Key Points
-
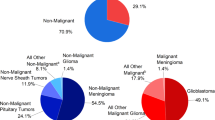
Malignant gliomas and medulloblastomas — the most common brain tumours affecting adults and children, respectively — remain responsible for a disproportionate level of morbidity and mortality among cancer patients.
-
The morphological histopathology traditionally used for the subclassification of these brain tumour variants is gradually giving way to more molecularly grounded criteria that better reflect the underlying biology.
-
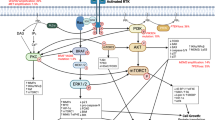
Recent integrated genomics has further implicated specific molecular networks in the pathogenesis of gliomas and medulloblastomas. These most prominently include receptor tyrosine kinase (RTK) signalling through the Ras–MAPK and PI3K–AKT–mTOR pathways, Wnt signalling and sonic hedgehog (SHH) signalling, along with the cell cycle-regulating RB and p53 pathways.
-
Expression analysis has recently defined transcriptional subclasses for both malignant gliomas and medulloblastomas that seem to be driven by distinct abnormalities in core signalling pathways. Such findings suggest that tumours in a particular molecular subgroup would preferentially respond to different targeted therapies.
-
Malignant gliomas and medulloblastomas also exhibit heterogeneity at the cellular level, with subpopulations of tumour cells harbouring stem-like properties rendering them more resistant to therapy. Such stem-like pools seem to reside in specialized microenvironments that actively maintain their biological characteristics.
-
Treatment challenges posed by malignant gliomas and medulloblastomas remain considerable, and many derive from the molecular and cellular heterogeneity inherent to these tumour variants. They include innate and acquired resistance and the obstacle to effective drug delivery posed by the blood–brain barrier.
Abstract
Malignant brain tumours continue to be the cause of a disproportionate level of morbidity and mortality across a wide range of individuals. The most common variants in the adult and paediatric populations — malignant glioma and medulloblastoma, respectively — have been the subject of increasingly intensive research over the past two decades that has led to considerable advances in the understanding of their basic biology and pathogenesis. This Review summarizes these developments in the context of the evolving notion of molecular pathology and discusses the implications that this work has on the design of new treatment regimens.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Stupp, R. et al. Radiotherapy plus concomitant and adjuvant temozolomide for glioblastoma. N. Engl. J. Med. 352, 987–996 (2005).
Wen, P. Y. & Kesari, S. Malignant gliomas in adults. N. Engl. J. Med. 359, 492–507 (2008).
Gilbertson, R. J. Medulloblastoma: signalling a change in treatment. Lancet Oncol. 5, 209–218 (2004).
Bailey, P. & Cushing, H. A Classification of Tumoursof the Glioma Group on a Histogenic Basis (J. B Lippincott, Philadelphia, 1926).
Kernohan, J. W., Mabon, R. F. & Svien, H. J. A simplified classification of gliomas. Proc. Staff Meetings Mayo Clinic 24, 71–74 (1949).
Ringertz, N. Grading of gliomas. Acta Pathol Microbiol Scand. 27, 51–65 (1950).
Zülch, K. J. (ed.) Histological typing of tumours of the central nervous system (World Health Organization, Switzerland, 1979).
Louis, D. N. et al. The 2007 WHO classification of tumours of the central nervous system. Acta Neuropathol. 114, 97–109 (2007). This paper establishes the current morphological and molecular criteria for the subclassification of brain tumours.
Dropcho, E. J. & Soong, S. J. The prognostic impact of prior low grade histology in patients with anaplastic gliomas: a case-control study. Neurology 47, 684–690 (1996).
Ohgaki, H. & Kleihues, P. Population-based studies on incidence, survival rates, and genetic alterations in astrocytic and oligodendroglial gliomas. J. Neuropathol. Exp. Neurol. 64, 479–489 (2005).
Ohgaki, H. et al. A case history of glioma progression. Acta Neuropathol. 97, 525–532 (1999).
Scherer, H. J. Cerebral astrocytomas and their derivatives. Am. J. Cancer 40, 159–198 (1940).
Rutkowski, S. et al. Treatment of early childhood medulloblastoma by postoperative chemotherapy alone. N. Engl. J. Med. 352, 978–986 (2005).
Sure, U. et al. Staging, scoring and grading of medulloblastoma. A postoperative prognosis predicting system based on the cases of a single institute. Acta Neurochir. (Wien) 132, 59–65 (1995).
Brown, H. G. et al. “Large cell/anaplastic” medulloblastomas: a Pediatric Oncology Group Study. J. Neuropathol. Exp. Neurol. 59, 857–865 (2000).
Giangaspero, F. et al. Large-cell medulloblastomas. A distinct variant with highly aggressive behavior. Am. J. Surg. Pathol. 16, 687–693 (1992).
Giangaspero, F. et al. Stratification of medulloblastoma on the basis of histopathological grading. Acta Neuropathol. 112, 5–12 (2006).
McManamy, C. S. et al. Morphophenotypic variation predicts clinical behavior in childhood non-desmoplastic medulloblastomas. J. Neuropathol. Exp. Neurol. 62, 627–632 (2003).
Malkin, D. et al. Germ line p53 mutations in a familial syndrome of breast cancer, sarcomas, and other neoplasms. Science 250, 1233–1238 (1990).
Srivastava, S., Zou, Z. Q., Pirollo, K., Blattner, W. & Chang, E. H. Germ-line transmission of a mutated p53 gene in a cancer-prone family with Li-Fraumeni syndrome. Nature 348, 747–749 (1990).
Louis, D. N. The p53 gene and protein in human brain tumors. J. Neuropathol. Exp. Neurol. 53, 11–21 (1994).
Libermann, T. A. et al. Amplification, enhanced expression and possible rearrangement of EGF receptor gene in primary human brain tumours of glial origin. Nature 313, 144–147 (1985).
Libermann, T. A. et al. Expression of epidermal growth factor receptors in human brain tumors. Cancer Res. 44, 753–760 (1984).
Wong, A. J. et al. Structural alterations of the epidermal growth factor receptor gene in human gliomas. Proc. Natl Acad. Sci. USA 89, 2965–2969 (1992).
Frederick, L., Wang, X. Y., Eley, G. & James, C. D. Diversity and frequency of epidermal growth factor receptor mutations in human glioblastomas. Cancer Res. 60, 1383–1387 (2000).
Sugawa, N., Ekstrand, A. J., James, C. D. & Collins, V. P. Identical splicing of aberrant epidermal growth factor receptor transcripts from amplified rearranged genes in human glioblastomas. Proc. Natl Acad. Sci. USA 87, 8602–8606 (1990).
Cairncross, J. G. et al. Specific genetic predictors of chemotherapeutic response and survival in patients with anaplastic oligodendrogliomas. J. Natl Cancer Inst. 90, 1473–1479 (1998).
Reifenberger, G. et al. Molecular genetic analysis of oligodendroglial tumors shows preferential allelic deletions on 19q and 1p. Am. J. Pathol. 145, 1175–1190 (1994).
Smith, J. S. et al. Localization of common deletion regions on 1p and 19q in human gliomas and their association with histological subtype. Oncogene 18, 4144–4152 (1999).
Barbashina, V., Salazar, P., Holland, E. C., Rosenblum, M. K. & Ladanyi, M. Allelic losses at 1p36 and 19q13 in gliomas: correlation with histologic classification, definition of a 150-kb minimal deleted region on 1p36, and evaluation of CAMTA1 as a candidate tumor suppressor gene. Clin. Cancer Res. 11, 1119–1128 (2005).
Costello, J. F. et al. Cyclin-dependent kinase 6 (CDK6) amplification in human gliomas identified using two-dimensional separation of genomic DNA. Cancer Res. 57, 1250–1254 (1997).
Henson, J. W. et al. The retinoblastoma gene is involved in the malignant progression of astrocytomas. Ann. Neurol. 36, 714–721 (1994).
Reifenberger, G., Reifenberger, J., Ichimura, K., Meltzer, P. S. & Collins, V. P. Amplification of multiple genes from chromosomal region 12q13–14 in human malignant gliomas: preliminary mapping of the amplicons shows preferential involvement of CDK4, SAS, and MDM2. Cancer Res. 54, 4299–4303 (1994).
Watanabe, T. et al. Concurrent inactivation of RB1 and TP53 pathways in anaplastic oligodendroglioma. J. Neuropathol. Exp. Neurol. 60, 1181–1190 (2001).
Shete, S. et al. Genome-wide association study identifies five susceptibility loci for glioma. Nature Genet. 41, 899–904 (2009).
Wrensch, M. et al. Variants in the CDKN2B and RTEL1 regions are associated with high-grade glioma susceptibility. Nature Genet. 41, 905–908 (2009). References 35 and 36 are large genome-wide association screens identifying low-penetrance susceptibility alleles that increase glioma risk.
Dai, C. et al. PDGF autocrine stimulation dedifferentiates cultured astrocytes and induces oligodendrogliomas and oligoastrocytomas from neural progenitors and astrocytes in vivo. Genes Dev. 15, 1913–1925 (2001).
Hu, X. et al. mTOR promotes survival and astrocytic characteristics induced by Pten/AKT signaling in glioblastoma. Neoplasia 7, 356–368 (2005).
Uhrbom, L., Nerio, E. & Holland, E. C. Dissecting tumor maintenance requirements using bioluminescence imaging of cell proliferation in a mouse glioma model. Nature Med. 10, 1257–1260 (2004).
Marino, S., Vooijs, M., van Der Gulden, H., Jonkers, J. & Berns, A. Induction of medulloblastomas in p53-null mutant mice by somatic inactivation of Rb in the external granular layer cells of the cerebellum. Genes Dev. 14, 994–1004 (2000).
Reilly, K. M., Loisel, D. A., Bronson, R. T., McLaughlin, M. E. & Jacks, T. Nf1; Trp53 mutant mice develop glioblastoma with evidence of strain-specific effects. Nature Genet. 26, 109–113 (2000).
Weiss, W. A. et al. Genetic determinants of malignancy in a mouse model for oligodendroglioma. Cancer Res. 63, 1589–1595 (2003).
Xiao, A., Wu, H., Pandolfi, P. P., Louis, D. N. & Van Dyke, T. Astrocyte inactivation of the pRb pathway predisposes mice to malignant astrocytoma development that is accelerated by PTEN mutation. Cancer Cell 1, 157–168 (2002).
Xiao, A. et al. Somatic induction of Pten loss in a preclinical astrocytoma model reveals major roles in disease progression and avenues for target discovery and validation. Cancer Res. 65, 5172–5180 (2005).
Zheng, H. et al. p53 and Pten control neural and glioma stem/progenitor cell renewal and differentiation. Nature 455, 1129–1133 (2008).
Zhu, Y. et al. Early inactivation of p53 tumor suppressor gene cooperating with NF1 loss induces malignant astrocytoma. Cancer Cell 8, 119–130 (2005).
Di Rocco, F., Carroll, R. S., Zhang, J. & Black, P. M. Platelet-derived growth factor and its receptor expression in human oligodendrogliomas. Neurosurgery 42, 341–346 (1998).
Westermark, B., Heldin, C. H. & Nister, M. Platelet-derived growth factor in human glioma. Glia 15, 257–263 (1995).
Clarke, I. D. & Dirks, P. B. A human brain tumor-derived PDGFR-α deletion mutant is transforming. Oncogene 22, 722–733 (2003).
Abounader, R. & Laterra, J. Scatter factor/hepatocyte growth factor in brain tumor growth and angiogenesis. Neuro Oncol. 7, 436–451 (2005).
Guha, A., Feldkamp, M. M., Lau, N., Boss, G. & Pawson, A. Proliferation of human malignant astrocytomas is dependent on Ras activation. Oncogene 15, 2755–2765 (1997).
Wang, H. et al. Analysis of the activation status of Akt, NFkB, and Stat3 in human diffuse gliomas. Lab. Invest. 84, 941–951 (2004).
James, C. D. et al. Clonal genomic alterations in glioma malignancy stages. Cancer Res. 48, 5546–5551 (1988).
Ding, H. et al. Oligodendrogliomas result from the expression of an activated mutant epidermal growth factor receptor in a RAS transgenic mouse astrocytoma model. Cancer Res. 63, 1106–1113 (2003).
Shih, A. H. et al. Dose-dependent effects of platelet-derived growth factor-B on glial tumorigenesis. Cancer Res. 64, 4783–4789 (2004).
Uhrbom, L. et al. Ink4a-Arf loss cooperates with KRas activation in astrocytes and neural progenitors to generate glioblastomas of various morphologies depending on activated Akt. Cancer Res. 62, 5551–5558 (2002).
Uhrbom, L., Hesselager, G., Nister, M. & Westermark, B. Induction of brain tumors in mice using a recombinant platelet-derived growth factor B-chain retrovirus. Cancer Res. 58, 5275–5279 (1998).
Wei, Q. et al. High-grade glioma formation results from postnatal pten loss or mutant epidermal growth factor receptor expression in a transgenic mouse glioma model. Cancer Res. 66, 7429–7437 (2006).
Zhu, H. et al. Oncogenic EGFR signaling cooperates with loss of tumor suppressor gene functions in gliomagenesis. Proc. Natl Acad. Sci. USA 106, 2712–2716 (2009).
Ding, H. et al. Astrocyte-specific expression of activated p21-ras results in malignant astrocytoma formation in a transgenic mouse model of human gliomas. Cancer Res. 61, 3826–3836 (2001).
Holland, E. C. et al. Combined activation of Ras and Akt in neural progenitors induces glioblastoma formation in mice. Nature Genet. 25, 55–57 (2000).
Kwon, C. H. et al. Pten haploinsufficiency accelerates formation of high-grade astrocytomas. Cancer Res. 68, 3286–3294 (2008).
Llaguno, S. A. et al. Malignant astrocytomas originate from neural stem/progenitor cells in a somatic tumor suppressor mouse model. Cancer Cell 15, 45–56 (2009).
Burton, E. C. et al. Genetic aberrations defined by comparative genomic hybridization distinguish long-term from typical survivors of glioblastoma. Cancer Res. 62, 6205–6210 (2002).
Hui, A. B., Lo, K. W., Yin, X. L., Poon, W. S. & Ng, H. K. Detection of multiple gene amplifications in glioblastoma multiforme using array-based comparative genomic hybridization. Lab. Invest. 81, 717–723 (2001).
Maher, E. A. et al. Marked genomic differences characterize primary and secondary glioblastoma subtypes and identify two distinct molecular and clinical secondary glioblastoma entities. Cancer Res. 66, 11502–11513 (2006).
Misra, A. et al. Array comparative genomic hybridization identifies genetic subgroups in grade 4 human astrocytoma. Clin. Cancer Res. 11, 2907–2918 (2005).
The Cancer Genome Atlas Research Network. Comprehensive genomic characterization defines human glioblastoma genes and core pathways. Nature 455, 1061–1068 (2008). This study reports large-scale, integrated genomic analysis revealing the central importance of certain core signalling pathways in the pathogenesis of malignant glioma.
Nigro, J. M. et al. Integrated array-comparative genomic hybridization and expression array profiles identify clinically relevant molecular subtypes of glioblastoma. Cancer Res. 65, 1678–1686 (2005).
Nishizaki, T. et al. Clinical evidence of distinct subgroups of astrocytic tumors defined by comparative genomic hybridization. Hum. Pathol. 31, 608–614 (2000).
Parsons, D. W. et al. An integrated genomic analysis of human glioblastoma multiforme. Science 321, 1807–1812 (2008). This study reached many of the same conclusions as reference 68 but also identified pathogenic mutations in IDH1.
Bredel, M. et al. A network model of a cooperative genetic landscape in brain tumors. JAMA 302, 261–275 (2009).
Yan, H. et al. IDH1 and IDH2 mutations in gliomas. N. Engl. J. Med. 360, 765–773 (2009).
Hartmann, C. et al. Type and frequency of IDH1 and IDH2 mutations are related to astrocytic and oligodendroglial differentiation and age: a study of 1,010 diffuse gliomas. Acta Neuropathol. 118, 469–474 (2009).
Sanson, M. et al. Isocitrate dehydrogenase 1 codon 132 mutation is an important prognostic biomarker in gliomas. J. Clin. Oncol. 27, 4150–4154 (2009).
Zhao, S. et al. Glioma-derived mutations in IDH1 dominantly inhibit IDH1 catalytic activity and induce HIF-1α. Science 324, 261–265 (2009). This study identifies a potential pathogenic mechanism for mutations in IDH proteins relating to a loss of enzymatic function.
Jensen, R. L. Brain tumor hypoxia: tumorigenesis, angiogenesis, imaging, pseudoprogression, and as a therapeutic target. J. Neurooncol. 92, 317–335 (2009).
Dang, L. et al. Cancer-associated IDH1 mutations produce 2-hydroxyglutarate. Nature 462, 739–744 (2009). This study identifies the abnormal production of 2-HG as a potential pathogenic mechanism for IDH mutations in glioma.
Mardis, E. R. et al. Recurring mutations found by sequencing an acute myeloid leukemia genome. N. Engl. J. Med. 361, 1058–1066 (2009).
Huse, J. T. et al. The PTEN-regulating microRNA miR-26a is amplified in high-grade glioma and facilitates gliomagenesis in vivo. Genes Dev. 23, 1327–1337 (2009).
Kim, H. et al. Integrative genome analysis reveals an oncomir/oncogene cluster regulating glioblastoma survivorship. Proc. Natl Acad. Sci. USA 107, 2183–2188 (2010).
Li, Y. et al. MicroRNA-34a inhibits glioblastoma growth by targeting multiple oncogenes. Cancer Res. 69, 7569–7576 (2009).
Novakova, J., Slaby, O., Vyzula, R. & Michalek, J. MicroRNA involvement in glioblastoma pathogenesis. Biochem. Biophys. Res. Commun. 386, 1–5 (2009).
Godard, S. et al. Classification of human astrocytic gliomas on the basis of gene expression: a correlated group of genes with angiogenic activity emerges as a strong predictor of subtypes. Cancer Res. 63, 6613–6625 (2003).
Nutt, C. L. et al. Gene expression-based classification of malignant gliomas correlates better with survival than histological classification. Cancer Res. 63, 1602–1607 (2003).
Li, A. et al. Unsupervised analysis of transcriptomic profiles reveals six glioma subtypes. Cancer Res. 69, 2091–2099 (2009).
Liang, Y. et al. Gene expression profiling reveals molecularly and clinically distinct subtypes of glioblastoma multiforme. Proc. Natl Acad. Sci. USA 102, 5814–5819 (2005).
Phillips, H. S. et al. Molecular subclasses of high-grade glioma predict prognosis, delineate a pattern of disease progression, and resemble stages in neurogenesis. Cancer Cell 9, 157–173 (2006). This study uses transcriptional profiling to subdivide malignant gliomas into distinct categories that differ in their neurobiological characteristics and clinical prognoses.
Horvath, S. et al. Analysis of oncogenic signaling networks in glioblastoma identifies ASPM as a molecular target. Proc. Natl Acad. Sci. USA 103, 17402–17407 (2006).
Brennan, C. et al. Glioblastoma subclasses can be defined by activity among signal transduction pathways and associated genomic alterations. PLoS ONE 4, e7752 (2009). This study uses an integrated genomics analysis of TCGA data coupled with proteomics in a second large tumour set to link glioma transcriptional subclasses with defined core signalling abnormalities.
Verhaak, R. G. W. et al. An integrated genomic analysis identifies clinically relevant subtypes of glioblastoma characterized by abnormalities in PDGFRA, IDH1, EGFR, and NF1. Cancer Cell 17, 98–110 (2009). This study uses TCGA data and an integrated genomics approach to associate glioma transcriptional subclasses with distinct abnormalities in specific genes.
Carro, M. S. et al. The transcriptional network for mesenchymal transformation of brain tumours. Nature 463, 318–325 (2010).
Entz-Werle, N. et al. Medulloblastoma: what is the role of molecular genetics? Expert Rev. Anticancer Ther. 8, 1169–1181 (2008).
Pietsch, T. et al. Medulloblastomas of the desmoplastic variant carry mutations of the human homologue of Drosophila patched. Cancer Res. 57, 2085–2088 (1997).
Raffel, C. et al. Sporadic medulloblastomas contain PTCH mutations. Cancer Res. 57, 842–845 (1997).
Reifenberger, J. et al. Missense mutations in SMOH in sporadic basal cell carcinomas of the skin and primitive neuroectodermal tumors of the central nervous system. Cancer Res. 58, 1798–1803 (1998).
Taylor, M. D. et al. Mutations in SUFU predispose to medulloblastoma. Nature Genet. 31, 306–310 (2002).
Hahn, H. et al. Mutations of the human homolog of Drosophila patched in the nevoid basal cell carcinoma syndrome. Cell 85, 841–851 (1996).
Pomeroy, S. L. & Sturla, L. M. Molecular biology of medulloblastoma therapy. Pediatr. Neurosurg. 39, 299–304 (2003).
Schuller, U. et al. Acquisition of granule neuron precursor identity is a critical determinant of progenitor cell competence to form Shh-induced medulloblastoma. Cancer Cell 14, 123–134 (2008).
Yang, Z. J. et al. Medulloblastoma can be initiated by deletion of Patched in lineage-restricted progenitors or stem cells. Cancer Cell 14, 135–145 (2008).
Browd, S. R. et al. N-myc can substitute for insulin-like growth factor signaling in a mouse model of sonic hedgehog-induced medulloblastoma. Cancer Res. 66, 2666–2672 (2006).
Goodrich, L. V., Milenkovic, L., Higgins, K. M. & Scott, M. P. Altered neural cell fates and medulloblastoma in mouse patched mutants. Science 277, 1109–1113 (1997).
Hallahan, A. R. et al. The SmoA1 mouse model reveals that notch signaling is critical for the growth and survival of sonic hedgehog-induced medulloblastomas. Cancer Res. 64, 7794–7800 (2004).
Hatton, B. A. et al. The Smo/Smo model: hedgehog-induced medulloblastoma with 90% incidence and leptomeningeal spread. Cancer Res. 68, 1768–1776 (2008).
Lee, Y. et al. Patched2 modulates tumorigenesis in patched1 heterozygous mice. Cancer Res. 66, 6964–6971 (2006).
McCall, T. D., Pedone, C. A. & Fults, D. W. Apoptosis suppression by somatic cell transfer of Bcl-2 promotes Sonic hedgehog-dependent medulloblastoma formation in mice. Cancer Res. 67, 5179–5185 (2007).
Pazzaglia, S. et al. High incidence of medulloblastoma following X-ray-irradiation of newborn Ptc1 heterozygous mice. Oncogene 21, 7580–7584 (2002).
Pazzaglia, S. et al. Linking DNA damage to medulloblastoma tumorigenesis in patched heterozygous knockout mice. Oncogene 25, 1165–1173 (2006).
Rao, G. et al. Sonic hedgehog and insulin-like growth factor signaling synergize to induce medulloblastoma formation from nestin-expressing neural progenitors in mice. Oncogene 23, 6156–6162 (2004).
Rao, R. D. et al. Disruption of parallel and converging signaling pathways contributes to the synergistic antitumor effects of simultaneous mTOR and EGFR inhibition in GBM cells. Neoplasia 7, 921–929 (2005).
Wetmore, C., Eberhart, D. E. & Curran, T. Loss of p53 but not ARF accelerates medulloblastoma in mice heterozygous for patched. Cancer Res. 61, 513–516 (2001).
Zurawel, R. H., Allen, C., Wechsler-Reya, R., Scott, M. P. & Raffel, C. Evidence that haploinsufficiency of Ptch leads to medulloblastoma in mice. Genes Chromosom. Cancer 28, 77–81 (2000).
Eberhart, C. G., Tihan, T. & Burger, P. C. Nuclear localization and mutation of beta-catenin in medulloblastomas. J. Neuropathol. Exp. Neurol. 59, 333–337 (2000).
Rossi, A., Caracciolo, V., Russo, G., Reiss, K. & Giordano, A. Medulloblastoma: from molecular pathology to therapy. Clin. Cancer Res. 14, 971–976 (2008).
Baeza, N., Masuoka, J., Kleihues, P. & Ohgaki, H. AXIN1 mutations but not deletions in cerebellar medulloblastomas. Oncogene 22, 632–636 (2003).
Huang, H. et al. APC mutations in sporadic medulloblastomas. Am. J. Pathol. 156, 433–437 (2000).
Koch, A. et al. Mutations of the Wnt antagonist AXIN2 (Conductin) result in TCF-dependent transcription in medulloblastomas. Int. J. Cancer 121, 284–291 (2007).
Koch, A. et al. Somatic mutations of WNT/wingless signaling pathway components in primitive neuroectodermal tumors. Int. J. Cancer 93, 445–449 (2001).
Zurawel, R. H., Chiappa, S. A., Allen, C. & Raffel, C. Sporadic medulloblastomas contain oncogenic β-catenin mutations. Cancer Res. 58, 896–899 (1998).
Ellison, D. W. et al. β-catenin status predicts a favorable outcome in childhood medulloblastoma: the United Kingdom Children's Cancer Study Group Brain Tumour Committee. J. Clin. Oncol. 23, 7951–7957 (2005).
Momota, H., Shih, A. H., Edgar, M. A. & Holland, E. C. c-Myc and β-catenin cooperate with loss of p53 to generate multiple members of the primitive neuroectodermal tumor family in mice. Oncogene 27, 4392–4401 (2008).
Biegel, J. A. et al. Prognostic significance of chromosome 17p deletions in childhood primitive neuroectodermal tumors (medulloblastomas) of the central nervous system. Clin. Cancer Res. 3, 473–478 (1997).
Bigner, S. H., Mark, J., Friedman, H. S., Biegel, J. A. & Bigner, D. D. Structural chromosomal abnormalities in human medulloblastoma. Cancer Genet. Cytogenet. 30, 91–101 (1988).
Cogen, P. H. & McDonald, J. D. Tumor suppressor genes and medulloblastoma. J. Neurooncol. 29, 103–112 (1996).
Lamont, J. M., McManamy, C. S., Pearson, A. D., Clifford, S. C. & Ellison, D. W. Combined histopathological and molecular cytogenetic stratification of medulloblastoma patients. Clin. Cancer Res. 10, 5482–5493 (2004).
Nicholson, J., Wickramasinghe, C., Ross, F., Crolla, J. & Ellison, D. Imbalances of chromosome 17 in medulloblastomas determined by comparative genomic hybridisation and fluorescence in situ hybridisation. Mol. Pathol. 53, 313–319 (2000).
Steichen-Gersdorf, E., Baumgartner, M., Kreczy, A., Maier, H. & Fink, F. M. Deletion mapping on chromosome 17p in medulloblastoma. Br. J. Cancer 76, 1284–1287 (1997).
Vagner-Capodano, A. M. et al. Detection of i(17q) chromosome by fluorescent in situ hybridization (FISH) with interphase nuclei in medulloblastoma. Cancer Genet. Cytogenet. 78, 1–6 (1994).
Tomlinson, F. H. et al. Aggressive medulloblastoma with high-level N-myc amplification. Mayo Clin. Proc. 69, 359–365 (1994).
Lee, Y. et al. Loss of suppressor-of-fused function promotes tumorigenesis. Oncogene 26, 6442–6447 (2007).
Uziel, T. et al. The tumor suppressors Ink4c and p53 collaborate independently with Patched to suppress medulloblastoma formation. Genes Dev. 19, 2656–2667 (2005).
Aldosari, N. et al. MYCC and MYCN oncogene amplification in medulloblastoma. A fluorescence in situ hybridization study on paraffin sections from the Children's Oncology Group. Arch. Pathol. Lab. Med. 126, 540–544 (2002).
Mendell, J. T. miRiad roles for the miR-17-92 cluster in development and disease. Cell 133, 217–222 (2008).
Rao, G., Pedone, C. A., Coffin, C. M., Holland, E. C. & Fults, D. W. c-Myc enhances sonic hedgehog-induced medulloblastoma formation from nestin-expressing neural progenitors in mice. Neoplasia 5, 198–204 (2003).
MacDonald, T. J. et al. Expression profiling of medulloblastoma: PDGFRA and the RAS/MAPK pathway as therapeutic targets for metastatic disease. Nature Genet. 29, 143–152 (2001).
Gilbertson, R. et al. Clinical and molecular stratification of disease risk in medulloblastoma. Br. J. Cancer 85, 705–712 (2001).
Calabrese, C., Frank, A., Maclean, K. & Gilbertson, R. Medulloblastoma sensitivity to 17-allylamino-17-demethoxygeldanamycin requires MEK/ERKM. J. Biol. Chem. 278, 24951–24959 (2003).
Hernan, R. et al. ERBB2 up-regulates S100A4 and several other prometastatic genes in medulloblastoma. Cancer Res. 63, 140–148 (2003).
Gilbertson, R. et al. Novel ERBB4 juxtamembrane splice variants are frequently expressed in childhood medulloblastoma. Genes Chromosom. Cancer 31, 288–294 (2001).
Gajjar, A. et al. Clinical, histopathologic, and molecular markers of prognosis: toward a new disease risk stratification system for medulloblastoma. J. Clin. Oncol. 22, 984–993 (2004).
Gilbertson, R. J., Pearson, A. D., Perry, R. H., Jaros, E. & Kelly, P. J. Prognostic significance of the c-erbB-2 oncogene product in childhood medulloblastoma. Br. J. Cancer 71, 473–477 (1995).
Gilbertson, R. J., Perry, R. H., Kelly, P. J., Pearson, A. D. & Lunec, J. Prognostic significance of HER2 and HER4 coexpression in childhood medulloblastoma. Cancer Res. 57, 3272–3280 (1997).
Pomeroy, S. L. et al. Prediction of central nervous system embryonal tumour outcome based on gene expression. Nature 415, 436–442 (2002).
Kool, M. et al. Integrated genomics identifies five medulloblastoma subtypes with distinct genetic profiles, pathway signatures and clinicopathological features. PLoS ONE 3, e3088 (2008).
Thompson, M. C. et al. Genomics identifies medulloblastoma subgroups that are enriched for specific genetic alterations. J. Clin. Oncol. 24, 1924–1931 (2006). References 145 and 146 establish transcriptional subclasses of medulloblastoma and links each to specific genomic and/or transcriptional characteristics.
Reya, T., Morrison, S. J., Clarke, M. F. & Weissman, I. L. Stem cells, cancer, and cancer stem cells. Nature 414, 105–111 (2001).
Galli, R. et al. Isolation and characterization of tumorigenic, stem-like neural precursors from human glioblastoma. Cancer Res. 64, 7011–7021 (2004).
Singh, S. K. et al. Identification of a cancer stem cell in human brain tumors. Cancer Res. 63, 5821–5828 (2003).
Singh, S. K. et al. Identification of human brain tumour initiating cells. Nature 432, 396–401 (2004).
Bao, S. et al. Glioma stem cells promote radioresistance by preferential activation of the DNA damage response. Nature 444, 756–760 (2006).
Blazek, E. R., Foutch, J. L. & Maki, G. Daoy medulloblastoma cells that express CD133 are radioresistant relative to CD133- cells, and the CD133+ sector is enlarged by hypoxia. Int. J. Radiat. Oncol. Biol. Phys. 67, 1–5 (2007).
Ogden, A. T. et al. Identification of A2B5+CD133- tumor-initiating cells in adult human gliomas. Neurosurgery 62, 505–515 (2008).
Wang, J. et al. CD133 negative glioma cells form tumors in nude rats and give rise to CD133 positive cells. Int. J. Cancer 122, 761–768 (2008).
Cheng, J. X., Liu, B. L. & Zhang, X. How powerful is CD133 as a cancer stem cell marker in brain tumors? Cancer Treat Rev. 35, 403–408 (2009).
Gilbertson, R. J. & Rich, J. N. Making a tumour's bed: glioblastoma stem cells and the vascular niche. Nature Rev. Cancer 7, 733–736 (2007).
Altman, J. & Das, G. D. Autoradiographic and histological evidence of postnatal hippocampal neurogenesis in rats. J. Comp. Neurol. 124, 319–335 (1965).
Lois, C. & Alvarez-Buylla, A. Proliferating subventricular zone cells in the adult mammalian forebrain can differentiate into neurons an glia. Proc. Natl Acad. Sci. USA 90, 2074–2077 (1993).
Riquelme, P. A., Drapeau, E. & Doetsch, F. Brain micro-ecologies: neural stem cell niches in the adult mammalian brain. Philos. Trans. R. Soc. Lond. B Biol. Sci. 363, 123–137 (2008).
Calabrese, C. et al. A perivascular niche for brain tumor stem cells. Cancer Cell 11, 69–82 (2007). This study demonstrates that brain tumour cells with stem cell-like characteristics localize in a perivascular distribution and require microvasculature for their tumour-initiating capacity.
Bao, S. et al. Stem cell-like glioma cells promote tumor angiogenesis through vascular endothelial growth factor. Cancer Res. 66, 7843–7848 (2006).
Li, Z., Wang, H., Eyler, C. E., Hjelmeland, A. B. & Rich, J. N. Turning cancer stem cells inside out: an exploration of glioma stem cell signaling pathways. J. Biol. Chem. 284, 16705–16709 (2009).
Becher, O. J. et al. Gli activity correlates with tumor grade in platelet-derived growth factor-induced gliomas. Cancer Res. 68, 2241–2249 (2008).
Li, Z. et al. Hypoxia-inducible factors regulate tumorigenic capacity of glioma stem cells. Cancer Cell 15, 501–513 (2009).
Covello, K. L. et al. HIF-2α regulates Oct-4: effects of hypoxia on stem cell function, embryonic development, and tumor growth. Genes Dev. 20, 557–570 (2006).
Heddleston, J. M., Li, Z., McLendon, R. E., Hjelmeland, A. B. & Rich, J. N. The hypoxic microenvironment maintains glioblastoma stem cells and promotes reprogramming towards a cancer stem cell phenotype. Cell Cycle 8, 3274–3284 (2009).
Wang, Z., Li, Y., Banerjee, S. & Sarkar, F. H. Emerging role of Notch in stem cells and cancer. Cancer Lett. 279, 8–12 (2009).
Shih, A. H. & Holland, E. C. Notch signaling enhances nestin expression in gliomas. Neoplasia 8, 1072–1082 (2006).
Bhattacharya, S., Das, A., Mallya, K. & Ahmad, I. Maintenance of retinal stem cells by Abcg2 is regulated by notch signaling. J. Cell Sci. 120, 2652–2662 (2007).
Fan, X. et al. Notch pathway inhibition depletes stem-like cells and blocks engraftment in embryonal brain tumors. Cancer Res. 66, 7445–7452 (2006).
Bleau, A. M. et al. PTEN/PI3K/Akt pathway regulates the side population phenotype and ABCG2 activity in glioma tumor stem-like cells. Cell Stem Cell 4, 226–235 (2009). This study demonstrates that PI3K–AKT–mTOR signalling regulates the expression of stem-like characteristics in glioma cells.
Charles, N. et al. Nitric oxide drives notch signaling and stem cell-like character via PKG in the perivascular niche of PDGF gliomas. Cell Stem Cell. 6, 141–152 (2010).
Hambardzumyan, D. et al. PI3K pathway regulates survival of cancer stem cells residing in the perivascular niche following radiation in medulloblastoma in vivo. Genes Dev. 22, 436–448 (2008).
Hambardzumyan, D., Amankulor, N. M., Helmy, K. Y., Becher, O. J. & Holland, E. C. Modeling Adult Gliomas Using RCAS/t-va Technology. Transl. Oncol. 2, 89–95 (2009).
Lindberg, N., Kastemar, M., Olofsson, T., Smits, A. & Uhrbom, L. Oligodendrocyte progenitor cells can act as cell of origin for experimental glioma. Oncogene 28, 2266–2275 (2009).
Assanah, M. et al. Glial progenitors in adult white matter are driven to form malignant gliomas by platelet-derived growth factor-expressing retroviruses. J. Neurosci. 26, 6781–6790 (2006).
Bleau, A. M., Huse, J. T. & Holland, E. C. The ABCG2 resistance network of glioblastoma. Cell Cycle 8, 2936–2944 (2009).
Wick, W., Platten, M. & Weller, M. New (alternative) temozolomide regimens for the treatment of glioma. Neuro Oncol. 11, 69–79 (2009).
Mellinghoff, I. K. et al. Molecular determinants of the response of glioblastomas to EGFR kinase inhibitors. N. Engl. J. Med. 353, 2012–2024 (2005). This trial of EGFR kinase inhibitors suggests that patients with both the EGFR vIII mutation and intact PTEN may be a molecularly defined subgroup that is more likely to respond to targeted kinase inhibition.
Yauch, R. L. et al. Smoothened mutation confers resistance to a Hedgehog pathway inhibitor in medulloblastoma. Science 326, 572–574 (2009).
Huse, J. T. & Holland, E. C. Genetically engineered mouse models of brain cancer and the promise of preclinical testing. Brain Pathol. 19, 132–143 (2009).
Charest, A. et al. ROS fusion tyrosine kinase activates a SH2 domain-containing phospatase-2/phosphatidylinositol 3-kinase/mammalian target of rapamycin signaling axis to form glioblastoma in mice. Cancer Res. 66, 7473–7481 (2006).
Lassman, A. B. et al. Molecular study of malignant gliomas treated with epidermal growth factor receptor inhibitors: tissue analysis from North American Brain Tumor Consortium Trials 01–03 and 00–01. Clin. Cancer Res. 11, 7841–7850 (2005).
Liebner, S. et al. Claudin-1 and claudin-5 expression and tight junction morphology are altered in blood vessels of human glioblastoma multiforme. Acta Neuropathol. 100, 323–331 (2000).
de Vries, N. A., Beijnen, J. H., Boogerd, W. & van Tellingen, O. Blood-brain barrier and chemotherapeutic treatment of brain tumors. Expert Rev. Neurother 6, 1199–1209 (2006).
Laquintana, V. et al. New strategies to deliver anticancer drugs to brain tumors. Expert Opin. Drug Deliv. 6, 1017–1032 (2009).
Acknowledgements
The authors would like to thank M. Rosenblum for providing representative tumour micrographs.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
Glossary
- Malignant glioma
-
Diffuse glioma of astrocytic, oligodendroglial or mixed lineage with a World Health Organization grade of either III or IV.
- Glial
-
Pertaining to glia, the non-neuronal support cells in the nervous system.
- Neuronal
-
Pertaining to neurons, the primary functional unit of the nervous system.
- Histogenesis
-
The origin of a tissue or tumour especially with regard to its development and formation.
- Variant III deletion
-
Pathogenic deletion mutant of EGFR involving exons 2–7 that leads to a constitutively active truncated protein.
- Neurofibromatosis type 1
-
Hereditary cancer-predisposing syndrome caused by mutations in NF1 and characterized most commonly by neurofibromas, optic gliomas and malignant peripheral nerve sheath tumours.
- Turcot's syndrome
-
Hereditary cancer-predisposing syndrome caused by mutations in APC and most commonly characterized by adenomatous polyposis of the colon and an increased incidence of neuroepithelial tumours.
- Supratentorial PNET
-
A class of PNET arising in the forebrain that is distinct from medulloblastoma.
- Atypical teratoid/rhabdoid tumour
-
An aggressive brain tumour variant that occurs in young children and is characterized by loss of the transcription factor integrase interactor 1 (INI1).
- Perivascular niche
-
A specialized microenvironment intimately associated with the microvasculature where the plurality of brain tumour stem-like cells seem to reside.
Rights and permissions
About this article
Cite this article
Huse, J., Holland, E. Targeting brain cancer: advances in the molecular pathology of malignant glioma and medulloblastoma. Nat Rev Cancer 10, 319–331 (2010). https://doi.org/10.1038/nrc2818
Issue Date:
DOI: https://doi.org/10.1038/nrc2818