Abstract
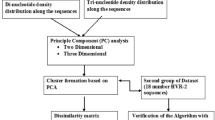
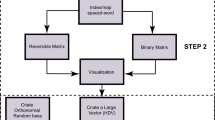
This paper presents a novel method to segment/decode DNA sequences based on n-gram statistical language model. Firstly, we find the length of most DNA “words” is 12 to 15 bps by analyzing the genomes of 12 model species. The bound of language entropy of DNA sequence is about 1.5674 bits. After building an n-gram biology languages model, we design an unsupervised ‘probability approach to word segmentation’ method to segment the DNA sequences. The benchmark of segmenting method is also proposed. In cross segmenting test, we find different genomes may use the similar language, but belong to different branches, just like the English and French/Latin. We present some possible applications of this method at last.
Similar content being viewed by others
Article PDF
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Liang, W. Segmenting DNA sequence into words based on statistical language model. Nat Prec (2012). https://doi.org/10.1038/npre.2012.6939.1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/npre.2012.6939.1