12 (2014).
Cook, C. et al. Acetylation of the KXGS motifs in tau is a critical determinant in modulation of tau aggregation and clearance. Hum. Mol. Genet. 23, 104–116 (2014).
Simoes-Pires, C. et al. HDAC6 as a target for neurodegenerative diseases: what makes it different from the other HDACs? Mol. Neurodegener. 8, 7 (2013).
Jeong, H. et al. Acetylation targets mutant huntingtin to autophagosomes for degradation. Cell 137, 60–72 (2009). This is a landmark demonstration of the role of acetylated HTT in Huntington's disease.
Govindarajan, N. et al. Reducing HDAC6 ameliorates cognitive deficits in a mouse model for Alzheimer's disease. EMBO Mol. Med. 5, 52–63 (2013).
Dompierre, J. P. et al. Histone deacetylase 6 inhibition compensates for the transport deficit in Huntington's disease by increasing tubulin acetylation. J. Neurosci. 27, 3571–3583 (2007).
Bobrowska, A., Paganetti, P., Matthias, P. & Bates, G. P. Hdac6 knock-out increases tubulin acetylation but does not modify disease progression in the R6/2 mouse model of Huntington's disease. PLoS ONE 6, e20696 (2011).
Pandey, U. B. et al. HDAC6 rescues neurodegeneration and provides an essential link between autophagy and the UPS. Nature 447, 859–863 (2007).
Brady, R. O., Kanfer, J. N., Bradley, R. M. & Shapiro, D. Demonstration of a deficiency of glucocerebroside-cleaving enzyme in Gaucher's disease. J. Clin. Invest. 45, 1112–1115 (1966).
Lu, J. et al. Histone deacetylase inhibitors prevent the degradation and restore the activity of glucocerebrosidase in Gaucher disease. Proc. Natl Acad. Sci. USA 108, 21200–21205 (2011).
Kovacs, J. J. et al. HDAC6 regulates Hsp90 acetylation and chaperone-dependent activation of glucocorticoid receptor. Mol. Cell 18, 601–607 (2005).
Shakespear, M. R., Halili, M. A., Irvine, K. M., Fairlie, D. P. & Sweet, M. J. Histone deacetylases as regulators of inflammation and immunity. Trends Immunol. 32, 335–343 (2011).
Hancock, W. W., Akimova, T., Beier, U. H., Liu, Y. & Wang, L. HDAC inhibitor therapy in autoimmunity and transplantation. Ann. Rheum. Dis. 71 (Suppl. 2), 46–54 (2012).
Sweet, M. J., Shakespear, M. R., Kamal, N. A. & Fairlie, D. P. HDAC inhibitors: modulating leukocyte differentiation, survival, proliferation and inflammation. Immunol. Cell Biol. 90, 14–22 (2012).
Xu, M., Nie, L., Kim, S. H. & Sun, X. H. STAT5-induced Id-1 transcription involves recruitment of HDAC1 and deacetylation of C/EBPβ. EMBO J. 22, 893–904 (2003).
Kramer, O. H. et al. A phosphorylation-acetylation switch regulates STAT1 signaling. Genes Dev. 23, 223–235 (2009).
Klampfer, L., Huang, J., Swaby, L. A. & Augenlicht, L. Requirement of histone deacetylase activity for signaling by STAT1. J. Biol. Chem. 279, 30358–30368 (2004).
Chang, H. M. et al. Induction of interferon-stimulated gene expression and antiviral responses require protein deacetylase activity. Proc. Natl Acad. Sci. USA 101, 9578–9583 (2004).
Nusinzon, I. & Horvath, C. M. Interferon-stimulated transcription and innate antiviral immunity require deacetylase activity and histone deacetylase 1. Proc. Natl Acad. Sci. USA 100, 14742–14747 (2003).
Lobera, M. et al. Selective class IIa histone deacetylase inhibition via a nonchelating zinc-binding group. Nature Chem. Biol. 9, 319–325 (2013).
Shakespear, M. R. et al. Histone deacetylase 7 promotes Toll-like receptor 4-dependent proinflammatory gene expression in macrophages. J. Biol. Chem. 288, 25362–25374 (2013).
Barneda-Zahonero, B. et al. HDAC7 is a repressor of myeloid genes whose downregulation is required for transdifferentiation of pre-B cells into macrophages. PLoS Genet. 9, e1003503 (2013).
Navarro, M. N., Goebel, J., Feijoo-Carnero, C., Morrice, N. & Cantrell, D. A. Phosphoproteomic analysis reveals an intrinsic pathway for the regulation of histone deacetylase 7 that controls the function of cytotoxic T lymphocytes. Nature Immunol. 12, 352–361 (2011).
de Zoeten, E. F., Wang, L., Sai, H., Dillmann, W. H. & Hancock, W. W. Inhibition of HDAC9 increases T regulatory cell function and prevents colitis in mice. Gastroenterology 138, 583–594 (2010).
Tao, R. et al. Deacetylase inhibition promotes the generation and function of regulatory T cells. Nature Med. 13, 1299–1307 (2007). This is an important study demonstrating the role of HDACs in regulating immune responses.
de Zoeten, E. F. et al. Histone deacetylase 6 and heat shock protein 90 control the functions of Foxp3+ T-regulatory cells. Mol. Cell. Biol. 31, 2066–2078 (2011).
Serrador, J. M. et al. HDAC6 deacetylase activity links the tubulin cytoskeleton with immune synapse organization. Immunity 20, 417–428 (2004).
Cabrero, J. R. et al. Lymphocyte chemotaxis is regulated by histone deacetylase 6, independently of its deacetylase activity. Mol. Biol. Cell 17, 3435–3445 (2006).
Yamaguchi, T. et al. Histone deacetylases 1 and 2 act in concert to promote the G1-to-S progression. Genes Dev. 24, 455–469 (2010).
Grausenburger, R. et al. Conditional deletion of histone deacetylase 1 in T cells leads to enhanced airway inflammation and increased Th2 cytokine production. J. Immunol. 185, 3489–3497 (2010).
Villagra, A. et al. The histone deacetylase HDAC11 regulates the expression of interleukin 10 and immune tolerance. Nature Immunol. 10, 92–100 (2009).
Margolis, D. M. Histone deacetylase inhibitors and HIV latency. Curr. Opin. HIV AIDS 6, 25–29 (2011).
Bullen, C. K., Laird, G. M., Durand, C. M., Siliciano, J. D. & Siliciano, R. F. New ex vivo approaches distinguish effective and ineffective single agents for reversing HIV-1 latency in vivo. Nature Med. 20, 425–429 (2014).
Zhou, G., Du, T. & Roizman, B. The role of the CoREST/REST repressor complex in herpes simplex virus 1 productive infection and in latency. Viruses 5, 1208–1218 (2013).
Johnstone, R. W. Histone-deacetylase inhibitors: novel drugs for the treatment of cancer. Nature Rev. Drug Discov. 1, 287–299 (2002).
Fischle, W. et al. Enzymatic activity associated with class II HDACs is dependent on a multiprotein complex containing HDAC3 and SMRT/N-CoR. Mol. Cell 9, 45–57 (2002).
Backs, J., Backs, T., Bezprozvannaya, S., McKinsey, T. A. & Olson, E. N. Histone deacetylase 5 acquires calcium/calmodulin-dependent kinase II responsiveness by oligomerization with histone deacetylase 4. Mol. Cell. Biol. 28, 3437–3445 (2008).
Kong, H. S. et al. Preclinical studies of YK-4-272, an inhibitor of class II histone deacetylases by disruption of nucleocytoplasmic shuttling. Pharm. Res. 29, 3373–3383 (2012).
Lahm, A. et al. Unraveling the hidden catalytic activity of vertebrate class IIa histone deacetylases. Proc. Natl Acad. Sci. USA 104, 17335–17340 (2007).
Herman, D. et al. Histone deacetylase inhibitors reverse gene silencing in Friedreich's ataxia. Nature Chem. Biol. 2, 551–558 (2006).
Rai, M. et al. HDAC inhibitors correct frataxin deficiency in a Friedreich ataxia mouse model. PLoS ONE 3, e1958 (2008).
Sandi, C. et al. Prolonged treatment with pimelic O-aminobenzamide HDAC inhibitors ameliorates the disease phenotype of a Friedreich ataxia mouse model. Neurobiol. Dis. 42, 496–505 (2011).
Soragni, E. et al. Rationale for the development of 2-aminobenzamide histone deacetylase inhibitors as therapeutics for Friedreich ataxia. J. Child Neurol. 27, 1164–1173 (2012).
Wells, C. E. et al. Inhibition of histone deacetylase 3 causes replication stress in cutaneous T cell lymphoma. PLoS ONE 8, e68915 (2013).
Minami, J. et al. Histone deacetylase 3 as a novel therapeutic target in multiple myeloma. Leukemia 28, 680–689 (2013).
Santo, L. et al. Preclinical activity, pharmacodynamic, and pharmacokinetic properties of a selective HDAC6 inhibitor, ACY-1215, in combination with bortezomib in multiple myeloma. Blood 119, 2579–2589 (2012).
McConkey, D. J., White, M. & Yan, W. HDAC inhibitor modulation of proteotoxicity as a therapeutic approach in cancer. Adv. Cancer Res. 116, 131–163 (2012).
Newbold, A. et al. Molecular and biological analysis of histone deacetylase inhibitors with diverse specificities. Mol. Cancer Ther. 12, 2709–2721 (2013).
Schroeder, F. A. et al. A selective HDAC 1/2 inhibitor modulates chromatin and gene expression in brain and alters mouse behavior in two mood-related tests. PLoS ONE 8, e71323 (2013).
Ai, T., Cui, H. & Chen, L. Multi-targeted histone deacetylase inhibitors in cancer therapy. Curr. Med. Chem. 19, 475–487 (2012).
Qian, C. et al. Cancer network disruption by a single molecule inhibitor targeting both histone deacetylase activity and phosphatidylinositol 3-kinase signaling. Clin. Cancer Res. 18, 4104–4113 (2012).
Lai, C. J. et al. CUDC-101, a multitargeted inhibitor of histone deacetylase, epidermal growth factor receptor, and human epidermal growth factor receptor 2, exerts potent anticancer activity. Cancer Res. 70, 3647–3656 (2010).
Wang, J. et al. Potential advantages of CUDC-101, a multitargeted HDAC, EGFR, and HER2 inhibitor, in treating drug resistance and preventing cancer cell migration and invasion. Mol. Cancer Ther. 12, 925–936 (2013).
Needham, L. A. et al. Drug targeting to monocytes and macrophages using esterase-sensitive chemical motifs. J. Pharmacol. Exp. Ther. 339, 132–142 (2011).
Ossenkoppele, G. J. et al. A phase I first-in-human study with tefinostat - a monocyte/macrophage targeted histone deacetylase inhibitor - in patients with advanced haematological malignancies. Br. J. Haematol. 162, 191–201 (2013).
Guerrant, W., Patil, V., Canzoneri, J. C. & Oyelere, A. K. Dual targeting of histone deacetylase and topoisomerase II with novel bifunctional inhibitors. J. Med. Chem. 55, 1465–1477 (2012).
Guerrant, W. et al. Dual-acting histone deacetylase-topoisomerase I inhibitors. Bioorg. Med. Chem. Lett. 23, 3283–3287 (2013).
Chen, G. L. et al. Discovery of a small molecular compound simultaneously targeting RXR and HADC: design, synthesis, molecular docking and bioassay. Bioorg. Med. Chem. Lett. 23, 3891–3895 (2013).
Gryder, B. E. et al. Histone deacetylase inhibitors equipped with estrogen receptor modulation activity. J. Med. Chem. 56, 5782–5796 (2013).
Chen, J. B. et al. Design and synthesis of dual-action inhibitors targeting histone deacetylases and 3-hydroxy-3-methylglutaryl coenzyme A reductase for cancer treatment. J. Med. Chem. 56, 3645–3655 (2013).
Tavera-Mendoza, L. E. et al. Incorporation of histone deacetylase inhibition into the structure of a nuclear receptor agonist. Proc. Natl Acad. Sci. USA 105, 8250–8255 (2008).
Lamblin, M. et al. Vitamin D receptor agonist/histone deacetylase inhibitor molecular hybrids. Bioorg. Med. Chem. 18, 4119–4137 (2010).
Ko, K. S., Steffey, M. E., Brandvold, K. R. & Soellner, M. B. Development of a chimeric c-Src kinase and HDAC inhibitor. ACS Med. Chem. Lett. 4, 779–783 (2013).
Patel, H. K. et al. A chimeric SERM-histone deacetylase inhibitor approach to breast cancer therapy. ChemMedChem 9, 602–613 (2013).
Weinstein, I. B. Cancer. Addiction to oncogenes — the Achilles heal of cancer. Science 297, 63–64 (2002).
Bolden, J. E. et al. HDAC inhibitors induce tumor-cell-selective pro-apoptotic transcriptional responses. Cell Death Dis. 4, e519 (2013).
Insinga, A. et al. Inhibitors of histone deacetylases induce tumor-selective apoptosis through activation of the death receptor pathway. Nature Med. 11, 71–76 (2005).
Nebbioso, A. et al. Tumor-selective action of HDAC inhibitors involves TRAIL induction in acute myeloid leukemia cells. Nature Med. 11, 77–84 (2005). References 130 and 131 demonstrate a role for the death receptor pathway in mediating apoptosis induced by HDAC inhibitors.
Ungerstedt, J. S. et al. Role of thioredoxin in the response of normal and transformed cells to histone deacetylase inhibitors. Proc. Natl Acad. Sci. USA 102, 673–678 (2005).
Fuino, L. et al. Histone deacetylase inhibitor LAQ824 down-regulates Her-2 and sensitizes human breast cancer cells to trastuzumab, taxotere, gemcitabine, and epothilone B. Mol. Cancer Ther. 2, 971–984 (2003).
Bali, P. et al. Inhibition of histone deacetylase 6 acetylates and disrupts the chaperone function of heat shock protein 90: a novel basis for antileukemia activity of histone deacetylase inhibitors. J. Biol. Chem. 280, 26729–26734 (2005).
Nimmanapalli, R. et al. Histone deacetylase inhibitor LAQ824 both lowers expression and promotes proteasomal degradation of Bcr-Abl and induces apoptosis of imatinib mesylate-sensitive or -refractory chronic myelogenous leukemia-blast crisis cells. Cancer Res. 63, 5126–5135 (2003).
Yu, W. et al. Heat shock protein 90 inhibition results in altered downstream signaling of mutant KIT and exerts synergistic effects on Kasumi-1 cells when combining with histone deacetylase inhibitor. Leuk. Res. 35, 1212–1218 (2011).
Wang, Y. et al. FK228 inhibits Hsp90 chaperone function in K562 cells via hyperacetylation of Hsp70. Biochem. Biophys. Res. Commun. 356, 998–1003 (2007).
Nguyen, T. et al. HDAC inhibitors potentiate the activity of the BCR/ABL kinase inhibitor KW-2449 in imatinib-sensitive or -resistant BCR/ABL+ leukemia cells in vitro and in vivo. Clin. Cancer Res. 17, 3219–3232 (2011).
Jaboin, J. et al. MS-27-275, an inhibitor of histone deacetylase, has marked in vitro and in vivo antitumor activity against pediatric solid tumors. Cancer Res. 62, 6108–6115 (2002).
Stumpel, D. J. et al. Connectivity mapping identifies HDAC inhibitors for the treatment of t(4;11)-positive infant acute lymphoblastic leukemia. Leukemia 26, 682–692 (2012).
Marshall, G. M. et al. Transcriptional upregulation of histone deacetylase 2 promotes Myc-induced oncogenic effects. Oncogene 29, 5957–5968 (2010).
Zhang, X. et al. Coordinated silencing of MYC-mediated miR-29 by HDAC3 and EZH2 as a therapeutic target of histone modification in aggressive B-cell lymphomas. Cancer Cell 22, 506–523 (2012).
Bolden, J. E., Peart, M. J. & Johnstone, R. W. Anticancer activities of histone deacetylase inhibitors. Nature Rev. Drug Discov. 5, 769–784 (2006).
Ellis, L. et al. The histone deacetylase inhibitors LAQ824 and LBH589 do not require death receptor signaling or a functional apoptosome to mediate tumor cell death or therapeutic efficacy. Blood 114, 380–393 (2009).
Kroemer, G., Galluzzi, L., Kepp, O. & Zitvogel, L. Immunogenic cell death in cancer therapy. Annu. Rev. Immunol. 31, 51–72 (2013).
Setiadi, A. F. et al. Epigenetic enhancement of antigen processing and presentation promotes immune recognition of tumors. Cancer Res. 68, 9601–9607 (2008).
Christiansen, A. J. et al. Eradication of solid tumors using histone deacetylase inhibitors combined with immune-stimulating antibodies. Proc. Natl Acad. Sci. USA 108, 4141–4146 (2011).
West, A. C. et al. An intact immune system is required for the anti-cancer activities of histone deacetylase inhibitors. Cancer Res. 73, 7265–7276 (2013).
Schwartz, B. E. et al. Differentiation of NUT midline carcinoma by epigenomic reprogramming. Cancer Res. 71, 2686–2696 (2011).
Bots, M. et al. Differentiation therapy for the treatment of t(8;21) acute myeloid leukemia using histone deacetylase inhibitors. Blood 123, 1341–1352 (2014).
Duvic, M. et al. Phase 2 trial of oral vorinostat (suberoylanilide hydroxamic acid, SAHA) for refractory cutaneous T-cell lymphoma (CTCL). Blood 109, 31–39 (2007).
VanderMolen, K. M., McCulloch, W., Pearce, C. J. & Oberlies, N. H. Romidepsin (Istodax, NSC 630176, FR901228, FK228, depsipeptide): a natural product recently approved for cutaneous T-cell lymphoma. J. Antibiot. 64, 525–531 (2011).
New, M., Olzscha, H. & La Thangue, N. B. HDAC inhibitor-based therapies: can we interpret the code? Mol. Oncol. 6, 637–656 (2012).
Nebbioso, A., Carafa, V., Benedetti, R. & Altucci, L. Trials with 'epigenetic' drugs: an update. Mol. Oncol. 6, 657–682 (2012).
Qiu, T. et al. Effects of treatment with histone deacetylase inhibitors in solid tumors: a review based on 30 clinical trials. Future Oncol. 9, 255–269 (2013).
Garcia-Manero, G. et al. Phase II trial of vorinostat with idarubicin and cytarabine for patients with newly diagnosed acute myelogenous leukemia or myelodysplastic syndrome. J. Clin. Oncol. 30, 2204–2210 (2012).
Bots, M. & Johnstone, R. W. Rational combinations using HDAC inhibitors. Clin. Cancer Res. 15, 3970–3977 (2009).
Thurn, K. T., Thomas, S., Moore, A. & Munster, P. N. Rational therapeutic combinations with histone deacetylase inhibitors for the treatment of cancer. Future Oncol. 7, 263–283 (2011).
Blum, W. et al. Phase I study of decitabine alone or in combination with valproic acid in acute myeloid leukemia. J. Clin. Oncol. 25, 3884–3891 (2007).
Badros, A. et al. Phase I study of vorinostat in combination with bortezomib for relapsed and refractory multiple myeloma. Clin. Cancer Res. 15, 5250–5257 (2009).
Weber, D. M. et al. Phase I trial of vorinostat combined with bortezomib for the treatment of relapsing and/or refractory multiple myeloma. Clin. Lymphoma Myeloma Leuk. 12, 319–324 (2012).
Millward, M. et al. Phase 1 clinical trial of the novel proteasome inhibitor marizomib with the histone deacetylase inhibitor vorinostat in patients with melanoma, pancreatic and lung cancer based on in vitro assessments of the combination. Invest. New Drugs 30, 2303–2317 (2012).
Dasmahapatra, G. et al. The pan-HDAC inhibitor vorinostat potentiates the activity of the proteasome inhibitor carfilzomib in human DLBCL cells in vitro and in vivo. Blood 115, 4478–4487 (2010).
Dasmahapatra, G. et al. Carfilzomib interacts synergistically with histone deacetylase inhibitors in mantle cell lymphoma cells in vitro and in vivo. Mol. Cancer Ther. 10, 1686–1697 (2011).
Munster, P. N. et al. A phase II study of the histone deacetylase inhibitor vorinostat combined with tamoxifen for the treatment of patients with hormone therapy-resistant breast cancer. Br. J. Cancer 104, 1828–1835 (2011).
Faller, D. V., Mentzer, S. J. & Perrine, S. P. Induction of the Epstein-Barr virus thymidine kinase gene with concomitant nucleoside antivirals as a therapeutic strategy for Epstein-Barr virus-associated malignancies. Curr. Opin. Oncol. 13, 360–367 (2001).
Perrine, S. P. et al. A phase 1/2 trial of arginine butyrate and ganciclovir in patients with Epstein-Barr virus-associated lymphoid malignancies. Blood 109, 2571–2578 (2007).
Ghosh, S. K., Perrine, S. P., Williams, R. M. & Faller, D. V. Histone deacetylase inhibitors are potent inducers of gene expression in latent EBV and sensitize lymphoma cells to nucleoside antiviral agents. Blood 119, 1008–1017 (2012).
Johnstone, R. W., Frew, A. J. & Smyth, M. J. The TRAIL apoptotic pathway in cancer onset, progression and therapy. Nature Rev. Cancer 8, 782–798 (2008).
Frew, A. J. et al. Combination therapy of established cancer using a histone deacetylase inhibitor and a TRAIL receptor agonist. Proc. Natl Acad. Sci. USA 105, 11317–11322 (2008).
Martin, B. P. et al. Antitumor activities and on-target toxicities mediated by a TRAIL receptor agonist following cotreatment with panobinostat. Int. J. Cancer 128, 2735–2747 (2011).
Whitecross, K. F. et al. Defining the target specificity of ABT-737 and synergistic antitumor activities in combination with histone deacetylase inhibitors. Blood 113, 1982–1991 (2009).
Chuang, D. M., Leng, Y., Marinova, Z., Kim, H. J. & Chiu, C. T. Multiple roles of HDAC inhibition in neurodegenerative conditions. Trends Neurosci. 32, 591–601 (2009).
Jia, H. et al. Histone deacetylase (HDAC) inhibitors targeting HDAC3 and HDAC1 ameliorate polyglutamine-elicited phenotypes in model systems of Huntington's disease. Neurobiol. Dis. 46, 351–361 (2012).
Steffan, J. S. et al. Histone deacetylase inhibitors arrest polyglutamine-dependent neurodegeneration in Drosophila. Nature 413, 739–743 (2001).
Kim, D. et al. Deregulation of HDAC1 by p25/Cdk5 in neurotoxicity. Neuron 60, 803–817 (2008).
Kozikowski, A. P. et al. Functional differences in epigenetic modulators-superiority of mercaptoacetamide-based histone deacetylase inhibitors relative to hydroxamates in cortical neuron neuroprotection studies. J. Med. Chem. 50, 3054–3061 (2007).
Butler, K. V. et al. Rational design and simple chemistry yield a superior, neuroprotective HDAC6 inhibitor, tubastatin A. J. Am. Chem. Soc. 132, 10842–10846 (2010).
Zhang, L. et al. Tubastatin A/ACY-1215 improves cognition in alzheimer's disease transgenic mice. J. Alzheimers Dis. http://dx.doi.org/10.3233/JAD-140066 (2014).
Zhang, L., Sheng, S. & Qin, C. The role of HDAC6 in Alzheimer's disease. J. Alzheimers Dis. 33, 283–295 (2013).
Subramanian, S., Bates, S., Wright, J., Espinoza-Delgado, I. & Piekarz, R. Clinical toxicities of histone deacetylase inhibitors. Pharmaceuticals 3, 2751–2767 (2010).
Seo, J., Howell, M. D., Singh, N. N. & Singh, R. N. Spinal muscular atrophy: an update on therapeutic progress. Biochim. Biophys. Acta 1832, 2180–2190 (2013).
Harahap, I. S. et al. Valproic acid increases SMN2 expression and modulates SF2/ASF and hnRNPA1 expression in SMA fibroblast cell lines. Brain Dev. 34, 213–222 (2012).
Evans, M. C., Cherry, J. J. & Androphy, E. J. Differential regulation of the SMN2 gene by individual HDAC proteins. Biochem. Biophys. Res. Commun. 414, 25–30 (2011).
Kwon, D. Y., Motley, W. W., Fischbeck, K. H. & Burnett, B. G. Increasing expression and decreasing degradation of SMN ameliorate the spinal muscular atrophy phenotype in mice. Hum. Mol. Genet. 20, 3667–3677 (2011).
Akimova, T., Beier, U. H., Liu, Y., Wang, L. & Hancock, W. W. Histone/protein deacetylases and T-cell immune responses. Blood 119, 2443–2451 (2012).
Halili, M. A., Andrews, M. R., Sweet, M. J. & Fairlie, D. P. Histone deacetylase inhibitors in inflammatory disease. Curr. Top. Med. Chem. 9, 309–319 (2009).
Hsieh, I. N. et al. Preclinical anti-arthritic study and pharmacokinetic properties of a potent histone deacetylase inhibitor MPT0G009. Cell Death Dis. 5, e1166 (2014).
Joosten, L. A., Leoni, F., Meghji, S. & Mascagni, P. Inhibition of HDAC activity by ITF2357 ameliorates joint inflammation and prevents cartilage and bone destruction in experimental arthritis. Mol. Med. 17, 391–396 (2011).
Archin, N. M. et al. Expression of latent human immunodeficiency type 1 is induced by novel and selective histone deacetylase inhibitors. AIDS 23, 1799–1806 (2009).
Archin, N. M. et al. Administration of vorinostat disrupts HIV-1 latency in patients on antiretroviral therapy. Nature 487, 482–485 (2012). This is an important demonstration that HDAC inhibitors can affect viral latency.
Ritchie, D. et al. Reactivation of DNA viruses in association with histone deacetylase inhibitor therapy: a case series report. Haematologica 94, 1618–1622 (2009).
Scala, S. et al. P-glycoprotein substrates and antagonists cluster into two distinct groups. Mol. Pharmacol. 51, 1024–1033 (1997).
Xiao, J. J. et al. Efflux of depsipeptide FK228 (FR901228, NSC-630176) is mediated by P-glycoprotein and multidrug resistance-associated protein 1. J. Pharmacol. Exp. Ther. 313, 268–276 (2005).
Ruefli, A. A. et al. Suberoylanilide hydroxamic acid (SAHA) overcomes multidrug resistance and induces cell death in P-glycoprotein-expressing cells. Int. J. Cancer 99, 292–298 (2002).
Lindemann, R. K. et al. Analysis of the apoptotic and therapeutic activities of histone deacetylase inhibitors by using a mouse model of B cell lymphoma. Proc. Natl Acad. Sci. USA 104, 8071–8076 (2007).
Newbold, A. et al. Characterisation of the novel apoptotic and therapeutic activities of the histone deacetylase inhibitor romidepsin. Mol. Cancer Ther. 7, 1066–1079 (2008).
Fantin, V. R. et al. Constitutive activation of signal transducers and activators of transcription predicts vorinostat resistance in cutaneous T-cell lymphoma. Cancer Res. 68, 3785–3794 (2008).
Fotheringham, S. et al. Genome-wide loss-of-function screen reveals an important role for the proteasome in HDAC inhibitor-induced apoptosis. Cancer Cell 15, 57–66 (2009). References 198 and 199 provide the first evidence for predictive biomarkers of tumour cell sensitivity to HDAC inhibitors.
Khan, O. et al. HR23B is a biomarker for tumor sensitivity to HDAC inhibitor-based therapy. Proc. Natl Acad. Sci. USA 107, 6532–6537 (2010).
Yeo, W. et al. Epigenetic therapy using belinostat for patients with unresectable hepatocellular carcinoma: a multicenter phase I/II study with biomarker and pharmacokinetic analysis of tumors from patients in the Mayo Phase II Consortium and the Cancer Therapeutics Research Group. J. Clin. Oncol. 30, 3361–3367 (2012).
Chen, L., Shinde, U., Ortolan, T. G. & Madura, K. Ubiquitin-associated (UBA) domains in Rad23 bind ubiquitin and promote inhibition of multi-ubiquitin chain assembly. EMBO Rep. 2, 933–938 (2001).
Chen, L. & Madura, K. Rad23 promotes the targeting of proteolytic substrates to the proteasome. Mol. Cell. Biol. 22, 4902–4913 (2002).
New, M. et al. A regulatory circuit that involves HR23B and HDAC6 governs the biological response to HDAC inhibitors. Cell Death Differ. 20, 1306–1316 (2013).
Xu, W., Ngo, L., Perez, G., Dokmanovic, M. & Marks, P. A. Intrinsic apoptotic and thioredoxin pathways in human prostate cancer cell response to histone deacetylase inhibitor. Proc. Natl Acad. Sci. USA 103, 15540–15545 (2006).
Garcia-Manero, G. et al. Phase 1 study of the histone deacetylase inhibitor vorinostat (suberoylanilide hydroxamic acid [SAHA]) in patients with advanced leukemias and myelodysplastic syndromes. Blood 111, 1060–1066 (2008).
Hu, Y. et al. Overcoming resistance to histone deacetylase inhibitors in human leukemia with the redox modulating compound β-phenylethyl isothiocyanate. Blood 116, 2732–2741 (2010).
Hug, B. A. & Lazar, M. A. ETO interacting proteins. Oncogene 23, 4270–4274 (2004).
Liu, S. et al. Interplay of RUNX1/MTG8 and DNA methyltransferase 1 in acute myeloid leukemia. Cancer Res. 65, 1277–1284 (2005).
Shia, W. J. et al. PRMT1 interacts with AML1-ETO to promote its transcriptional activation and progenitor cell proliferative potential. Blood 119, 4953–4962 (2012).
Roudaia, L. et al. CBFβ is critical for AML1-ETO and TEL-AML1 activity. Blood 113, 3070–3079 (2009).
Rice, K. L. & de The, H. The acute promyelotic leukemia (APL) success story: curing leukemia through targeted therapies. J. Intern. Med. 276, 61–70 (2014).
Villa, R. et al. Role of the polycomb repressive complex 2 in acute promyelocytic leukemia. Cancer Cell 11, 513–525 (2007).
Gupta, P., Reid, R. C., Iyer, A., Sweet, M. J. & Fairlie, D. P. Towards isozyme-selective HDAC inhibitors for interrogating disease. Curr. Top. Med. Chem. 12, 1479–1499 (2012).
Salisbury, C. M. & Cravatt, B. F. Activity-based probes for proteomic profiling of histone deacetylase complexes. Proc. Natl Acad. Sci. USA 104, 1171–1176 (2007).
Tessier, P. et al. Diphenylmethylene hydroxamic acids as selective class IIa histone deacetylase inhibitors. Bioorg. Med. Chem. Lett. 19, 5684–5688 (2009).
Malvaez, M. et al. HDAC3-selective inhibitor enhances extinction of cocaine-seeking behavior in a persistent manner. Proc. Natl Acad. Sci. USA 110, 2647–2652 (2013).
Jochems, J. et al. Antidepressant-like properties of novel HDAC6 Selective inhibitors with improved brain bioavailability. Neuropsychopharmacology 39, 389–400 (2013).
Haggarty, S. J., Koeller, K. M., Wong, J. C., Grozinger, C. M. & Schreiber, S. L. Domain-selective small-molecule inhibitor of histone deacetylase 6 (HDAC6)-mediated tubulin deacetylation. Proc. Natl Acad. Sci. USA 100, 4389–4394 (2003).
Namdar, M., Perez, G., Ngo, L. & Marks, P. A. Selective inhibition of histone deacetylase 6 (HDAC6) induces DNA damage and sensitizes transformed cells to anticancer agents. Proc. Natl Acad. Sci. USA 107, 20003–20008 (2010).
Vishwakarma, S. et al. Tubastatin, a selective histone deacetylase 6 inhibitor shows anti-inflammatory and anti-rheumatic effects. Int. Immunopharmacol. 16, 72–78 (2013).
Kaliszczak, M. et al. A novel small molecule hydroxamate preferentially inhibits HDAC6 activity and tumour growth. Br. J. Cancer 108, 342–350 (2013).
Lee, J. H. et al. Development of a histone deacetylase 6 inhibitor and its biological effects. Proc. Natl Acad. Sci. USA 110, 15704–15709 (2013).
Yu, C. W., Chang, P. T., Hsin, L. W. & Chern, J. W. Quinazolin-4-one derivatives as selective histone deacetylase-6 inhibitors for the treatment of Alzheimer's disease. J. Med. Chem. 56, 6775–6791 (2013).
Balasubramanian, S. et al. A novel histone deacetylase 8 (HDAC8)-specific inhibitor PCI-34051 induces apoptosis in T-cell lymphomas. Leukemia 22, 1026–1034 (2008).
Suzuki, T. et al. Rapid discovery of highly potent and selective inhibitors of histone deacetylase 8 using click chemistry to generate candidate libraries. J. Med. Chem. 55, 9562–9575 (2012).
Saha, A. et al. Synthesis and biological evaluation of a targeted DNA-binding transcriptional activator with HDAC8 inhibitory activity. Bioorg. Med. Chem. 21, 4201–4209 (2013).
Olson, D. E. et al. Discovery of the first histone deacetylase 6/8 dual inhibitors. J. Med. Chem. 56, 4816–4820 (2013).
Ruefli, A. A. et al. The histone deacetylase inhibitor and chemotherapeutic agent suberoylanilide hydroxamic acid (SAHA) induces a cell-death pathway characterized by cleavage of Bid and production of reactive oxygen species. Proc. Natl Acad. Sci. USA 98, 10833–10838 (2001).
Rosato, R. R., Almenara, J. A. & Grant, S. The histone deacetylase inhibitor MS-275 promotes differentiation or apoptosis in human leukemia cells through a process regulated by generation of reactive oxygen species and induction of p21CIP1/WAF1 1. Cancer Res. 63, 3637–3645 (2003).
Butler, L. M. et al. The histone deacetylase inhibitor SAHA arrests cancer cell growth, up-regulates thioredoxin-binding protein-2, and down-regulates thioredoxin. Proc. Natl Acad. Sci. USA 99, 11700–11705 (2002).
Robert, C. & Rassool, F. V. HDAC inhibitors: roles of DNA damage and repair. Adv. Cancer Res. 116, 87–129 (2012).
Kachhap, S. K. et al. Downregulation of homologous recombination DNA repair genes by HDAC inhibition in prostate cancer is mediated through the E2F1 transcription factor. PLoS ONE 5, e11208 (2010).
Petruccelli, L. A. et al. Vorinostat induces reactive oxygen species and DNA damage in acute myeloid leukemia cells. PLoS ONE 6, e20987 (2011).
Conti, C. et al. Inhibition of histone deacetylase in cancer cells slows down replication forks, activates dormant origins, and induces DNA damage. Cancer Res. 70, 4470–4480 (2010).
Dai, Y., Rahmani, M., Dent, P. & Grant, S. Blockade of histone deacetylase inhibitor-induced RelA/p65 acetylation and NF-κB activation potentiates apoptosis in leukemia cells through a process mediated by oxidative damage, XIAP downregulation, and c-Jun N-terminal kinase 1 activation. Mol. Cell. Biol. 25, 5429–5444 (2005).
Chen, C. S. et al. Histone deacetylase inhibitors sensitize prostate cancer cells to agents that produce DNA double-strand breaks by targeting Ku70 acetylation. Cancer Res. 67, 5318–5327 (2007).
Newbold, A., Salmon, J., Stanley, K. & Johnstone, R. The role of p21waf1/cip1 and p27Kip1 in HDACi-mediated tumor cell death and cell cycle arrest. Oncogene http://dx.doi.org/10.1038/onc.2013.482 (2013).
Lindemann, R. K., Gabrielli, B. & Johnstone, R. W. Histone-deacetylase inhibitors for the treatment of cancer. Cell Cycle 3, 779–788 (2004).
Gabrielli, B. & Brown, M. Histone deacetylase inhibitors disrupt the mitotic spindle assembly checkpoint by targeting histone and nonhistone proteins. Adv. Cancer Res. 116, 1–37 (2012).
Qiu, L. et al. Histone deacetylase inhibitors trigger a G2 checkpoint in normal cells that is defective in tumor cells. Mol. Biol. Cell 11, 2069–2083 (2000).
Munro, J., Barr, N. I., Ireland, H., Morrison, V. & Parkinson, E. K. Histone deacetylase inhibitors induce a senescence-like state in human cells by a p16-dependent mechanism that is independent of a mitotic clock. Exp. Cell Res. 295, 525–538 (2004).
Rebbaa, A., Zheng, X., Chu, F. & Mirkin, B. L. The role of histone acetylation versus DNA damage in drug-induced senescence and apoptosis. Cell Death Differ. 13, 1960–1967 (2006).
Place, R. F., Noonan, E. J. & Giardina, C. HDACs and the senescent phenotype of WI-38 cells. BMC Cell Biol. 6, 37 (2005).
Terao, Y. et al. Sodium butyrate induces growth arrest and senescence-like phenotypes in gynecologic cancer cells. Int. J. Cancer 94, 257–267 (2001).
Pazolli, E. et al. Chromatin remodeling underlies the senescence-associated secretory phenotype of tumor stromal fibroblasts that supports cancer progression. Cancer Res. 72, 2251–2261 (2012).
Ablain, J. & de The, H. Revisiting the differentiation paradigm in acute promyelocytic leukemia. Blood 117, 5795–5802 (2011).
Gottlicher, M. et al. Valproic acid defines a novel class of HDAC inhibitors inducing differentiation of transformed cells. EMBO J. 20, 6969–6978 (2001).
Leiva, M. et al. Valproic acid induces differentiation and transient tumor regression, but spares leukemia-initiating activity in mouse models of APL. Leukemia 26, 1630–1637 (2012).
Lin, R. J. et al. Role of the histone deacetylase complex in acute promyelocytic leukaemia. Nature 391, 811–814 (1998).
Fredly, H. et al. The combination of valproic acid, all-trans retinoic acid and low-dose cytarabine as disease-stabilizing treatment in acute myeloid leukemia. Clin. Epigenet. 5, 13 (2013).
Cimino, G. et al. Sequential valproic acid/all-trans retinoic acid treatment reprograms differentiation in refractory and high-risk acute myeloid leukemia. Cancer Res. 66, 8903–8911 (2006).
Lee, Y. J. et al. Molecular mechanism of SAHA on regulation of autophagic cell death in tamoxifen-resistant MCF-7 breast cancer cells. Int. J. Med. Sci. 9, 881–893 (2012).
Robert, T. et al. HDACs link the DNA damage response, processing of double-strand breaks and autophagy. Nature 471, 74–79 (2011).
Shao, Y., Gao, Z., Marks, P. A. & Jiang, X. Apoptotic and autophagic cell death induced by histone deacetylase inhibitors. Proc. Natl Acad. Sci. USA 101, 18030–18035 (2004).
Liu, Y. L. et al. Autophagy potentiates the anti-cancer effects of the histone deacetylase inhibitors in hepatocellular carcinoma. Autophagy 6, 1057–1065 (2010).
Gammoh, N. et al. Role of autophagy in histone deacetylase inhibitor-induced apoptotic and nonapoptotic cell death. Proc. Natl Acad. Sci. USA 109, 6561–6565 (2012).
Dupere-Richer, D. et al. Vorinostat-induced autophagy switches from a death-promoting to a cytoprotective signal to drive acquired resistance. Cell Death Dis. 4, e486 (2013).
Song, W. et al. HDAC inhibition by LBH589 affects the phenotype and function of human myeloid dendritic cells. Leukemia 25, 161–168 (2011).
Ning, Z. Q. et al. Chidamide (CS055/HBI-8000): a new histone deacetylase inhibitor of the benzamide class with antitumor activity and the ability to enhance immune cell-mediated tumor cell cytotoxicity. Cancer Chemother. Pharmacol. 69, 901–909 (2012).
Murakami, T. et al. Transcriptional modulation using HDACi depsipeptide promotes immune cell-mediated tumor destruction of murine B16 melanoma. J. Invest. Dermatol. 128, 1506–1516 (2008).
Northrop, J. K., Wells, A. D. & Shen, H. Cutting edge: chromatin remodeling as a molecular basis for the enhanced functionality of memory CD8 T cells. J. Immunol. 181, 865–868 (2008).
Fann, M. et al. Histone acetylation is associated with differential gene expression in the rapid and robust memory CD8+ T-cell response. Blood 108, 3363–3370 (2006).
Shen, L. et al. Class I histone deacetylase inhibitor entinostat suppresses regulatory T cells and enhances immunotherapies in renal and prostate cancer models. PLoS ONE 7, e30815 (2012).
Bridle, B. W. et al. HDAC inhibition suppresses primary immune responses, enhances secondary immune responses, and abrogates autoimmunity during tumor immunotherapy. Mol. Ther. 21, 887–894 (2013).
Shen, L. & Pili, R. Class I histone deacetylase inhibition is a novel mechanism to target regulatory T cells in immunotherapy. Oncoimmunology 1, 948–950 (2012).
Villagra, A., Sotomayor, E. M. & Seto, E. Histone deacetylases and the immunological network: implications in cancer and inflammation. Oncogene 29, 157–173 (2010).
Cantley, M. D. & Haynes, D. R. Epigenetic regulation of inflammation: progressing from broad acting histone deacetylase (HDAC) inhibitors to targeting specific HDACs. Inflammopharmacology 21, 301–307 (2013).
Schmudde, M., Friebe, E., Sonnemann, J., Beck, J. F. & Broker, B. M. Histone deacetylase inhibitors prevent activation of tumour-reactive NK cells and T cells but do not interfere with their cytolytic effector functions. Cancer Lett. 295, 173–181 (2010).
Reddy, P. et al. Histone deacetylase inhibitor suberoylanilide hydroxamic acid reduces acute graft-versus-host disease and preserves graft-versus-leukemia effect. Proc. Natl Acad. Sci. USA 101, 3921–3926 (2004).
Reddy, P. et al. Histone deacetylase inhibition modulates indoleamine 2,3-dioxygenase-dependent DC functions and regulates experimental graft-versus-host disease in mice. J. Clin. Invest. 118, 2562–2573 (2008). This study provides preclinical evidence that HDAC inhibitors may be effective in immune-based disorders.
Kwon, H. J., Kim, M. S., Kim, M. J., Nakajima, H. & Kim, K. W. Histone deacetylase inhibitor FK228 inhibits tumor angiogenesis. Int. J. Cancer 97, 290–296 (2002).
Williams, R. J. Trichostatin A, an inhibitor of histone deacetylase, inhibits hypoxia-induced angiogenesis. Expert Opin. Investig. Drugs 10, 1571–1573 (2001).
Deroanne, C. F. et al. Histone deacetylases inhibitors as anti-angiogenic agents altering vascular endothelial growth factor signaling. Oncogene 21, 427–436 (2002).
Ellis, L., Hammers, H. & Pili, R. Targeting tumor angiogenesis with histone deacetylase inhibitors. Cancer Lett. 280, 145–153 (2009).
Ryu, H. et al. Histone deacetylase inhibitors prevent oxidative neuronal death independent of expanded polyglutamine repeats via an Sp1-dependent pathway. Proc. Natl Acad. Sci. USA 100, 4281–4286 (2003).
Langley, B. et al. Pulse inhibition of histone deacetylases induces complete resistance to oxidative death in cortical neurons without toxicity and reveals a role for cytoplasmic p21(waf1/cip1) in cell cycle-independent neuroprotection. J. Neurosci. 28, 163–176 (2008).
Leng, Y. & Chuang, D. M. Endogenous α-synuclein is induced by valproic acid through histone deacetylase inhibition and participates in neuroprotection against glutamate-induced excitotoxicity. J. Neurosci. 26, 7502–7512 (2006).
Saunders, K. O., Freel, S. A., Overman, R. G., Cunningham, C. K. & Tomaras, G. D. Epigenetic regulation of CD8+ T-lymphocyte mediated suppression of HIV-1 replication. Virology 405, 234–242 (2010).
Acknowledgements
R.W.J. is a Principal Research Fellow of the National Health and Medical Research Council (NHMRC) of Australia and his research is supported by the NHMRC Program and Project Grants, Cancer Council Victoria, the Leukaemia Foundation of Australia and the Victorian Cancer Agency. The research of K.J.F. was supported by a postdoctoral fellowship from Cancer Council Victoria. The authors thank S. Mincci for helpful comments and advice.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The Johnstone laboratory has received grant funding from Novartis and Merck for studies using panobinostat and vorinostat, respectively. R.W.J. has received speaker's honoraria from Novartis. K.J.F. declares no competing interests.
Related links
FURTHER INFORMATION
Supplementary information
Supplementary information S1 (table)
Classical Zn2+-dependent HDACs (PDF 107 kb)
Supplementary information S2 (table)
Broad specificity HDAC inhibitors (PDF 110 kb)
Glossary
- Histone deacetylases
-
(HDACs). A family of 18 proteins in humans, consisting of class I proteins (HDAC1, HDAC2, HDAC3 and HDAC8), class IIa proteins (HDAC4, HDAC5, HDAC7 and HDAC9), class IIb proteins (HDAC6 and HDAC10), class III proteins (sirtuins 1–7) and class IV proteins (HDAC11). These enzymes remove acetyl groups from lysine on histones and other proteins.
- Epigenetic modifications
-
Reversible, heritable genetic changes that occur without changes in DNA sequence.
- Chromatin remodelling
-
An alteration in chromatin structure that affects the nuclease sensitivity of a region of chromatin. Accomplished by covalent modification of histones and/or the action of ATP-dependent remodelling complexes.
- Epigenetic writers
-
Enzymes such as histone acetyltransferases, methylases and kinases, which covalently modify the amino-terminal 'tails' of histone proteins, and DNA methyltransferases, which modify the DNA itself.
- Epigenetic erasers
-
Enzymes such as histone deacetylases, phosphatases, deubiquitylases and demethylases that reverse covalent modifications within the amino-terminal 'tails' of histone proteins.
- Epigenetic readers
-
Proteins containing chromodomains, bromodomains, Tudor domains and DNA methyl-binding domains that recognize specific histone marks and recruit other chromatin modifiers and remodelling proteins to alter chromatin architecture and function.
- Tauopathies
-
A class of neurodegenerative diseases associated with the pathological aggregation of tau protein in the human brain.
- Regulatory T cell
-
(TReg cell). A T cell type that suppresses the immune responses of other cells to maintain tolerance to self-antigens and abrogate autoimmune disease.
- Polypharmacological molecules
-
Single drug molecules that bind to multiple targets.
- Oncogene addiction
-
The reliance of tumours on a single dominant oncogene for growth and survival, so that inhibition of this specific oncogene is sufficient to halt the neoplastic phenotype.
Rights and permissions
About this article
Cite this article
Falkenberg, K., Johnstone, R. Histone deacetylases and their inhibitors in cancer, neurological diseases and immune disorders. Nat Rev Drug Discov 13, 673–691 (2014). https://doi.org/10.1038/nrd4360
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrd4360
This article is cited by
-
HDAC1/2 control mesothelium/ovarian cancer adhesive interactions impacting on Talin-1-α5β1-integrin-mediated actin cytoskeleton and extracellular matrix protein remodeling
Journal of Experimental & Clinical Cancer Research (2024)
-
Dysregulation of histone deacetylases in ocular diseases
Archives of Pharmacal Research (2024)
-
Epigenetics as a versatile regulator of fibrosis
Journal of Translational Medicine (2023)
-
AF9 targets acetyl-modified STAT6 to diminish purine metabolism and accelerate cell apoptosis during metastasis
Cell Death & Differentiation (2023)
-
Histone 4 lysine 5/12 acetylation enables developmental plasticity of Pristionchus mouth form
Nature Communications (2023)