Abstract
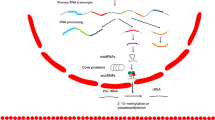
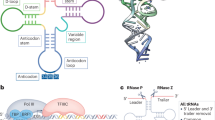
Small nucleolar RNAs (snoRNAs) have long been considered important but unglamorous elements in the production of the protein synthesis machinery of the cell. Recently, however, several independent lines of evidence have indicated that these non-coding RNAs might have crucial roles in controlling cell behaviour, and snoRNA dysfunction could consequently contribute to oncogenesis in previously unsuspected ways.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Lestrade, L. & Weber, M. J. snoRNA-LBME-db, a comprehensive database of human H/ACA and C/D box snoRNAs. Nucleic Acids Res. 34, D158–D162 (2006).
Kiss-Laszlo, Z., Henry, Y., Bachellerie, J., Caizergues-Ferrer, M. & Kiss, T. Site-specific ribose methylation of preribosomal RNA: a novel function for small nucleolar RNAs. Cell 85, 1077–1088 (1996).
Tollervey, D. & Kiss, T. Function and synthesis of small nucleolar RNAs. Curr. Opin. Cell Biol. 9, 337–342 (1997).
Weinstein L. B., Steitz J. A. Guided tours: from precursor snoRNA to functional snoRNP. Curr. Opin. Cell Biol. 11, 378–384 (1999).
Mattick, J. & Makunin, I. Non-coding RNA. Hum. Mol. Genet. 15, R17–R29 (2006).
Decatur, W. & Fournier, M. rRNA modifications and ribosome function. Trends Biochem. Sci. 27, 344–351 (2002).
Tycowski K. T., Shu M., Kukoyi A., Steitz J. A. A conserved WD40 protein binds the cajal body localization signal of scaRNP particles. Mol. Cell 34, 47–57 (2009).
Tycowski, K. T., Shu, M. D. & Steitz, J. A. A mammalian gene with introns instead of exons generating stable RNA products. Nature 379, 464–466 (1996).
Smith, C. M. & Steitz, J. A. Classification of gas5 as a multi-small-nucleolar-RNA (snoRNA) host gene and a member of the 5 ′-terminal oligopyrimidine gene family reveals common features of snoRNA host genes. Mol. Cell. Biol. 18, 6897–6909 (1998).
Dieci, G., Preti, M. & Montanini, B. Eukaryotic snoRNAs: a paradigm for gene expression flexibility. Genomics 94, 83–88 (2009).
Hoeppner, M. P., White, S., Jeffares, D. C. & Poole, A. M. Evolutionarily stable association of intronic snornas and micrornas with their host genes. Genome Biol. Evol. 1, 420–428 (2009).
Kino, T., Hurt, D. E., Ichijo, T., Nader, N. & Chrousos, G. P. Noncoding RNA Gas5 is a growth arrest- and starvation-associated repressor of the glucocorticoid receptor. Sci. Signal. 3, ra8 (2010).
Askarian-Amiri, M. E. et al. SNORD-host RNA Zfas1 is a regulator of mammary development and a potential marker for breast cancer. RNA 17, 878–891 (2011).
Williams, G. T. et al. Isolation of genes controlling apoptosis through their effects on cell survival. Gene Ther. Mol. Biol. 10B, 255–261 (2006).
Mourtada-Maarabouni, M., Hedge, V. L., Kirkham, L., Farzaneh, F. & Williams, G. T. Growth arrest in human T-cells is controlled by the non-coding RNA growth-arrest-specific transcript 5 (GAS5) J. Cell Sci. 121, 939–946 (2008).
Mourtada-Maarabouni, M., Pickard, M. R., Hedge, V. L., Farzaneh, F. & Williams, G. T. GAS5, a non-protein-coding RNA, controls apoptosis and is downregulated in breast cancer. Oncogene 28, 195–208 (2009).
Dong, X. et al. Implication of snoRNA U50 in human breast cancer. J. Genet. Genomics 36, 447–454 (2009).
Dong, X. et al. SnoRNA U50 is a candidate tumor-suppressor gene at 6q14.3 with a mutation associated with clinically significant prostate cancer. Hum. Mol. Genet. 17, 1031–1042 (2008).
Gee, H. E. et al. The small-nucleolar RNAs commonly used for microRNA normalisation correlate with tumour pathology and prognosis. Br. J. Cancer 104, 1168–1177 (2011).
Liao, J. et al. Small nucleolar RNA signatures as biomarkers for non-small-cell lung cancer. Mol. Cancer 9, 198 (2010).
Martens-Uzunova, E. S. et al. Diagnostic and prognostic signatures from the small non-coding RNA transcriptome in prostate cancer. Oncogene 18 Jul 2011 (doi:10.1038/onc.2011.304).
Scott, M. S. & Ono, M. From snoRNA to miRNA: dual function regulatory non-coding RNAs. Biochimie 93, 1987–1992 (2011).
Kishore, S. & Stamm, S. The snoRNA HBII-52 regulates alternative splicing of the serotonin receptor 2C. Science 311, 230–232 (2006).
Sahoo, T. et al. Prader-Willi phenotype caused by paternal deficiency for the HBII-85C/D box small nucleolar RNA cluster. Nature Genet. 40, 719–721 (2008).
Kishore, S. et al. The snoRNA MBII-52 (SNORD 115) is processed into smaller RNAs and regulates alternative splicing. Hum. Mol. Genet. 19, 1153–1164 (2010).
Tanaka, R. et al. Intronic U50 small-nucleolar-RNA (snoRNA) host gene of no protein-coding potential is mapped at the chromosome breakpoint t(3;6)(q27;q15) of human B-cell lymphoma. Genes Cells 5, 277–287 (2000).
Mei, Y.-P. et al. Small nucleolar RNA 42 acts as an oncogene in lung tumorigenesis. Oncogene 10 Oct 2011 (doi:10.1038/onc.2011.449).
Testa, J. et al. Advances in the analysis of chromosome alterations in human lung carcinomas. Cancer Genet. Cytogenet. 95, 20–32 (1997).
Michel, C. I. et al. Small Nucleolar RNAs U32a, U33, and U35a are critical mediators of metabolic stress. Cell Metab. 14, 33–44 (2011).
Montanaro, L. Dyskerin and cancer: more than telomerase. The defect in mRNA translation helps in explaining how a proliferative defect leads to cancer. J. Pathol. 222, 345–349 (2010).
Batista, L. F. Z. et al. Telomere shortening and loss of self-renewal in dyskeratosis congenita induced pluripotent stem cells. Nature 474, 399–402 (2011).
Hirose, T. & Steitz, J. Position within the host intron is critical for efficient processing of box C/D snoRNAs in mammalian cells. Proc. Natl Acad. Sci. USA 98, 12914–12919 (2001).
Meyuhas, O. Synthesis of the translational apparatus is regulated at the translational level. Eur. J. Biochem. 267, 6321–6330 (2000).
Yamashita, A. et al. SMG-8 and SMG-9, two novel subunits of the SMG-1 complex, regulate remodeling of the mRNA surveillance complex during nonsense-mediated mRNA decay. Genes Dev. 23, 1091–1105 (2009).
Schneider, C., King, R. M. & Philipson, L. Genes specifically expressed at growth arrest of mammalian-cells. Cell 54, 787–793 (1988).
Mourtada-Maarabouni, M., Hasan, A. M., Farzaneh, F. & Williams, G. T. Inhibition of human T-cell proliferation by mammalian target of rapamycin (mTOR) antagonists requires noncoding RNA growth-arrest-specific transcript 5 (GAS5). Mol. Pharmacol. 78, 19–28 (2010).
Williams, G. T., Mourtada-Maarabouni, M. & Farzaneh, F. A critical role for non-coding RNA GAS5 in growth arrest and rapamycin inhibition in human T-lymphocytes. Biochem. Soc. Trans. 39, 482–486 (2011).
Meyuhas, O. & Dreazen, A. Ribosomal protein S6 kinase: from TOP mRNAs to cell size. Prog. Mol. Biol. Transl. Sci. 90, 109–153 (2009).
Loreni, F., Iadevaia, V., Tino, E., Caldarola, S. & Amaldi, F. RACK1 mRNA translation is regulated via a rapamycin-sensitive pathway and coordinated with ribosomal protein synthesis. FEBS Lett. 579, 5517–5520 (2005).
Chitpatima, S., Makrides, S., Bandyopadhyay, R. & Brawerman, G. Nucleotide-sequence of a major messenger-RNA for a 21 kilodalton polypeptide that is under translational control in mouse-tumor cells. Nucleic Acids Res. 16, 2350–2350 (1988).
Michiels, L., Vanderrauwelaert, E., Vanhasselt, F., Kas, K. & Merregaert, J. Fau cDNA encodes a ubiquitin-like-S30 fusion protein and is expressed as an antisense sequence in the finkel-biskis-reilly murine sarcoma-virus. Oncogene 8, 2537–2546 (1993).
Mourtada-Maarabouni, M., Kirkham, L., Farzaneh, F. & Williams, G. T. Functional expression cloning reveals a central role for the receptor for activated protein kinase C 1 (RACK1) in T cell apoptosis. J. Leukoc. Biol. 78, 503–514 (2005).
Mourtada-Maarabouni, M., Kirkham, L., Farzaneh, F. & Williams, G. T. Regulation of apoptosis by fau revealed by functional expression cloning and antisense expression. Oncogene 23, 9419–9426 (2004).
Telerman, A. & Amson, R. The molecular programme of tumour reversion: the steps beyond malignant transformation. Nature Rev. Cancer 9, 206–215 (2009).
Dean, J. L. E., Sully, G., Clark, A. R. & Saklatvala, J. The involvement of AU-rich element-binding proteins in p38 mitogen-activated protein kinase pathway-mediated mRNA stabilisation. Cell. Signal. 16, 1113–1121 (2004).
Kakegawa, T. et al. Identification of AUF1 as a rapamycin-responsive binding protein to the 5′-terminal oligopyrimidine element of mRNAs. Arch. Biochem. Biophys. 465, 274–281 (2007).
Schultze, F. C., Petrova, D. T., Oellerich, M., Armstrong, V. W. & Asif, A. R. Differential proteome and phosphoproteome signatures in human T-lymphoblast cells induced by sirolimus. Cell Prolif. 43, 396–404 (2010).
Shchors, K. et al. Cell death inhibiting RNA (CDIR) derived from a 3 '-untranslated region binds AUF1 and heat shock protein 27. J. Biol. Chem. 277, 47061–47072 (2002).
Ender, C. et al. A human snoRNA with microRNA-like functions. Mol. Cell 32, 519–528 (2008).
Scott, M. S., Avolio, F., Ono, M., Lamond, A. I. & Barton, G. J. Human miRNA precursors with box H/ACA snoRNA features. PLoS Comput. Biol. 5, e1000507 (2009).
Ono, M. et al. Identification of human miRNA precursors that resemble box C/D snoRNAs. Nucleic Acids Res. 39, 3879–3891 (2011).
Esquela-Kerscher, A. & Slack, F. Oncomirs - microRNAs with a role in cancer. Nature Rev. Cancer 6, 259–269 (2006).
Brameier, M., Herwig, A., Reinhardt, R., Walter, L. & Gruber, J. Human box C/D snoRNAs with miRNA like functions: expanding the range of regulatory RNAs. Nucleic Acids Res. 39, 675–686 (2011).
Xiao, J., Lin, H., Luo, X., Luo, X. & Wang, Z. miR-605 joins p53 network to form a p53:miR-605:Mdm2 positive feedback loop in response to stress. EMBO J. 30, 524–532 (2011).
Montanaro, L., Trere, D. & Derenzini, M. Nucleolus, ribosomes, and cancer. Am. J. Pathol. 173, 301–310 (2008).
Shiue, C.-N., Berkson, R. G. & Wright, A. P. H. c-Myc induces changes in higher order rDNA structure on stimulation of quiescent cells. Oncogene 28, 1833–1842 (2009).
Novina, C. D. & Chabner, B. A. RNA-directed therapy: the next step in the miRNA revolution. Oncologist 13, 1–3 (2008).
Liu, X., Song, W., Sun, T., Zhang, P. & Wang, J. Targeted delivery of antisense inhibitor of miRNA for antiangiogenesis therapy using cRGD-functionalized nanoparticles. Mol. Pharm. 8, 250–259 (2011).
Doniger, T., Michaeli, S. & Unger, R. Families of H/ACA ncRNA molecules in trypanosomatids. RNA Biol. 6, 370–374 (2009).
Liang, X., Liu, Q., Liu, Q., King, T. H. & Fournier, M. J. Strong dependence between functional domains in a dual-function snoRNA infers coupling of rRNA processing and modification events. Nucleic Acids Res. 38, 3376–3387 (2010).
Acknowledgements
The authors would like to thank Leukaemia and Lymphoma Research, Breast Cancer Campaign and the National Cancer Research Institute UK Prostate Cancer Collaborative for financial support and apologize to those authors whose work has not been cited because of space constraints. The authors also acknowledge financial support from the Department of Health via the National Institute for Health Research (NIHR) comprehensive Biomedical Research Centre award to Guy's & St Thomas' NHS Foundation Trust in partnership with King's College London and King's College Hospital NHS Foundation Trust. Work in F.F.'s laboratory is also supported by Rosetrees Trust, Elimination of Leukaemia Fund and the Experimental Cancer Medicine Centre at King's College London.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
Rights and permissions
About this article
Cite this article
Williams, G., Farzaneh, F. Are snoRNAs and snoRNA host genes new players in cancer?. Nat Rev Cancer 12, 84–88 (2012). https://doi.org/10.1038/nrc3195
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrc3195
This article is cited by
-
Changes in the small noncoding RNA transcriptome in osteosarcoma cells
Journal of Orthopaedic Surgery and Research (2023)
-
LncRNA SNHG4 promotes prostate cancer cell survival and resistance to enzalutamide through a let-7a/RREB1 positive feedback loop and a ceRNA network
Journal of Experimental & Clinical Cancer Research (2023)
-
Long noncoding RNA SNHG16 regulates TLR4-mediated autophagy and NETosis formation in alveolar hemorrhage associated with systemic lupus erythematosus
Journal of Biomedical Science (2023)
-
SNORA56-mediated pseudouridylation of 28 S rRNA inhibits ferroptosis and promotes colorectal cancer proliferation by enhancing GCLC translation
Journal of Experimental & Clinical Cancer Research (2023)
-
Involvement of the oncogenic small nucleolar RNA SNORA24 in regulation of p53 stability in colorectal cancer
Cell Biology and Toxicology (2023)