Abstract
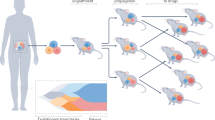
Patient-derived xenografts (PDXs) have emerged as an important platform to elucidate new treatments and biomarkers in oncology. PDX models are used to address clinically relevant questions, including the contribution of tumour heterogeneity to therapeutic responsiveness, the patterns of cancer evolutionary dynamics during tumour progression and under drug pressure, and the mechanisms of resistance to treatment. The ability of PDX models to predict clinical outcomes is being improved through mouse humanization strategies and the implementation of co-clinical trials, within which patients and PDXs reciprocally inform therapeutic decisions. This Opinion article discusses aspects of PDX modelling that are relevant to these questions and highlights the merits of shared PDX resources to advance cancer medicine from the perspective of EurOPDX, an international initiative devoted to PDX-based research.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Change history
15 September 2017
In the online html version of this article, Joan Seoane's affiliations were not correct. He is also a member of the EurOPDX Consortium and is at the Vall d'Hebron Institute of Oncology, 08035 Barcelona, the Universitat Autònoma de Barcelona, 08193 Bellaterra, and the Institució Catalana de Recerca i Estudis Avançats (ICREA), 08010 Barcelona, Spain. This is correct in the print and PDF versions of the article.
References
de Bono, J. S. & Ashworth, A. Translating cancer research into targeted therapeutics. Nature 467, 543–549 (2010).
Daniel, V. C. et al. A primary xenograft model of small-cell lung cancer reveals irreversible changes in gene expression imposed by culture in vitro. Cancer Res. 69, 3364–3373 (2009).
Arrowsmith, J. Trial watch: phase II failures: 2008–2010. Nat. Rev. Drug Discov. 10, 328–329 (2011).
Arrowsmith, J. & Miller, P. Trial watch: phase II and phase III attrition rates 2011–2012. Nat. Rev. Drug Discov. 12, 569 (2013).
Paul, S. M. et al. How to improve R&D productivity: the pharmaceutical industry's grand challenge. Nat. Rev. Drug Discov. 9, 203–214 (2010).
Bertotti, A. et al. A molecularly annotated platform of patient-derived xenografts (“xenopatients”) identifies HER2 as an effective therapeutic target in cetuximab-resistant colorectal cancer. Cancer Discov. 1, 508–523 (2011).
Bertotti, A. et al. The genomic landscape of response to EGFR blockade in colorectal cancer. Nature 526, 263–267 (2015).
DeRose, Y. S. et al. Tumor grafts derived from women with breast cancer authentically reflect tumor pathology, growth, metastasis and disease outcomes. Nat. Med. 17, 1514–1520 (2011).
Gao, H. et al. High-throughput screening using patient-derived tumor xenografts to predict clinical trial drug response. Nat. Med. 21, 1318–1325 (2015).
Hidalgo, M. et al. Patient-derived xenograft models: an emerging platform for translational cancer research. Cancer Discov. 4, 998–1013 (2014).
Siolas, D. & Hannon, G. J. Patient-derived tumor xenografts: transforming clinical samples into mouse models. Cancer Res. 73, 5315–5319 (2013).
Tentler, J. J. et al. Patient-derived tumour xenografts as models for oncology drug development. Nat. Rev. Clin. Oncol. 9, 338–350 (2012).
Day, C. P., Merlino, G. & Van Dyke, T. Preclinical mouse cancer models: a maze of opportunities and challenges. Cell 163, 39–53 (2015).
Tabassum, D. P. & Polyak, K. Tumorigenesis: it takes a village. Nat. Rev. Cancer 15, 473–483 (2015).
Aparicio, S. & Caldas, C. The implications of clonal genome evolution for cancer medicine. N. Engl. J. Med. 368, 842–851 (2013).
Almendro, V. et al. Inference of tumor evolution during chemotherapy by computational modeling and in situ analysis of genetic and phenotypic cellular diversity. Cell Rep. 6, 514–527 (2014).
Kreso, A. & Dick, J. E. Evolution of the cancer stem cell model. Cell Stem Cell 14, 275–291 (2014).
Kreso, A. et al. Variable clonal repopulation dynamics influence chemotherapy response in colorectal cancer. Science 339, 543–548 (2013).
Marusyk, A., Almendro, V. & Polyak, K. Intra-tumour heterogeneity: a looking glass for cancer? Nat. Rev. Cancer 12, 323–334 (2012).
Maley, C. C. et al. Genetic clonal diversity predicts progression to esophageal adenocarcinoma. Nat. Genet. 38, 468–473 (2006).
Curtis, C. et al. The genomic and transcriptomic architecture of 2,000 breast tumours reveals novel subgroups. Nature 486, 346–352 (2012).
Dawson, S. J., Rueda, O. M., Aparicio, S. & Caldas, C. A new genome-driven integrated classification of breast cancer and its implications. EMBO J. 32, 617–628 (2013).
Shah, S. P. et al. The clonal and mutational evolution spectrum of primary triple-negative breast cancers. Nature 486, 395–399 (2012).
Eirew, P. et al. Dynamics of genomic clones in breast cancer patient xenografts at single-cell resolution. Nature 518, 422–426 (2015).
Nik-Zainal, S. et al. The life history of 21 breast cancers. Cell 149, 994–1007 (2012).
Bhang, H. E. et al. Studying clonal dynamics in response to cancer therapy using high-complexity barcoding. Nat. Med. 21, 440–448 (2015).
Jeselsohn, R. et al. Emergence of constitutively active estrogen receptor-α mutations in pretreated advanced estrogen receptor-positive breast cancer. Clin. Cancer Res. 20, 1757–1767 (2014).
Murtaza, M. et al. Multifocal clonal evolution characterized using circulating tumour DNA in a case of metastatic breast cancer. Nat. Commun. 6, 8760 (2015).
Bruna, A. et al. A biobank of breast cancer explants with preserved intra-tumor heterogeneity to screen anticancer compounds. Cell 167, 1–15 (2016).
Marangoni, E. et al. A new model of patient tumor-derived breast cancer xenografts for preclinical assays. Clin. Cancer Res. 13, 3989–3998 (2007).
Li, S. et al. Endocrine-therapy-resistant ESR1 variants revealed by genomic characterization of breast-cancer-derived xenografts. Cell Rep. 4, 1116–1130 (2013).
Cassidy, J. W., Caldas, C. & Bruna, A. Maintaining tumor heterogeneity in patient-derived tumor xenografts. Cancer Res. 75, 2963–2968 (2015).
Cottu, P. et al. Acquired resistance to endocrine treatments is associated with tumor-specific molecular changes in patient-derived luminal breast cancer xenografts. Clin. Cancer Res. 20, 4314–4325 (2014).
Ter Brugge, P. et al. Mechanisms of therapy resistance in patient-derived xenograft models of BRCA1-deficient breast cancer. J. Natl Cancer Inst. 108, djw148 (2016).
Kemper, K. et al. Intra- and inter-tumor heterogeneity in a vemurafenib-resistant melanoma patient and derived xenografts. EMBO Mol. Med. 7, 1104–1118 (2015).
Shi, H. et al. Acquired resistance and clonal evolution in melanoma during BRAF inhibitor therapy. Cancer Discov. 4, 80–93 (2014).
Tirosh, I. et al. Dissecting the multicellular ecosystem of metastatic melanoma by single-cell RNA-seq. Science 352, 189–196 (2016).
Kemper, K. et al. BRAFV600E kinase domain duplication identified in therapy-refractory melanoma patient-derived xenografts. Cell Rep. 16, 263–277 (2016).
Nguyen, L. V. et al. DNA barcoding reveals diverse growth kinetics of human breast tumour subclones in serially passaged xenografts. Nat. Commun. 5, 5871 (2014).
Joosse, S. A., Gorges, T. M. & Pantel, K. Biology, detection, and clinical implications of circulating tumor cells. EMBO Mol. Med. 7, 1–11 (2015).
Massague, J. & Obenauf, A. C. Metastatic colonization by circulating tumour cells. Nature 529, 298–306 (2016).
Lapidot, T. et al. A cell initiating human acute myeloid leukaemia after transplantation into SCID mice. Nature 367, 645–648 (1994).
Pece, S. et al. Biological and molecular heterogeneity of breast cancers correlates with their cancer stem cell content. Cell 140, 62–73 (2010).
Reya, T., Morrison, S. J., Clarke, M. F. & Weissman, I. L. Stem cells, cancer, and cancer stem cells. Nature 414, 105–111 (2001).
Li, C., Lee, C. J. & Simeone, D. M. Identification of human pancreatic cancer stem cells. Methods Mol. Biol. 568, 161–173 (2009).
Lawson, D. A. et al. Single-cell analysis reveals a stem-cell program in human metastatic breast cancer cells. Nature 526, 131–135 (2015).
Li, X. et al. Intrinsic resistance of tumorigenic breast cancer cells to chemotherapy. J. Natl Cancer Inst. 100, 672–679 (2008).
Todaro, M. et al. Colon cancer stem cells dictate tumor growth and resist cell death by production of interleukin-4. Cell Stem Cell 1, 389–402 (2007).
Al-Hajj, M., Wicha, M. S., Benito-Hernandez, A., Morrison, S. J. & Clarke, M. F. Prospective identification of tumorigenic breast cancer cells. Proc. Natl Acad. Sci. USA 100, 3983–3988 (2003).
Fan, F. et al. The requirement for freshly isolated human colorectal cancer (CRC) cells in isolating CRC stem cells. Br. J. Cancer 112, 539–546 (2015).
Borovski, T., De Sousa, E. M. F., Vermeulen, L. & Medema, J. P. Cancer stem cell niche: the place to be. Cancer Res. 71, 634–639 (2011).
Charafe-Jauffret, E. et al. ALDH1-positive cancer stem cells predict engraftment of primary breast tumors and are governed by a common stem cell program. Cancer Res. 73, 7290–7300 (2013).
Miranda-Lorenzo, I. et al. Intracellular autofluorescence: a biomarker for epithelial cancer stem cells. Nat. Methods 11, 1161–1169 (2014).
Sainz, B. Jr et al. Microenvironmental hCAP-18/LL-37 promotes pancreatic ductal adenocarcinoma by activating its cancer stem cell compartment. Gut 64, 1921–1935 (2015).
Magee, J. A., Piskounova, E. & Morrison, S. J. Cancer stem cells: impact, heterogeneity, and uncertainty. Cancer Cell 21, 283–296 (2012).
Rottenberg, S. et al. Selective induction of chemotherapy resistance of mammary tumors in a conditional mouse model for hereditary breast cancer. Proc. Natl Acad. Sci. USA 104, 12117–12122 (2007).
Castillo-Avila, W. et al. Sunitinib inhibits tumor growth and synergizes with cisplatin in orthotopic models of cisplatin-sensitive and cisplatin-resistant human testicular germ cell tumors. Clin. Cancer Res. 15, 3384–3395 (2009).
Juliachs, M. et al. The PDGFRβ–AKT pathway contributes to CDDP-acquired resistance in testicular germ cell tumors. Clin. Cancer Res. 20, 658–667 (2014).
Simoes, B. M. et al. Anti-estrogen resistance in human breast tumors is driven by JAG1-NOTCH4-dependent cancer stem cell activity. Cell Rep. 12, 1968–1977 (2015).
Herrera-Abreu, M. T. et al. Early adaptation and acquired resistance to CDK4/6 inhibition in estrogen receptor-positive breast cancer. Cancer Res. 76, 2301–2313 (2016).
Kim, K. T. et al. Single-cell mRNA sequencing identifies subclonal heterogeneity in anti-cancer drug responses of lung adenocarcinoma cells. Genome Biol. 16, 127 (2015).
Cottu, P. et al. Modeling of response to endocrine therapy in a panel of human luminal breast cancer xenografts. Breast Cancer Res. Treat. 133, 595–606 (2012).
Zhang, X. et al. A renewable tissue resource of phenotypically stable, biologically and ethnically diverse, patient-derived human breast cancer xenograft models. Cancer Res. 73, 4885–4897 (2013).
Das Thakur, M. et al. Modelling vemurafenib resistance in melanoma reveals a strategy to forestall drug resistance. Nature 494, 251–255 (2013).
Sun, C. et al. Reversible and adaptive resistance to BRAFV600E inhibition in melanoma. Nature 508, 118–122 (2014).
Stewart, E. L. et al. Clinical utility of patient-derived xenografts to determine biomarkers of prognosis and map resistance pathways in EGFR-mutant lung adenocarcinoma. J. Clin. Oncol. 33, 2472–2480 (2015).
Stebbing, J. et al. Patient-derived xenografts for individualized care in advanced sarcoma. Cancer 120, 2006–2015 (2014).
Balko, J. M. et al. Molecular profiling of the residual disease of triple-negative breast cancers after neoadjuvant chemotherapy identifies actionable therapeutic targets. Cancer Discov. 4, 232–245 (2014).
Zacarias-Fluck, M. F. et al. Effect of cellular senescence on the growth of HER2-positive breast cancers. J. Natl Cancer Inst. 107, djv020 (2015).
Bankert, R. B., Egilmez, N. K. & Hess, S. D. Human-SCID mouse chimeric models for the evaluation of anti-cancer therapies. Trends Immunol. 22, 386–393 (2001).
Hylander, B. L. et al. Origin of the vasculature supporting growth of primary patient tumor xenografts. J. Transl Med. 11, 110 (2013).
Schreiber, R. D., Old, L. J. & Smyth, M. J. Cancer immunoediting: integrating immunity's roles in cancer suppression and promotion. Science 331, 1565–1570 (2011).
Guichelaar, T. et al. Human regulatory T cells do not suppress the antitumor immunity in the bone marrow: a role for bone marrow stromal cells in neutralizing regulatory T cells. Clin. Cancer Res. 19, 1467–1475 (2013).
King, M. A. et al. Human peripheral blood leucocyte non-obese diabetic-severe combined immunodeficiency interleukin-2 receptor gamma chain gene mouse model of xenogeneic graft-versus-host-like disease and the role of host major histocompatibility complex. Clin. Exp. Immunol. 157, 104–118 (2009).
Holzapfel, B. M., Wagner, F., Thibaudeau, L., Levesque, J. P. & Hutmacher, D. W. Concise review: humanized models of tumor immunology in the 21st century: convergence of cancer research and tissue engineering. Stem Cells 33, 1696–1704 (2015).
Drake, A. C., Chen, Q. & Chen, J. Engineering humanized mice for improved hematopoietic reconstitution. Cell. Mol. Immunol. 9, 215–224 (2012).
Reinisch, A., Gratzinger, D., Hong, W.-J. & Majeti, R. A. Novel humanized bone marrow niche xenotransplantation model allows superior engraftment of human normal and malignant hematopoietic cells and reveals myelofibrosis-initiating cells in the HSC compartment. Blood 124, 349 (2014).
Rongvaux, A. et al. Human hemato-lymphoid system mice: current use and future potential for medicine. Annu. Rev. Immunol. 31, 635–674 (2013).
Voloshin, T. et al. G-CSF supplementation with chemotherapy can promote revascularization and subsequent tumor regrowth: prevention by a CXCR4 antagonist. Blood 118, 3426–3435 (2011).
Morton, J. J. et al. XactMice: humanizing mouse bone marrow enables microenvironment reconstitution in a patient-derived xenograft model of head and neck cancer. Oncogene 35, 290–300 (2016).
Takenaka, K. et al. Polymorphism in Sirpa modulates engraftment of human hematopoietic stem cells. Nat. Immunol. 8, 1313–1323 (2007).
Du, Q. et al. Establishment of and comparison between orthotopic xenograft and subcutaneous xenograft models of gallbladder carcinoma. Asian Pac. J. Cancer Prev. 15, 3747–3752 (2014).
Hoffman, R. M. Patient-derived orthotopic xenografts: better mimic of metastasis than subcutaneous xenografts. Nat. Rev. Cancer 15, 451–452 (2015).
Dai, L., Lu, C., Yu, X. I., Dai, L. J. & Zhou, J. X. Construction of orthotopic xenograft mouse models for human pancreatic cancer. Exp. Ther. Med. 10, 1033–1038 (2015).
Ambrogio, C. et al. Combined inhibition of DDR1 and Notch signaling is a therapeutic strategy for KRAS-driven lung adenocarcinoma. Nat. Med. 22, 270–277 (2016).
de Jong, M., Essers, J. & van Weerden, W. M. Imaging preclinical tumour models: improving translational power. Nat. Rev. Cancer 14, 481–493 (2014).
Iorns, E. et al. A new mouse model for the study of human breast cancer metastasis. PLoS ONE 7, e47995 (2012).
Gupta, P., Adkins, C., Lockman, P. & Srivastava, S. K. Metastasis of breast tumor cells to brain is suppressed by phenethyl isothiocyanate in a novel metastasis model. PLoS ONE 8, e67278 (2013).
Lee, H. W. et al. Patient-derived xenografts from non-small cell lung cancer brain metastases are valuable translational platforms for the development of personalized targeted therapy. Clin. Cancer Res. 21, 1172–1182 (2015).
Chen, H. J. et al. Comprehensive models of human primary and metastatic colorectal tumors in immunodeficient and immunocompetent mice by chemokine targeting. Nat. Biotechnol. 33, 656–660 (2015).
Girotti, M. R. et al. Application of sequencing, liquid biopsies, and patient-derived xenografts for personalized medicine in melanoma. Cancer Discov. 6, 286–299 (2016).
Nunes, M. et al. Evaluating patient-derived colorectal cancer xenografts as preclinical models by comparison with patient clinical data. Cancer Res. 75, 1560–1566 (2015).
Sartore-Bianchi, A. et al. Dual-targeted therapy with trastuzumab and lapatinib in treatment-refractory, KRAS codon 12/13 wild-type, HER2-positive metastatic colorectal cancer (HERACLES): a proof-of-concept, multicentre, open-label, phase 2 trial. Lancet Oncol. 17, 738–746 (2016).
Krebs, M. G. et al. Evaluation and prognostic significance of circulating tumor cells in patients with non-small-cell lung cancer. J. Clin. Oncol. 29, 1556–1563 (2011).
Scher, H. I. et al. Circulating tumour cells as prognostic markers in progressive, castration-resistant prostate cancer: a reanalysis of IMMC38 trial data. Lancet Oncol. 10, 233–239 (2009).
Zhang, L. et al. Meta-analysis of the prognostic value of circulating tumor cells in breast cancer. Clin. Cancer Res. 18, 5701–5710 (2012).
Baccelli, I. et al. Identification of a population of blood circulating tumor cells from breast cancer patients that initiates metastasis in a xenograft assay. Nat. Biotechnol. 31, 539–544 (2013).
Hodgkinson, C. L. et al. Tumorigenicity and genetic profiling of circulating tumor cells in small-cell lung cancer. Nat. Med. 20, 897–903 (2014).
Yap, T. A., Lorente, D., Omlin, A., Olmos, D. & de Bono, J. S. Circulating tumor cells: a multifunctional biomarker. Clin. Cancer Res. 20, 2553–2568 (2014).
Alix-Panabieres, C. & Pantel, K. Challenges in circulating tumour cell research. Nat. Rev. Cancer 14, 623–631 (2014).
Ignatiadis, M., Lee, M. & Jeffrey, S. S. Circulating tumor cells and circulating tumor DNA: challenges and opportunities on the path to clinical utility. Clin. Cancer Res. 21, 4786–4800 (2015).
Williams, E. S. et al. Generation of prostate cancer patient derived xenograft models from circulating tumor cells. J. Vis. Exp. 104, e53182 (2015).
Toyoshima, K. et al. Analysis of circulating tumor cells derived from advanced gastric cancer. Int. J. Cancer 137, 991–998 (2015).
Yu, M. et al. Cancer therapy. Ex vivo culture of circulating breast tumor cells for individualized testing of drug susceptibility. Science 345, 216–220 (2014).
Cayrefourcq, L. et al. Establishment and characterization of a cell line from human circulating colon cancer cells. Cancer Res. 75, 892–901 (2015).
Aggarwal, C. et al. Relationship among circulating tumor cells, CEA and overall survival in patients with metastatic colorectal cancer. Ann. Oncol. 24, 420–428 (2013).
Vishnoi, M. et al. The isolation and characterization of CTC subsets related to breast cancer dormancy. Sci. Rep. 5, 17533 (2015).
Krebs, M. G. et al. Molecular analysis of circulating tumour cells-biology and biomarkers. Nat. Rev. Clin. Oncol. 11, 129–144 (2014).
Markou, A. et al. PIK3CA mutational status in circulating tumor cells can change during disease recurrence or progression in patients with breast cancer. Clin. Cancer Res. 20, 5823–5834 (2014).
Giuliano, M. et al. Circulating and disseminated tumor cells from breast cancer patient-derived xenograft-bearing mice as a novel model to study metastasis. Breast Cancer Res. 17, 3 (2015).
Torphy, R. J. et al. Circulating tumor cells as a biomarker of response to treatment in patient-derived xenograft mouse models of pancreatic adenocarcinoma. PLoS ONE 9, e89474 (2014).
Jordan, N. V. et al. HER2 expression identifies dynamic functional states within circulating breast cancer cells. Nature 537, 102–106 (2016).
Garraway, L. A. & Lander, E. S. Lessons from the cancer genome. Cell 153, 17–37 (2013).
Trusolino, L. & Bertotti, A. Compensatory pathways in oncogenic kinase signaling and resistance to targeted therapies: six degrees of separation. Cancer Discov. 2, 876–880 (2012).
Bardelli, A. et al. Amplification of the MET receptor drives resistance to anti-EGFR therapies in colorectal cancer. Cancer Discov. 3, 658–673 (2013).
Kavuri, S. M. et al. HER2 activating mutations are targets for colorectal cancer treatment. Cancer Discov. 5, 832–841 (2015).
Leto, S. M. et al. Sustained inhibition of HER3 and EGFR is necessary to induce regression of HER2-amplified gastrointestinal carcinomas. Clin. Cancer Res. 21, 5519–5531 (2015).
Zanella, E. R. et al. IGF2 is an actionable target that identifies a distinct subpopulation of colorectal cancer patients with marginal response to anti-EGFR therapies. Sci. Transl Med. 7, 272ra12 (2015).
Sun, C. et al. Intrinsic resistance to MEK inhibition in KRAS mutant lung and colon cancer through transcriptional induction of ERBB3. Cell Rep. 7, 86–93 (2014).
Weickhardt, A. J. et al. Dual targeting of the epidermal growth factor receptor using the combination of cetuximab and erlotinib: preclinical evaluation and results of the phase II DUX study in chemotherapy-refractory, advanced colorectal cancer. J. Clin. Oncol. 30, 1505–1512 (2012).
Long, G. V. et al. Combined BRAF and MEK inhibition versus BRAF inhibition alone in melanoma. N. Engl. J. Med. 371, 1877–1888 (2014).
Bossi, D. et al. In vivo genetic screens of patient-derived tumors revealed unexpected frailty of the transformed phenotype. Cancer Discov. 6, 650–663 (2016).
Carugo, A. et al. In vivo functional platform targeting patient-derived xenografts identifies WDR5-Myc association as a critical determinant of pancreatic cancer. Cell Rep. 16, 133–147 (2016).
Murphy, B. et al. Evaluation of alternative in vivo drug screening methodology: a single mouse analysis. Cancer Res. 76, 5798–5809 (2016).
Migliardi, G. et al. Inhibition of MEK and PI3K/mTOR suppresses tumor growth but does not cause tumor regression in patient-derived xenografts of RAS-mutant colorectal carcinomas. Clin. Cancer Res. 18, 2515–2525 (2012).
Boj, S. F. et al. Organoid models of human and mouse ductal pancreatic cancer. Cell 160, 324–338 (2015).
Gao, D. et al. Organoid cultures derived from patients with advanced prostate cancer. Cell 159, 176–187 (2014).
Huang, L. et al. Ductal pancreatic cancer modeling and drug screening using human pluripotent stem cell- and patient-derived tumor organoids. Nat. Med. 21, 1364–1371 (2015).
Sato, T. et al. Long-term expansion of epithelial organoids from human colon, adenoma, adenocarcinoma, and Barrett's epithelium. Gastroenterology 141, 1762–1772 (2011).
van de Wetering, M. et al. Prospective derivation of a living organoid biobank of colorectal cancer patients. Cell 161, 933–945 (2015).
Weeber, F. et al. Preserved genetic diversity in organoids cultured from biopsies of human colorectal cancer metastases. Proc. Natl Acad. Sci. USA 112, 13308–13311 (2015).
Hubert, C. G. et al. A three-dimensional organoid culture system derived from human glioblastomas recapitulates the hypoxic gradients and cancer stem cell heterogeneity of tumors found in vivo. Cancer Res. 76, 2465–2477 (2016).
Crystal, A. S. et al. Patient-derived models of acquired resistance can identify effective drug combinations for cancer. Science 346, 1480–1486 (2014).
Nardella, C., Lunardi, A., Patnaik, A., Cantley, L. C. & Pandolfi, P. P. The APL paradigm and the “co-clinical trial” project. Cancer Discov. 1, 108–116 (2011).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT02720796 (2016).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT02752932 (2016).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT02312245 (2016).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT02795650 (2016).
Azaro, A. et al. A first-in-human phase I trial of LY2780301, a dual p70 S6 kinase and Akt Inhibitor, in patients with advanced or metastatic cancer. Invest. New Drugs 33, 710–719 (2015).
Juric, D. et al. Convergent loss of PTEN leads to clinical resistance to a PI(3)Kα inhibitor. Nature 518, 240–244 (2015).
Morelli, M. P. et al. Prioritizing phase I treatment options through preclinical testing on personalized tumorgraft. J. Clin. Oncol. 30, e45–e48 (2012).
Zembutsu, H. et al. Genome-wide cDNA microarray screening to correlate gene expression profiles with sensitivity of 85 human cancer xenografts to anticancer drugs. Cancer Res. 62, 518–527 (2002).
Gupta, S. K. et al. Delineation of MGMT Hypermethylation as a biomarker for veliparib-mediated temozolomide-sensitizing therapy of glioblastoma. J. Natl Cancer Inst. 108, djv369 (2016).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT02152982 (2016).
Brown, K. E. et al. Proteomic profiling of patient-derived glioblastoma xenografts identifies a subset with activated EGFR: implications for drug development. J. Neurochem. 133, 730–738 (2015).
Grinde, M. T. et al. Interplay of choline metabolites and genes in patient-derived breast cancer xenografts. Breast Cancer Res. 16, R5 (2014).
Moestue, S. A. et al. Distinct choline metabolic profiles are associated with differences in gene expression for basal-like and luminal-like breast cancer xenograft models. BMC Cancer 10, 433 (2010).
Glunde, K., Jie, C. & Bhujwalla, Z. M. Molecular causes of the aberrant choline phospholipid metabolism in breast cancer. Cancer Res. 64, 4270–4276 (2004).
Nelson, S. J. et al. Metabolic imaging of patients with prostate cancer using hyperpolarized [1-13C]pyruvate. Sci. Transl. Med. 5, 198ra108 (2013).
Klomp, D. W. et al. 31P MRSI and 1H MRS at 7 T: initial results in human breast cancer. NMR Biomed. 24, 1337–1342 (2011).
Esmaeili, M. et al. In vivo31P magnetic resonance spectroscopic imaging (MRSI) for metabolic profiling of human breast cancer xenografts. J. Magn. Reson. Imaging 41, 601–609 (2015).
Eyre, R. et al. Patient-derived mammosphere and xenograft tumour initiation correlates with progression to metastasis. J. Mammary Gland Biol. Neoplasia http://dx.doi.org/10.1007/s10911-016-9361-8 (2016).
Moon, H. G. et al. Prognostic and functional importance of the engraftment-associated genes in the patient-derived xenograft models of triple-negative breast cancers. Breast Cancer Res. Treat. 154, 13–22 (2015).
Garrido-Laguna, I. et al. Tumor engraftment in nude mice and enrichment in stroma- related gene pathways predict poor survival and resistance to gemcitabine in patients with pancreatic cancer. Clin. Cancer Res. 17, 5793–5800 (2011).
Delitto, D. et al. Patient-derived xenograft models for pancreatic adenocarcinoma demonstrate retention of tumor morphology through incorporation of murine stromal elements. Am. J. Pathol. 185, 1297–1303 (2015).
Isella, C. et al. Stromal contribution to the colorectal cancer transcriptome. Nat. Genet. 47, 312–319 (2015).
Calon, A. et al. Stromal gene expression defines poor-prognosis subtypes in colorectal cancer. Nat. Genet. 47, 320–329 (2015).
Dunne, P. D. et al. Challenging the cancer molecular stratification dogma: intratumoral heterogeneity undermines consensus molecular subtypes and potential diagnostic value in colorectal cancer. Clin. Cancer Res. 22, 4095–4104 (2016).
Bhargava, M. et al. Scatter factor and hepatocyte growth factor: activities, properties, and mechanism. Cell Growth Differ. 3, 11–20 (1992).
Pennacchietti, S. et al. Microenvironment-derived HGF overcomes genetically determined sensitivity to anti-MET drugs. Cancer Res. 74, 6598–6609 (2014).
Mestas, J. & Hughes, C. C. Of mice and not men: differences between mouse and human immunology. J. Immunol. 172, 2731–2738 (2004).
Brodeur, J. et al. Knock-in of human HGF into the mouse genome maintains endogenous HGF regulation and supports the growth of HGF-dependent human cancer cell lines. Cancer Res. 69, abstr. 305 (2009).
Zimmer, L. et al. Phase I expansion and pharmacodynamic study of the oral MEK inhibitor RO4987655 (CH4987655) in selected patients with advanced cancer with RAS–RAF mutations. Clin. Cancer Res. 20, 4251–4261 (2014).
Eckhardt, S. G. et al. Challenges, opportunities, and lessons learned in the bench-to-bedside translation of xenopatient studies. Clin. Cancer Res. 22 (16 Suppl.), abstr. IA20 (2016).
Townsend, E. C. et al. The public repository of xenografts enables discovery and randomized phase II-like trials in mice. Cancer Cell 29, 574–586 (2016).
Baralis, E., Bertotti, A., Fiori, A. & Grand, A. LAS: a software platform to support oncological data management. J. Med. Syst. 36 (Suppl. 1), S81–S90 (2012).
Chou, J. et al. Phenotypic and transcriptional fidelity of patient-derived colon cancer xenografts in immune-deficient mice. PLoS ONE 8, e79874 (2013).
Ito, R., Takahashi, T., Katano, I. & Ito, M. Current advances in humanized mouse models. Cell. Mol. Immunol. 9, 208–214 (2012).
Conway, T. et al. Xenome — a tool for classifying reads from xenograft samples. Bioinformatics 28, i172–i178 (2012).
Bacac, M. et al. A novel carcinoembryonic antigen T-cell bispecific antibody (CEA TCB) for the treatment of solid tumors. Clin. Cancer Res. 22, 3286–3297 (2016).
Ito, M. et al. NOD/SCID/γcnull mouse: an excellent recipient mouse model for engraftment of human cells. Blood 100, 3175–3182 (2002).
Shultz, L. D. et al. Human lymphoid and myeloid cell development in NOD/LtSz-scid IL2Rγnull mice engrafted with mobilized human hemopoietic stem cells. J. Immunol. 174, 6477–6489 (2005).
Shultz, L. D., Brehm, M. A., Garcia-Martinez, J. V. & Greiner, D. L. Humanized mice for immune system investigation: progress, promise and challenges. Nat. Rev. Immunol. 12, 786–798 (2012).
Traggiai, E. et al. Development of a human adaptive immune system in cord blood cell-transplanted mice. Science 304, 104–107 (2004).
Ito, R. et al. Establishment of a human allergy model using human IL-3/GM-CSF-transgenic NOG mice. J. Immunol. 191, 2890–2899 (2013).
Billerbeck, E. et al. Development of human CD4+FoxP3+ regulatory T cells in human stem cell factor-, granulocyte-macrophage colony-stimulating factor-, and interleukin-3-expressing NOD-SCID IL2Rγnull humanized mice. Blood 117, 3076–3086 (2011).
Rongvaux, A. et al. Development and function of human innate immune cells in a humanized mouse model. Nat. Biotechnol. 32, 364–372 (2014).
Cunningham, D. et al. Cetuximab monotherapy and cetuximab plus irinotecan in irinotecan-refractory metastatic colorectal cancer. N. Engl. J. Med. 351, 337–345 (2004).
Kawazoe, A. et al. A retrospective observational study of clinicopathological features of KRAS, NRAS, BRAF and PIK3CA mutations in Japanese patients with metastatic colorectal cancer. BMC Cancer 15, 258 (2015).
Acknowledgements
The authors would like to thank all members of the EurOPDX Consortium who also contributed to this article, and in particular S. Corso, S. Giordano, P. P. López-Casas, K. Moran-Jones and F. Nemati. The Caldas laboratory would like to thank the PGE team for their support, especially Lisa, Steve and Yi. A.T.B. is supported by Science Foundation Ireland under grants 13/CDA/2183 and 15/TIDA/2963 and further receives funding from the Irish Cancer Society Collaborative Cancer Research Centre BREAST-PREDICT Grant CCRC13GAL. D.G.A. and R.B.C. are supported by Breast Cancer Now. F.A., E.H. and J.C.M. received KULeuven GOA funding (GOA/14/012) and a research grant from Stichting tegen Kanker. J.A. is funded by the Breast Cancer Research Foundation, the Spanish Association Against Cancer (AECC) and the Instituto de Salud Carlos III (PI16/00253 and CIBER-ONC). A.V.B. and D.K.C. are supported by Cancer Research UK (C29717/A17263), the Wellcome Trust (10372/Z/14/Z), the Scottish Genomes Partnership — SEHHD-CSO 1175759/2158447, the Howat Foundation and Pancreatic Cancer UK. A.B., C.C. and O.M.R. have been supported by funding from Cancer Research UK and by the European Union to the EUROCAN Network of Excellence (FP7; grant number 260791). E.B. is supported by the CETOCOEN PLUS project (CZ.02.1.01/0.0/0.0/15_003/0000469) and the RECETOX Research Infrastructure (LM2015051). G.C. and D.V. were funded by NIH transformative R01CA156695 and European Research Council (ERC) Advanced grant 1400206AdG-322875. S.G.E. receives support from NCI grant 1UM1CA186688 for early-phase trials through the ET-CTN. E.G.S. is supported by the Spanish Ministry of Economy and Competitivity MINECO and from the ISCIII (SAF2014-55997; PIE13/00022, co-funded by FEDER funds/ European Regional Development Fund (ERDF) — a way to build Europe), by a Career Catalyst Grant from the Susan Komen Foundation (CCR13262449) and by a European Research Council Consolidator grant (CoG682935). M.A.J. is supported by an Irish Health Research Board Health Research Award (#HRA-POR-2014-547). S.D.J. is supported by the Dutch Cancer Society (grants RUG 2010-4833, RUG 2011-5231, RUG 2012-5477 and RUG 2014-6691). J.J. is funded by the Dutch Cancer Society (NKI 2011-5197 and EMCR 2014-7048), the Netherlands Organisation for Scientific Research (Zenith 93512009, Vici 91814643, CancerGenomiCs.nl) and the European Research Council (ERC-SyG CombatCancer). K.K. and D.S.P. are supported by the Dutch Cancer Society (NKI-2013-5799). L.L. and P.G.P. are funded by ERC Advanced Grant 341131 and Italian Association for Cancer Research (AIRC) Investigator Investigator Grant 14216. G.M.M. receives funds from the Norwegian Cancer Society (421851) and the Research Council of Norway (222262/F20). J.H.N. is funded by the Research Council of Norway under grant 250459/F20. H.G.P. is supported by the Instituto de Salud Carlos III and the Miguel Servet Program (MSII14/00037). V.S. is supported by the Instituto de Salud Carlos III (PI13/01714 and the Miguel Servet Program CP14/00028), by a Career Catalyst Grant from the Susan Komen Foundation CCR15330331 and the FERO Foundation. L.S. was funded by Worldwide Cancer Research (WCR/AICR Grant #13-1182), the European Research Council (CoG Grant #617473), the Instituto de Salud Carlos III (FIS Grant #PI13/01705) and the FERO Foundation. A.V. is supported by the Instituto de Salud Carlos III (PI13/0133 and PIE13/00022 (Oncoprofile)), Fundación Mutua Madrileña AP150932014 and a grant from the Spanish Association Against Cancer from Barcelona, AECC. A.B. is supported by AIRC (Investigator Grant project 15571). L.T. and E.M. are supported by the AIRC (Special Programme Molecular Clinical Oncology 5 × 1000, project 9970, and Investigator Grant projects, 14205 to L.T. and 12944 to E.M.) and also receive funding from the Fondazione Piemontese per la Ricerca sul Cancro-ONLUS (5 × 1000 Italian Ministry of Health 2011).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
Grants from Celgene and Boehringer-Ingelheim, honoraria from Roche and Genentech, consultancy for Roche, Genentech, Novartis and Sanofi-Aventis (G.C.), consultancy for Oncodesign and funding by Novartis (S.R.R.), founder of the spin-off Xenopat S.L. (A.V.). The other authors declare no competing interests.
Related links
Supplementary information
Supplementary information S1 (table)
Examples of improvements in humanised mouse models for PDX studies (PDF 155 kb)
Rights and permissions
About this article
Cite this article
Byrne, A., Alférez, D., Amant, F. et al. Interrogating open issues in cancer precision medicine with patient-derived xenografts. Nat Rev Cancer 17, 254–268 (2017). https://doi.org/10.1038/nrc.2016.140
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrc.2016.140
This article is cited by
-
Magnetic resonance imaging-guided intracranial resection of glioblastoma tumors in patient-derived orthotopic xenografts leads to clinically relevant tumor recurrence
BMC Cancer (2024)
-
Simultaneous inhibition of FAK and ROS1 synergistically repressed triple-negative breast cancer by upregulating p53 signalling
Biomarker Research (2024)
-
Patient-derived organoids in human cancer: a platform for fundamental research and precision medicine
Molecular Biomedicine (2024)
-
Microfluidic high-throughput 3D cell culture
Nature Reviews Bioengineering (2024)
-
Omics-based molecular classifications empowering in precision oncology
Cellular Oncology (2024)