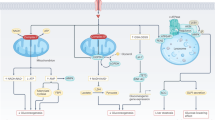
Key Points
-
Multiple molecular mechanisms, both intrinsic and extrinsic, converge to alter core cellular metabolism and provide support for the three basic needs of dividing cells: rapid ATP generation to maintain energy status; increased biosynthesis of macromolecules; and tightened maintenance of appropriate cellular redox status. Metabolic changes are a common feature of cancerous tissues, although it is unclear to what extent these metabolic changes are important in low-grade slow growing tumours.
-
The best characterized metabolic phenotype observed in tumour cells is the Warburg effect, which is a shift from ATP generation through oxidative phosphorylation to ATP generation through glycolysis, even under normal oxygen concentrations. This effect is regulated by the PI3K, hypoxia-indicible factor (HIF), p53, MYC and AMP-activated protein kinase (AMPK)–liver kinase B1 (LKB1) pathways.
-
Metabolic adaptation in tumours extends beyond the Warburg effect. It is becoming clear that alterations to metabolism balance the need of the cell for energy with its equally important need for macromolecular building blocks and maintenance of redox balance. To this end, a key molecule produced as a result of altered cancer metabolism is reduced nicotinamide adenine dinucleotide phosphate (NADPH), which functions as a cofactor and provides reducing power in many enzymatic reactions that are crucial for macromolecular biosynthesis. NADPH is also an antioxidant and forms part of the defence against reactive oxygen species (ROS) that are produced during rapid proliferation.
-
High levels of ROS can cause damage to macromolecules, which can induce senescence and apoptosis. Cells counteract the detrimental effects of ROS by producing antioxidant molecules, such as reduced glutathione (GSH) and thioredoxin (TRX). Several of these antioxidant systems, including GSH and TRX, rely on the reducing power of NADPH to maintain their activities.
-
In addition to the genetic changes that alter tumour cell metabolism, the abnormal tumour microenvironment — such as hypoxia, pH and low glucose concentrations — have a major role in determining the metabolic phenotype of tumour cells.
-
Mutations in oncogenes and tumour suppressor genes cause alterations to multiple intracellular signalling pathways that affect tumour cell metabolism and re-engineer it to allow enhanced survival and growth.
Abstract
Interest in the topic of tumour metabolism has waxed and waned over the past century of cancer research. The early observations of Warburg and his contemporaries established that there are fundamental differences in the central metabolic pathways operating in malignant tissue. However, the initial hypotheses that were based on these observations proved inadequate to explain tumorigenesis, and the oncogene revolution pushed tumour metabolism to the margins of cancer research. In recent years, interest has been renewed as it has become clear that many of the signalling pathways that are affected by genetic mutations and the tumour microenvironment have a profound effect on core metabolism, making this topic once again one of the most intense areas of research in cancer biology.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Stratton, M. R., Campbell, P. J. & Futreal, P. A. The cancer genome. Nature 458, 719–724 (2009).
The International Cancer Genome Consortium. International network of cancer genome projects. Nature 464, 993–998 (2010).
Parsons, D. W. et al. An integrated genomic analysis of human glioblastoma multiforme. Science 321, 1807–1812 (2008). Sequencing of the glioblastoma genome in which mutation of IDH1 was identified as a driver mutation.
Vander Heiden, M. G., Cantley, L. C. & Thompson, C. B. Understanding the Warburg effect: the metabolic requirements of cell proliferation. Science 324, 1029–1033 (2009). Provocative review advancing the concept that glycolytic metabolism supports biosynthetic pathways.
Newsholme, E. A., Crabtree, B. & Ardawi, M. S. The role of high rates of glycolysis and glutamine utilization in rapidly dividing cells. Biosci. Rep. 5, 393–400 (1985).
Tatum, J. L. et al. Hypoxia: importance in tumor biology, noninvasive measurement by imaging, and value of its measurement in the management of cancer therapy. Int. J. Radiat. Biol. 82, 699–757 (2006).
Warburg, O. On the origin of cancer cells. Science 123, 309–314 (1956).
Semenza, G. L. et al. 'The metabolism of tumours': 70 years later. Novartis Found. Symp. 240, 251–260; discussion 260–254 (2001).
Frezza, C. & Gottlieb, E. Mitochondria in cancer: not just innocent bystanders. Semin. Cancer Biol. 19, 4–11 (2009).
Weinhouse, S. The Warburg hypothesis fifty years later. Z. Krebsforsch. Klin. Onkol. Cancer Res. Clin. Oncol. 87, 115–126 (1976).
Funes, J. M. et al. Transformation of human mesenchymal stem cells increases their dependency on oxidative phosphorylation for energy production. Proc. Natl Acad. Sci. USA 104, 6223–6228 (2007).
Fogal, V. et al. Mitochondrial p32 protein is a critical regulator of tumor metabolism via maintenance of oxidative phosphorylation. Mol. Cell. Biol. 30, 1303–1318 (2010).
Weinberg, F. et al. Mitochondrial metabolism and ROS generation are essential for Kras-mediated tumorigenicity. Proc. Natl Acad. Sci. USA 107, 8788–8793 (2010).
Gatenby, R. A. & Gillies, R. J. Why do cancers have high aerobic glycolysis? Nature Rev. Cancer 4, 891–899 (2004).
Gillies, R. J., Robey, I. & Gatenby, R. A. Causes and consequences of increased glucose metabolism of cancers. J. Nucl. Med. 49 (Suppl. 2), 24S-42S (2008).
Gambhir, S. S. Molecular imaging of cancer with positron emission tomography. Nature Rev. Cancer 2, 683–693 (2002).
Gambhir, S. S. et al. A tabulated summary of the FDG PET literature. J. Nucl. Med. 42, 1S–93S (2001).
Jadvar, H., Alavi, A. & Gambhir, S. S. 18F-FDG uptake in lung, breast, and colon cancers: molecular biology correlates and disease characterization. J. Nucl. Med. 50, 1820–1827 (2009).
Czernin, J. & Phelps, M. E. Positron emission tomography scanning: current and future applications. Annu. Rev. Med. 53, 89–112 (2002).
Le, A. et al. Inhibition of lactate dehydrogenase A induces oxidative stress and inhibits tumor progression. Proc. Natl Acad. Sci. USA 107, 2037–2042 (2010).
Fantin, V. R., St-Pierre, J. & Leder, P. Attenuation of LDH-A expression uncovers a link between glycolysis, mitochondrial physiology, and tumor maintenance. Cancer Cell 9, 425–434 (2006).
Wong, K. K., Engelman, J. A. & Cantley, L. C. Targeting the PI3K signaling pathway in cancer. Curr. Opin. Genet. Dev. 20, 87–90 (2010).
Plas, D. R. & Thompson, C. B. Akt-dependent transformation: there is more to growth than just surviving. Oncogene 24, 7435–7442 (2005).
Elstrom, R. L. et al. Akt stimulates aerobic glycolysis in cancer cells. Cancer Res. 64, 3892–3899 (2004).
Fan, Y., Dickman, K. G. & Zong, W. X. Akt and c-Myc differentially activate cellular metabolic programs and prime cells to bioenergetic inhibition. J. Biol. Chem. 285, 7324–7333 (2010).
Robey, R. B. & Hay, N. Is Akt the “Warburg kinase”?-Akt-energy metabolism interactions and oncogenesis. Semin. Cancer Biol. 19, 25–31 (2009).
Khatri, S., Yepiskoposyan, H., Gallo, C. A., Tandon, P. & Plas, D. R. FOXO3a regulates glycolysis via transcriptional control of tumor suppressor TSC1. J. Biol. Chem. 285, 15960–15965 (2010).
Fang, M. et al. The ER UDPase ENTPD5 promotes protein N-glycosylation, the Warburg effect, and proliferation in the PTEN pathway. Cell 143, 711–724 (2010).
Guertin, D. A. & Sabatini, D. M. Defining the role of mTOR in cancer. Cancer Cell 12, 9–22 (2007).
Bertout, J. A., Patel, S. A. & Simon, M. C. The impact of O2 availability on human cancer. Nature Rev. Cancer 8, 967–975 (2008).
Inoki, K., Corradetti, M. N. & Guan, K. L. Dysregulation of the TSC-mTOR pathway in human disease. Nature Genet. 37, 19–24 (2005).
Kapitsinou, P. P. & Haase, V. H. The VHL tumor suppressor and HIF: insights from genetic studies in mice. Cell Death Differ. 15, 650–659 (2008).
Kaelin, W. G. The von Hippel-Lindau tumour suppressor protein: O2 sensing and cancer. Nature Rev. Cancer 8, 865–873 (2008).
Selak, M. A. et al. Succinate links TCA cycle dysfunction to oncogenesis by inhibiting HIF-α prolyl hydroxylase. Cancer Cell 7, 77–85 (2005).
King, A., Selak, M. A. & Gottlieb, E. Succinate dehydrogenase and fumarate hydratase: linking mitochondrial dysfunction and cancer. Oncogene 25, 4675–4682 (2006).
Semenza, G. L. HIF-1: upstream and downstream of cancer metabolism. Curr. Opin. Genet. Dev. 20, 51–56 (2010).
Papandreou, I., Cairns, R. A., Fontana, L., Lim, A. L. & Denko, N. C. HIF-1 mediates adaptation to hypoxia by actively downregulating mitochondrial oxygen consumption. Cell Metab. 3, 187–197 (2006).
Kim, J. W., Tchernyshyov, I., Semenza, G. L. & Dang, C. V. HIF-1-mediated expression of pyruvate dehydrogenase kinase: a metabolic switch required for cellular adaptation to hypoxia. Cell Metab. 3, 177–185 (2006). References 37 and 38 showed that HIF1 induces expression of PDK1, which limits the flow of pyruvate into the TCA cycle and decreases oxidative phosphorylation.
Lu, C. W., Lin, S. C., Chen, K. F., Lai, Y. Y. & Tsai, S. J. Induction of pyruvate dehydrogenase kinase-3 by hypoxia-inducible factor-1 promotes metabolic switch and drug resistance. J. Biol. Chem. 283, 28106–28114 (2008).
Cairns, R. A. et al. Pharmacologically increased tumor hypoxia can be measured by 18F-Fluoroazomycin arabinoside positron emission tomography and enhances tumor response to hypoxic cytotoxin PR-104. Clin. Cancer Res. 15, 7170–7174 (2009).
Michelakis, E. D., Webster, L. & Mackey, J. R. Dichloroacetate (DCA) as a potential metabolic-targeting therapy for cancer. Br. J. Cancer 99, 989–994 (2008).
Semenza, G. L. Defining the role of hypoxia-inducible factor 1 in cancer biology and therapeutics. Oncogene 29, 625–634 (2010).
Onnis, B., Rapisarda, A. & Melillo, G. Development of HIF-1 inhibitors for cancer therapy. J. Cell. Mol. Med. 13, 2780–2786 (2009).
Dang, C. V., Le, A. & Gao, P. MYC-induced cancer cell energy metabolism and therapeutic opportunities. Clin. Cancer Res. 15, 6479–6483 (2009).
Kim, J. W., Gao, P., Liu, Y. C., Semenza, G. L. & Dang, C. V. Hypoxia-inducible factor 1 and dysregulated c-Myc cooperatively induce vascular endothelial growth factor and metabolic switches hexokinase 2 and pyruvate dehydrogenase kinase 1. Mol. Cell. Biol. 27, 7381–7393 (2007).
Dang, C. V., Kim, J. W., Gao, P. & Yustein, J. The interplay between MYC and HIF in cancer. Nature Rev. Cancer 8, 51–56 (2008).
Li, F. et al. Myc stimulates nuclearly encoded mitochondrial genes and mitochondrial biogenesis. Mol. Cell. Biol. 25, 6225–6234 (2005).
Kuhajda, F. P. AMP-activated protein kinase and human cancer: cancer metabolism revisited. Int. J. Obes. 32 (Suppl. 4), S36–S41 (2008).
Shackelford, D. B. & Shaw, R. J. The LKB1-AMPK pathway: metabolism and growth control in tumour suppression. Nature Rev. Cancer 9, 563–575 (2009). A comprehensive review of AMPK and LKB1 in cancer metabolism.
Jones, R. G. et al. AMP-activated protein kinase induces a p53-dependent metabolic checkpoint. Mol. Cell 18, 283–293 (2005).
Jenne, D. E. et al. Peutz-Jeghers syndrome is caused by mutations in a novel serine threonine kinase. Nature Genet. 18, 38–43 (1998).
Ji, H. et al. LKB1 modulates lung cancer differentiation and metastasis. Nature 448, 807–810 (2007).
Wingo, S. N. et al. Somatic LKB1 mutations promote cervical cancer progression. PLoS ONE 4, e5137 (2009).
Wang, W. & Guan, K. L. AMP-activated protein kinase and cancer. Acta Physiol. 196, 55–63 (2009).
Libby, G. et al. New users of metformin are at low risk of incident cancer: a cohort study among people with type 2 diabetes. Diabetes Care 32, 1620–1625 (2009).
Anisimov, V. N. et al. Effect of metformin on life span and on the development of spontaneous mammary tumors in HER-2/neu transgenic mice. Exp. Gerontol. 40, 685–693 (2005).
Vousden, K. H. & Ryan, K. M. p53 and metabolism. Nature Rev. Cancer 9, 691–700 (2009).
Mathupala, S. P., Heese, C. & Pedersen, P. L. Glucose catabolism in cancer cells. The type II hexokinase promoter contains functionally active response elements for the tumor suppressor p53. J. Biol. Chem. 272, 22776–22780 (1997).
Bensaad, K. et al. TIGAR, a p53-inducible regulator of glycolysis and apoptosis. Cell 126, 107–120 (2006).
Stambolic, V. et al. Regulation of PTEN transcription by p53. Mol. Cell 8, 317–325 (2001).
Matoba, S. et al. p53 regulates mitochondrial respiration. Science 312, 1650–1653 (2006).
Almeida, R. et al. OCT-1 is over-expressed in intestinal metaplasia and intestinal gastric carcinomas and binds to, but does not transactivate, CDX2 in gastric cells. J. Pathol. 207, 396–401 (2005).
Jin, T. et al. Examination of POU homeobox gene expression in human breast cancer cells. Int. J. Cancer 81, 104–112 (1999).
Shakya, A. et al. Oct1 loss of function induces a coordinate metabolic shift that opposes tumorigenicity. Nature Cell Biol. 11, 320–327 (2009).
Mazurek, S., Boschek, C. B., Hugo, F. & Eigenbrodt, E. Pyruvate kinase type M2 and its role in tumor growth and spreading. Semin. Cancer Biol. 15, 300–308 (2005).
Mazurek, S., Zwerschke, W., Jansen-Durr, P. & Eigenbrodt, E. Metabolic cooperation between different oncogenes during cell transformation: interaction between activated ras and HPV-16 E7. Oncogene 20, 6891–6898 (2001).
Zwerschke, W. et al. Modulation of type M2 pyruvate kinase activity by the human papillomavirus type 16 E7 oncoprotein. Proc. Natl Acad. Sci. USA 96, 1291–1296 (1999).
Christofk, H. R., Vander Heiden, M. G., Wu, N., Asara, J. M. & Cantley, L. C. Pyruvate kinase M2 is a phosphotyrosine-binding protein. Nature 452, 181–186 (2008).
Marshall, S., Bacote, V. & Traxinger, R. R. Discovery of a metabolic pathway mediating glucose-induced desensitization of the glucose transport system. Role of hexosamine biosynthesis in the induction of insulin resistance. J. Biol. Chem. 266, 4706–4712 (1991).
Christofk, H. R. et al. The M2 splice isoform of pyruvate kinase is important for cancer metabolism and tumour growth. Nature 452, 230–233 (2008). The first mechanistic investigation of PKM2 using experimental cancer models, confirming the hypothesis that PKM2 expression provides an advantage for tumour growth.
David, C. J., Chen, M., Assanah, M., Canoll, P. & Manley, J. L. HnRNP proteins controlled by c-Myc deregulate pyruvate kinase mRNA splicing in cancer. Nature 463, 364–368 (2009). Discovery and explanation of the connection between the oncoprotein MYC and PKM2 expression.
Schneider, J. et al. Tumor M2-pyruvate kinase in lung cancer patients: immunohistochemical detection and disease monitoring. Anticancer Res. 22, 311–318 (2002).
Cerwenka, H. et al. TUM2-PK (pyruvate kinase type tumor M2), CA19–19 and CEA in patients with benign, malignant and metastasizing pancreatic lesions. Anticancer Res. 19, 849–851 (1999).
Luftner, D. et al. Tumor type M2 pyruvate kinase expression in advanced breast cancer. Anticancer Res. 20, 5077–5082 (2000).
Nathan, C. & Ding, A. SnapShot: reactive oxygen intermediates (ROI). Cell 140, 951 (2010).
Budihardjo, I. I. et al. 6-Aminonicotinamide sensitizes human tumor cell lines to cisplatin. Clin. Cancer Res. 4, 117–130 (1998).
Mardis, E. R. et al. Recurring mutations found by sequencing an acute myeloid leukemia genome. N. Engl. J. Med. 361, 1058–1066 (2009).
Yan, H. et al. IDH1 and IDH2 mutations in gliomas. N. Engl. J. Med. 360, 765–773 (2009).
Gross, S. et al. Cancer-associated metabolite 2-hydroxyglutarate accumulates in acute myelogenous leukemia with isocitrate dehydrogenase 1 and 2 mutations. J. Exp. Med. 207, 339–344 (2010).
Ward, P. S. et al. The common feature of leukemia-associated IDH1 and IDH2 mutations is a neomorphic enzyme activity converting α-ketoglutarate to 2-hydroxyglutarate. Cancer Cell 17, 225–234 (2010).
Zhao, S. et al. Glioma-derived mutations in IDH1 dominantly inhibit IDH1 catalytic activity and induce HIF-1α. Science 324, 261–265 (2009).
Dang, L. et al. Cancer-associated IDH1 mutations produce 2-hydroxyglutarate. Nature 462, 739–744 (2009). Discovery that driver mutations in IDH1 cause the acquisition of a novel enzymatic activity and production of 2-HG.
Bleeker, F. E. et al. IDH1 mutations at residue p.R132 (IDH1R132) occur frequently in high-grade gliomas but not in other solid tumors. Hum. Mutat. 30, 7–11 (2009).
Kang, M. R. et al. Mutational analysis of IDH1 codon 132 in glioblastomas and other common cancers. Int. J. Cancer 125, 353–355 (2009).
Giannoni, E., Buricchi, F., Raugei, G., Ramponi, G. & Chiarugi, P. Intracellular reactive oxygen species activate Src tyrosine kinase during cell adhesion and anchorage-dependent cell growth. Mol. Cell. Biol. 25, 6391–6403 (2005).
Lee, S. R. et al. Reversible inactivation of the tumor suppressor PTEN by H2O2. J. Biol. Chem. 277, 20336–20342 (2002).
Cao, J. et al. Prdx1 inhibits tumorigenesis via regulating PTEN/AKT activity. EMBO J. 28, 1505–1517 (2009).
Gao, P. et al. HIF-dependent antitumorigenic effect of antioxidants in vivo. Cancer Cell 12, 230–238 (2007).
Bell, E. L., Emerling, B. M. & Chandel, N. S. Mitochondrial regulation of oxygen sensing. Mitochondrion 5, 322–332 (2005).
Ramsey, M. R. & Sharpless, N. E. ROS as a tumour suppressor? Nature Cell Biol. 8, 1213–1215 (2006).
Takahashi, A. et al. Mitogenic signalling and the p16INK4a-Rb pathway cooperate to enforce irreversible cellular senescence. Nature Cell Biol. 8, 1291–1297 (2006).
Garrido, C. et al. Mechanisms of cytochrome c release from mitochondria. Cell Death Differ. 13, 1423–1433 (2006).
Han, D., Antunes, F., Canali, R., Rettori, D. & Cadenas, E. Voltage-dependent anion channels control the release of the superoxide anion from mitochondria to cytosol. J. Biol. Chem. 278, 5557–5563 (2003).
Fruehauf, J. P. & Meyskens, F. L. Reactive oxygen species: a breath of life or death? Clin. Cancer Res. 13, 789–794 (2007).
Bae, Y. S. et al. Epidermal growth factor (EGF)-induced generation of hydrogen peroxide. Role in EGF receptor-mediated tyrosine phosphorylation. J. Biol. Chem. 272, 217–221 (1997).
Sundaresan, M., Yu, Z. X., Ferrans, V. J., Irani, K. & Finkel, T. Requirement for generation of H2O2 for platelet-derived growth factor signal transduction. Science 270, 296–299 (1995).
Vaughn, A. E. & Deshmukh, M. Glucose metabolism inhibits apoptosis in neurons and cancer cells by redox inactivation of cytochrome c. Nature Cell Biol. 10, 1477–1483 (2008). Evidence that redox control by the GSH system is important in neurons and cancer cells and that reduction of cytochrome c prevents apoptosis.
Schafer, Z. T. et al. Antioxidant and oncogene rescue of metabolic defects caused by loss of matrix attachment. Nature 461, 109–113 (2009).
Li, B., Gordon, G. M., Du, C. H., Xu, J. & Du, W. Specific killing of Rb mutant cancer cells by inactivating TSC2. Cancer Cell 17, 469–480 (2010). Evidence that inappropriate activation of growth and proliferation pathways can lead to excessive stress and cell death.
Ozcan, U. et al. Loss of the tuberous sclerosis complex tumor suppressors triggers the unfolded protein response to regulate insulin signaling and apoptosis. Mol. Cell 29, 541–551 (2008).
Nogueira, V. et al. Akt determines replicative senescence and oxidative or oncogenic premature senescence and sensitizes cells to oxidative apoptosis. Cancer Cell 14, 458–470 (2008).
Liu, Y. et al. MnSOD inhibits proline oxidase-induced apoptosis in colorectal cancer cells. Carcinogenesis 26, 1335–1342 (2005).
Liu, Y., Borchert, G. L., Surazynski, A. & Phang, J. M. Proline oxidase, a p53-induced gene, targets COX-2/PGE2 signaling to induce apoptosis and inhibit tumor growth in colorectal cancers. Oncogene 27, 6729–6737 (2008).
Liu, Y. et al. Proline oxidase functions as a mitochondrial tumor suppressor in human cancers. Cancer Res. 69, 6414–6422 (2009).
Budanov, A. V., Sablina, A. A., Feinstein, E., Koonin, E. V. & Chumakov, P. M. Regeneration of peroxiredoxins by p53-regulated sestrins, homologs of bacterial AhpD. Science 304, 596–600 (2004).
Yoon, K. A., Nakamura, Y. & Arakawa, H. Identification of ALDH4 as a p53-inducible gene and its protective role in cellular stresses. J. Hum. Genet. 49, 134–140 (2004).
Suzuki, S. et al. Phosphate-activated glutaminase (GLS2), a p53-inducible regulator of glutamine metabolism and reactive oxygen species. Proc. Natl Acad. Sci. USA 107, 7461–7466 (2010).
Chen, W. et al. Direct interaction between Nrf2 and p21Cip1/WAF1 upregulates the Nrf2-mediated antioxidant response. Mol. Cell 34, 663–673 (2009).
Trachootham, D., Alexandre, J. & Huang, P. Targeting cancer cells by ROS-mediated mechanisms: a radical therapeutic approach? Nature Rev. Drug Discov. 8, 579–591 (2009).
Trachootham, D. et al. Effective elimination of fludarabine-resistant CLL cells by PEITC through a redox-mediated mechanism. Blood 112, 1912–1922 (2008).
Trachootham, D. et al. Selective killing of oncogenically transformed cells through a ROS-mediated mechanism by β-phenylethyl isothiocyanate. Cancer Cell 10, 241–252 (2006).
Clements, C. M., McNally, R. S., Conti, B. J., Mak, T. W. & Ting, J. P. DJ-1, a cancer- and Parkinson's disease-associated protein, stabilizes the antioxidant transcriptional master regulator Nrf2. Proc. Natl Acad. Sci. USA 103, 15091–15096 (2006).
Gasser, T. et al. Genetic complexity and Parkinson's disease. Science 277, 388–389 (1997).
Kim, R. H. et al. Hypersensitivity of DJ-1-deficient mice to 1-methyl-4-phenyl-1,2,3,6-tetrahydropyrindine (MPTP) and oxidative stress. Proc. Natl Acad. Sci. USA 102, 5215–5220 (2005).
Kim, R. H. et al. DJ-1, a novel regulator of the tumor suppressor PTEN. Cancer Cell 7, 263–273 (2005). Discovery that PARK7 , a gene mutated in Parkinson's disease, is an oncogene.
Davidson, B. et al. Expression and clinical role of DJ-1, a negative regulator of PTEN, in ovarian carcinoma. Hum. Pathol. 39, 87–95 (2008).
Yuen, H. F. et al. DJ-1 could predict worse prognosis in esophageal squamous cell carcinoma. Cancer Epidemiol. Biomarkers Prev. 17, 3593–3602 (2008).
Lin, M. T. & Beal, M. F. Mitochondrial dysfunction and oxidative stress in neurodegenerative diseases. Nature 443, 787–795 (2006).
Bajaj, A., Driver, J. A. & Schernhammer, E. S. Parkinson's disease and cancer risk: a systematic review and meta-analysis. Cancer Causes Control 21, 697–707 (2010).
Coles, N. W. & Johnstone, R. M. Glutamine metabolism in Ehrlich ascites-carcinoma cells. Biochem. J. 83, 284–291 (1962).
Reitzer, L. J., Wice, B. M. & Kennell, D. Evidence that glutamine, not sugar, is the major energy source for cultured HeLa cells. J. Biol. Chem. 254, 2669–2676 (1979).
Eagle, H. Nutrition needs of mammalian cells in tissue culture. Science 122, 501–514 (1955).
Wise, D. R. et al. Myc regulates a transcriptional program that stimulates mitochondrial glutaminolysis and leads to glutamine addiction. Proc. Natl Acad. Sci. USA 105, 18782–18787 (2008).
Gao, P. et al. c-Myc suppression of miR-23a/b enhances mitochondrial glutaminase expression and glutamine metabolism. Nature 458, 762–765 (2009). Strong mechanistic evidence that MYC participates in promoting mitochondrial glutaminase activity.
DeBerardinis, R. J. et al. Beyond aerobic glycolysis: transformed cells can engage in glutamine metabolism that exceeds the requirement for protein and nucleotide synthesis. Proc. Natl Acad. Sci. USA 104, 19345–19350 (2007).
Gallagher, F. A., Kettunen, M. I., Day, S. E., Lerche, M. & Brindle, K. M. 13C MR spectroscopy measurements of glutaminase activity in human hepatocellular carcinoma cells using hyperpolarized 13C-labeled glutamine. Magn. Reson. Med. 60, 253–257 (2008).
Richards, N. G. & Kilberg, M. S. Asparagine synthetase chemotherapy. Annu. Rev. Biochem. 75, 629–654 (2006).
Reinert, R. B. et al. Role of glutamine depletion in directing tissue-specific nutrient stress responses to L-asparaginase. J. Biol. Chem. 281, 31222–31233 (2006).
Lunt, S. J., Chaudary, N. & Hill, R. P. The tumor microenvironment and metastatic disease. Clin. Exp. Metastasis 26, 19–34 (2009).
Bristow, R. G. & Hill, R. P. Hypoxia and metabolism. Hypoxia, DNA repair and genetic instability. Nature Rev. Cancer 8, 180–192 (2008).
Semenza, G. L. Regulation of cancer cell metabolism by hypoxia-inducible factor 1. Semin. Cancer Biol. 19, 12–16 (2009).
Denko, N. C. Hypoxia, HIF1 and glucose metabolism in the solid tumour. Nature Rev. Cancer 8, 705–713 (2008).
Wouters, B. G. & Koritzinsky, M. Hypoxia signalling through mTOR and the unfolded protein response in cancer. Nature Rev. Cancer 8, 851–864 (2008).
Koritzinsky, M. et al. Gene expression during acute and prolonged hypoxia is regulated by distinct mechanisms of translational control. EMBO J. 25, 1114–1125 (2006).
Rouschop, K. M. & Wouters, B. G. Regulation of autophagy through multiple independent hypoxic signaling pathways. Curr. Mol. Med. 9, 417–424 (2009).
Koumenis, C. et al. Regulation of protein synthesis by hypoxia via activation of the endoplasmic reticulum kinase PERK and phosphorylation of the translation initiation factor eIF2α. Mol. Cell. Biol. 22, 7405–7416 (2002).
Bi, M. et al. ER stress-regulated translation increases tolerance to extreme hypoxia and promotes tumor growth. EMBO J. 24, 3470–3481 (2005).
Romero-Ramirez, L. et al. XBP1 is essential for survival under hypoxic conditions and is required for tumor growth. Cancer Res. 64, 5943–5947 (2004).
Cairns, R., Papandreou, I. & Denko, N. Overcoming physiologic barriers to cancer treatment by molecularly targeting the tumor microenvironment. Mol. Cancer Res. 4, 61–70 (2006).
Moreno-Sanchez, R., Rodriguez-Enriquez, S., Marin-Hernandez, A. & Saavedra, E. Energy metabolism in tumor cells. FEBS J. 274, 1393–1418 (2007).
Yang, C. et al. Glioblastoma cells require glutamate dehydrogenase to survive impairments of glucose metabolism or Akt signaling. Cancer Res. 69, 7986–7993 (2009).
Sonveaux, P. et al. Targeting lactate-fueled respiration selectively kills hypoxic tumor cells in mice. J. Clin. Invest. 118, 3930–3942 (2008).
Acknowledgements
The authors would like to thank M. Saunders for scientific editing of the Review.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
Glossary
- Redox status
-
Balance of the reduced state versus the oxidized state of a biochemical system. This balance is influenced by the level of reactive oxygen and nitrogen species (ROS and RNS) relative to the capacity of antioxidant systems to eliminate ROS and RNS.
- Oxidative phosphorylation
-
Oxygen-dependent process coupling the oxidation of macromolecules and the electron transport chain with ATP synthesis. In eukaryotic cells, it occurs within the mitochondria and is a source of ROS production.
- Glycolysis
-
Oxygen-independent metabolism of glucose and other sugars into pyruvate to produce energy in the form of ATP and intermediate substrates for other metabolic pathways.
- Pentose phosphate pathway
-
PPP. Biochemical pathway converting glucose into substrates for nucleotide biosynthesis and redox control, such as ribose and NADPH. Owing to multiple connections to the glycolytic pathway, the PPP can operate in various modes to allow the production of NADPH and/or ribose as required.
- Macromolecular biosynthesis
-
Biochemical synthesis of the carbohydrates, nucleotides, proteins and lipids that make up cells and tissues. These pathways require energy, reducing power and appropriate substrates.
- Reduced nicotinamide adenine dinucleotide phosphate
-
NADPH. Cofactor that drives anabolic biochemical reactions and provides reducing capacity to combat oxidative stress.
- 2-hydroxyglutarate
-
2-HG. A dicarboxylic acid metabolite produced from αKG by the NADPH-dependent reaction of the mutated forms of IDH1 and IDH2. It is also produced at low levels by other enzymes.
- Parkinson's disease
-
A neurodegenerative disorder affecting the CNS, which is characterized by muscle rigidity and the onset of tremors.
- Amyotrophic lateral sclerosis
-
ALS. Also known as Lou Gehrigs disease; it occurs owing to the degeneration of the CNS and leads to the inability to control muscles and eventual muscle atrophy.
- Glutaminolysis
-
The catabolic metabolism of glutamine, which yields substrates that replenish the TCA cycle, produce GSH and supply building blocks for amino acid and nucleotide synthesis.
- Anapleurosis
-
Category of reactions that serve to replenish the intermediate substrates of an anabolic biochemical pathway, especially important in the TCA cycle.
Rights and permissions
About this article
Cite this article
Cairns, R., Harris, I. & Mak, T. Regulation of cancer cell metabolism. Nat Rev Cancer 11, 85–95 (2011). https://doi.org/10.1038/nrc2981
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrc2981
This article is cited by
-
Lithium carbonate revitalizes tumor-reactive CD8+ T cells by shunting lactic acid into mitochondria
Nature Immunology (2024)
-
Impact of serum eicosapentaenoic acid/arachidonic acid ratio on overall survival in lung cancer patients treated with pembrolizumab: a pilot study
Scientific Reports (2024)
-
Therapeutic targeting nudix hydrolase 1 creates a MYC-driven metabolic vulnerability
Nature Communications (2024)
-
Antiproliferative, antimigratory, and apoptotic effects of diffractaic and vulpinic acids as thioredoxin reductase 1 inhibitors on cervical cancer
Naunyn-Schmiedeberg's Archives of Pharmacology (2024)
-
Nanoscale metal–organic frameworks as smart nanocarriers for cancer therapy
Journal of Nanostructure in Chemistry (2024)