Abstract
In a recent report of the North American Prodrome Longitudinal Study (NAPLS), clinical high-risk individuals who converted to psychosis showed a steeper rate of cortical gray matter reduction compared with non-converters and healthy controls, and the rate of cortical thinning was correlated with levels of proinflammatory cytokines at baseline. These findings suggest a critical role for microglia, the resident macrophages in the brain, in perturbations of cortical maturation processes associated with onset of psychosis. Elucidating gene expression pathways promoting microglial action prior to disease onset would inform potential preventative intervention targets. Here we used a forward stepwise regression algorithm to build a classifier of baseline microRNA expression in peripheral leukocytes associated with annualized rate of cortical thinning in a subsample of the NAPLS cohort (N=74). Our cortical thinning classifier included nine microRNAs, p=3.63 × 10-08, R2=0.358, permutation-based p=0.039, the gene targets of which were enriched for intracellular signaling pathways that are important to coordinating inflammatory responses within immune cells (p<0.05, Benjamini–Hochberg corrected). The classifier was also related to proinflammatory cytokine levels in serum (p=0.038). Furthermore, miRNAs that predicted conversion status were found to do so in a manner partially mediated by rate of cortical thinning (point estimate=0.078 (95% CIs: 0.003, 0.168), p=0.03). Many of the miRNAs identified here have been previously implicated in brain development, synaptic plasticity, immune function and/or schizophrenia, showing some convergence across studies and methodologies. Altered intracellular signaling within the immune system may interact with cortical maturation in individuals at high risk for schizophrenia promoting disease onset.
Similar content being viewed by others
INTRODUCTION
Schizophrenia is a highly heritable, genetically complex neurodevelopmental disorder that typically onsets in late adolescence (Cannon et al, 1998). Recent work suggests that cortical maturation during this period is perturbed in individuals at high risk for psychosis who transition to a psychotic disorder (Borgwardt et al, 2008; Cannon et al, 2015). The cortex typically thins during this developmental stage—ostensibly the aggregate result of synaptic pruning (Boksa, 2012)—and individuals who eventually develop psychosis show an exaggerated and accelerated trajectory of cortical thinning from prodromal phase to disease onset, compared with similarly high-risk individuals who ultimately do not develop psychosis and normal risk healthy controls (Cannon et al, 2015). This divergence in neurodevelopment does not appear to be a secondary effect of medication (Cannon et al, 2015) and thus may be a contributing aspect of the pathophysiology driving the conversion from high vulnerability to a disease state.
Mechanisms promoting this neurodevelopmental trajectory might be key targets for preventative intervention. Proximate changes in neuronal development are almost certainly central to the emergence of pathology (Feinberg, 1982; Glausier and Lewis, 2013; Monzón-Sandoval et al, 2015; Sekar et al, 2016). In addition, mounting evidence has tied increased action of microglia, the resident macrophages in the brain, with schizophrenia (van Berckel et al, 2008; Doorduin et al, 2009; Garay and McAllister, 2010; Khandaker et al, 2015; Monji et al, 2013)—a connection that reflects the crucial role of microglia in normal cortical synaptic pruning during late adolescence: coordinating the selection and removal of synapses. Indeed, the impact of schizophrenia-associated genetic variation on synaptic pruning during an analogous critical period in mice has suggested that dysregulation of gene expression within both microglia and neurons, leading to over-activation of microglia and excessive pruning, may be one key mechanism (Sekar et al, 2016). This can also occur in response to systemic inflammation, which erodes the blood-brain barrier and allows T cells to enter, signaling microglial activation and increasing neuroinflammation (Boulanger and Shatz, 2004; Garay and McAllister, 2010; Khandaker et al, 2015; Monji et al, 2013). In line with this, in Phase 2 the North American Prodrome Longitudinal Study (NAPLS2), peripheral proinflammatory cytokine levels—specifically those known to activate microglia resulting in synaptic pruning—were correlated with rate of cortical thinning across high-risk individuals and controls, an effect particularly pronounced in those individuals who developed psychosis (Cannon et al, 2015). Given this evidence, it is reasonable to hypothesize that cortical maturation in schizophrenia may be influenced by increased microglia activity resulting from genetic variation within immune cells, systemic inflammation, or both.
However, preliminary work linking immunological genetic abnormalities with cortical development in psychosis has been exclusively done in animal models (Lee et al, 2014; Sekar et al, 2016). An important next step in determining the mechanisms by which synaptic pruning is perturbed in schizophrenia is to identify gene expression networks altered prior to onset of illness that promote this neurodevelopmental trajectory in humans. Ruling out gene expression changes secondary to the illness or medication is critical to establish a plausible causal association, which is possible when using a prospective, longitudinal research design, as was done when these sources were ruled out as accounting for the steeper rate of cortical thinning during the ramp-up to full psychosis in clinical high risk (CHR) cases in the NAPLS2 cohort (Cannon et al, 2015).
Investigations of both rare mutations and common polymorphisms suggest genetic variation associated with schizophrenia disproportionally affects miRNAs (Hauberg et al, 2016; Warnica et al, 2015; Zhang et al, 2015). In addition, expression of multiple miRNAs differ by patient status in circulating leukocytes (Gardiner et al, 2012; Lai et al, 2011; Wei et al, 2015), cerebrospinal fluid (Gallego et al, 2012), and postmortem brain tissue (Kim et al, 2010; Miller et al, 2012; Moreau et al, 2011); peripheral expression also correlates with neurocognitive and physiological biomarkers of schizophrenia (Lai et al, 2011). These small, non-coding RNAs (~22 nucleotides) modulate gene expression of up to 60% of protein-coding genes by inhibiting translation of target messenger RNA (Friedman et al, 2009) and serve as regulatory hubs of gene expression networks (Lim et al, 2005; Ziats and Rennert, 2014). The functional implications of these disruptions in schizophrenia predominantly suggest abnormalities in gene pathways critical for neurodevelopment and synaptic plasticity (Caputo et al, 2015; Mellios and Sur, 2012). miRNA expression is thus a good candidate by which liability for schizophrenia could manifest to alter neurodevelopment prior to disease onset.
In the current study, we assessed how baseline patterns of miRNA expression in circulating leukocytes were correlated with annualized cortical gray matter reduction in a subsample of the NAPLS2 cohort, including both CHR individuals, some of whom converted to psychosis, as well as healthy controls. To interpret the functional relevance of any miRNAs associated with cortical thinning, we both established the corresponding miRNA gene targets and assessed enrichment of any canonical signaling pathways, as well as tested the relationship between associated miRNAs and plasma levels of proinflammatory cytokines, a marker of inflammatory response previously associated with cortical thinning (Cannon et al, 2015).
MATERIALS AND METHODS
Participants
Participants in NAPLS2 were evaluated at eight sites using the Structured Interview for Prodromal Syndromes (McGlashan et al, 2010) and the Structured Clinical Interview for Axis I (DSM-IV; First et al, 1995) at each assessment by trained interviewers who met high reliability standards (ICCs=0.92–0.96; Addington et al, 2012). Assessments were done every six months for up to two years. Subject recruitment criteria are listed in the Supplementary Materials and Methods.
In total, 764 CHR subjects and 279 demographically matched subjects ages 12–35 were included. Subjects were scanned using MRI at baseline and either at 12-month follow-up or at the point of conversion to psychosis (N=405). Those with longitudinal imaging data were included in our primary study reporting on differential rates of change in cortical thickness (Cannon et al, 2015). As part of a pilot study, a subset of subjects additionally submitted samples for RNA sequencing (N=101). Thirteen CHR cases who converted to psychosis, 34 CHR cases who did not convert within 2 years, and 27 healthy comparison subjects had usable and complete data after exclusions due to either missing (ie, only baseline MRI scan) or poor quality data (criteria described below). There were no significant differences in age, sex, education or family income by group (Table 1). Additional clinical characteristics are listed in Supplementary Table S1.
Longitudinal Reduction in Cortical Thickness
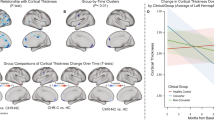
We previously showed that CHR subjects who converted to psychosis showed a steeper annualized rate of reduction in cortical thickness in the right superior frontal, middle frontal, and medial orbitofrontal regions relative to those who did not convert, accounting for baseline cortical thickness, gender, site, and multiple comparisons using a false-discovery rate correction (Cannon et al, 2015). Controls and non-converters did not differ in these regions. Annualized rate of thickness reduction observed in the right superior frontal cortex—the most pronounced difference between converters and non-converters found previously—was extracted for all individuals and used as the outcome of interest; rates for the other two areas were also extracted for supplementary analyses (Supplementary Information). Scanning parameters, image processing, quality control, and statistical analyses are detailed in the original publication (Cannon et al, 2015).
RNA Sequencing
miRNA expression in this cohort was assayed using RNA sequencing. Our group recently developed a miRNA classifier for conversion status among CHR subjects in this cohort (Jeffries et al, 2016). That publication included detailed descriptions of the assays and normalization procedures, as well as the exact data matrix of z-scored expression of the 136 miRNAs used in the current study; a summary is in Supplementary Materials and Methods. From an initial set of 101 samples, we excluded six (three controls, three CHR) with low abundance reads, leaving 27 controls and 68 CHR subjects (N=95). Of these, 74 had at least two MRI scans and thus were included in the current study.
Cytokines
Blood samples for a subset of subjects were obtained and processed using the Luminx Human DiscoveryMAP bead-based multiplex immunoassay (Luminex, Austin, TX). More detailed methods regarding processing and normalization have been published elsewhere (Cannon et al, 2015). For the current study, we calculated the same sum score of three proinflammatory cytokines (ie, cytokines known to consistently activate microglia) previously associated with cortical thinning among prodromal individuals: tumor necrosis factor-α, interleukin-2, and interferon-γ (Cannon et al, 2015).
Statistical Analyses
Classification algorithm
We detected one or more reads for 1569 of the 2588 canonical miRNAs in miRBase v21; however, we limited our analyses to miRNAs that were robustly expressed (>10 000 or more total reads), resulting in 136 miRNAs. We employed a forward stepwise regression algorithm (Liu et al, 2005; Perkins et al, 2015), the Coarse Approximation Linear Function (CALF; https://cran.r-project.org/web/packages/CALF/index.html), to develop our classifier, the rationale and detailed description of which is published elsewhere (Jeffries et al, 2016). Briefly, this algorithm selected first the best (miRNA) predictor of rate of cortical thickness reduction, based on a Pearson correlation, then added miRNAs to the model until no improvement in the metric was possible. The nonzero weights applied to selected miRNAs could be either +1 or −1; for ease of interpretation, the first choice is made such that the classifier positively correlates with the target. The restriction to coarse weights should improve generalizability (at a marginal cost to performance), as sample-specific β-weights are less likely to be stable across samples, especially when derived from small samples (Hand, 2006; Jeffries et al, 2016; Perkins et al, 2015).
Permutations
To assess the statistical significance of this classifier, we ran the CALF algorithm 2000 times on permutations of the cortical thinning phenotype, limiting the number of miRNAs that could be included to the number selected in the true data. Further, we applied the CALF algorithm to the true data in 2000 randomly selected 80% subsets of the subjects, again limiting to the number of miRNAs selected in the full data set. miRNAs selected more frequently should be more likely to generalize to independent samples, as their selection is less dependent on any particular combination of subjects. miRNAs selected in the full data set that were not selected frequently in random subsets (ie, in the top 15 miRNAs) were excluded from the final classifier. To confirm the effect of the classifier on cortical thinning in the presence of covariates, a weighted score was calculated by summing the values of the included miRNAs and was entered as a predictor into a linear regression of rate of superior frontal cortical thinning, along with age, sex, and diagnostic group (converters vs non-converters and controls). To assess any differential effects of the classifier by diagnostic group—a key test of the clinical relevance of such a classifier—we next tested the interaction effect between classifier and diagnostic group on cortical thinning, including age and sex in the regression as well.
Association with cytokines
To assess the relationship between the miRNA cortical thinning classifier and the proinflammatory cytokine score, we ran a robust linear regression including the miRNA classifier, age, sex, and diagnostic group as predictors using the MASS package in R for the regression and Wald test command for multiple coefficients (f.robftest) from sfsmisc to extract parameters for each predictor. We opted to use a robust linear regression due to the detection of cytokine score outliers within the sample (>5 SD of the mean).
Mediation
A classifier was built previously in this sample to predict conversion to psychosis in the CHR subjects (N=68) using miRNA expression (Jeffries et al, 2016). To test if the relationship between the conversion-based miRNA classifier and conversion status was mediated by annualized reduction in cortical thickness, we performed a mediation analysis in CHR subjects with longitudinal imaging data (N=47) using the mediation package in R (Tingley et al, 2014), implementing the Preacher & Hayes (Preacher and Hayes, 2008) method of bootstrapping (1000 simulations) to determine confidence intervals.
All analyses were implemented in the freely available R statistical environment (Version 3.2.2; http://cran.r-project.org/). R code developed for statistical modeling is available upon request (Zheutlin).
Functional Annotation of miRNA Gene Targets
Experimentally validated and high-confidence predicted gene targets of the nine miRNAs included in the classifier for cortical gray matter reduction were established using Ingenuity Pathway Analysis (IPA; Ingenuity Systems, USA, www.ingenuity.com). This gene list was assessed for significant enrichment of genes within canonical molecular pathways (p<0.05, Benjamini–Hochberg corrected for multiple comparisons). IPA also generates a list of top diseases and biological functions based on the output.
By default, enrichment analyses test whether an input gene list has relatively more genes from predefined sets of genes than would be expected by chance drawing a similarly sized list from the entire human genome (~20 000 genes). Since only a subset of genes is ever expressed in each tissue type, this genome-wide reference set can include thousands of genes that could never be identified by the experiment (ie, genes never expressed in that tissue) and thus, such an analysis inflates p-values (by using an unfairly large denominator) and is biased towards finding significant ‘enrichment’ of tissue-specific genes (eg, immune-related genes assayed from immune cells). To protect against this, we included only genes known to be expressed in immune cells (8828 IDs mapped to 8766 genes) in the reference gene set. This allows us to identify pathways affected by the miRNAs measured here in the tissue from which they were assayed. For reference, we also included the same analysis using the standard whole-genome reference set.
RESULTS
miRNA Classifier of Cortical Thinning
The CALF algorithm developed for rate of cortical thinning in the full data set included 10 miRNAs (Pearson r=0.695; Table 2). Testing this algorithm using permuted phenotypic data yielded 77 of 2000 iterations with an equivalent or higher Pearson correlation than the true data (r⩾0.695), resulting in an empirical p-value of 0.039 (Figure 1a). Across analyses of random 80% subsets of the true data, nine of the 10 miRNAs were among the top most frequently selected markers (Figure 1b and Supplementary Figure S1), and thus these nine were summed together (weights were always consistent across subsets) to create the final classifier function (Figure 2; Table 2). In a follow-up analysis with age, sex, and diagnostic outcomes as covariates, this classifier was confirmed to predict the rate of superior frontal cortical thinning, t=6.20, p=3.63 × 10-08, R2=0.358. Critically, the interaction between risk group and miRNA classifier was also significant, t=2.11, p=0.039, R2=0.061, such that the predictive association of the miRNA classifier and rate of cortical thinning was stronger among converters relative to non-converters and controls (Figure 2). Finally, the classifier was also related to serum levels of proinflammatory cytokines, F=4.49, p=0.038.
miRNA classifier development and validation. To test the internal robustness of the classifier derived using true data, permutations and analyses using only subsets of the data were employed. (a) 2000 permutations were run and the distribution of correlations between the permuted phenotype and corresponding miRNA classifier was plotted here. The dashed line represents the correlation in the true data (r=0.695). 77 permutations outperformed the true data by chance, yielding an empirical p-value of .039. (b) 2000 analyses using random 80% subsets of the true data were run. The 20 most frequently selected miRNAs identified across analyses were displayed here and colored according to whether or not they were selected in the model using all the data. It is likely that the ‘not selected’ miRNAs are frequently selected across subsets due to low signal to noise and/or that many miRNAs are not independent and thus, correlated miRNAs are all chosen more frequently, but each is chosen arbitrarily in a given subset. The nine miRNAs selected frequently in subsets as well as in the full data set were included in the final classifier.
miRNA classifier for annualized rate of cortical gray matter reduction. The classifier reflects expression across nine miRNAs identified using a forward stepwise linear algorithm. Slopes for diagnostic groups were plotted separately, as they differed significantly. Points represent subjects and were coded by shape and color to reflect diagnostic group.
Functional Annotation of miRNA Gene Targets
A total of 3,657 experimental and high-probability predicted gene targets were identified for the selected nine miRNAs, of which 2,517 were expressed in immune cells; 2,482 of these were mapped to functional groups and included for annotation. We found significant enrichment for genes within the following pathways: protein kinase A signaling, ERK5 signaling, factors promoting cardiogenesis in vertebrates, PPARα/RXRα activation, HGF signaling, and reelin signaling in neurons. The top biological functions identified were embryonic development, nervous system development, and function, and organismal development; top molecular and cellular functions included cellular development, cellular growth and proliferation, and cellular movement. Canonical pathways identified using the standard whole-genome reference set are listed in Supplementary Results.
Cortical Thinning and miRNA Predictors of Conversion to Psychosis
In a prior report, we developed a miRNA classifier of conversion status among CHR subjects in this sample. That classifier, which included five miRNAs, accounted for 41.9% of the variance in conversion to psychosis (Jeffries et al, 2016). A bootstrapped mediation analysis revealed that rate of cortical thinning partially mediated this relationship (point estimate=0.078 (95% CIs: 0.003, 0.168), p=0.03), despite the fact that the classifier predicting conversion and the classifier predicting cortical thinning included only one overlapping miRNA: hsa-miR-103a-3p. An estimated 7.8% of conversion status (18.7% of the miRNA classifier effect on conversion status) in this sample can be attributed to annualized reduction in cortical gray matter.
DISCUSSION
We have identified miRNA expression within peripheral leukocytes that correlated with the rate of superior frontal cortical thinning across CHR and control subjects and particularly in converters to psychosis. The gene targets of the identified miRNAs are enriched for intracellular signaling pathways primary to cellular development, growth, and proliferation. Many of the miRNAs identified here have been previously implicated in brain development, synaptic plasticity, immune function, and/or schizophrenia, showing some convergence across studies and methodologies. Previously, miR-339-3p was found to be differentially expressed in human adolescent prefrontal cortex (Ziats and Rennert, 2014), miR-501-3p was involved in postnatal brain development in mice via affecting NMDA-mediated synaptic plasticity (Hu et al, 2015), and miR-142-5p was one of the most highly expressed miRNAs in T cells (Sonkoly et al, 2008; Wu et al, 2007). Two more of the identified miRNAs, miR-26b-5p and miR-27b-3p, have been related to schizophrenia directly; the first was a top regulator of schizophrenia-related genes (Gumerov and Hegyi, 2015) and the second was differentially expressed in schizophrenia patient-derived neural progenitor cells (Topol et al, 2016). Taken together, these findings suggest that alterations in signal transduction may be particular mechanisms within the immune system that promote disease onset during the prodromal phase via affecting cortical development.
Specifically, these findings implicate molecular pathways that are broadly important to cellular signal transduction. Every canonical pathway identified by our enrichment analyses included MAPK and related molecules, many included adhesion molecules (eg, CAMK family), and most were involved in similar protein kinase pathways, all of which are core intracellular signaling pathways. One primary role of these pathways within immune cells is coordinating inflammatory responses (Loza et al, 2007; Medzhitov and Horng, 2009; Saklatvala et al, 2003; Sohn et al, 2005). Furthermore, we found a significant correlation between plasma levels of proinflammatory cytokines—known both to activate microglia and correlate with cortical thinning in prodromal subjects (Cannon et al, 2015)—and our miRNA classifier, indicating a potential link between miRNA-mediated inflammatory responses and cortical maturation. This gene expression dysregulation could be present in peripheral tissue exclusively, or both peripheral and central immune cells (Ponomarev et al, 2013), but in either case could promote an increased inflammatory response, which could ultimately result in over-pruning (Khandaker et al, 2015; Monji et al, 2013; Sekar et al, 2016). Prior PET imaging studies have established that recent-onset patients show neuroinflammation across gray matter and have posited that this could contribute to synaptic pruning (van Berckel et al, 2008; Doorduin et al, 2009; Khandaker et al, 2015). We can only speculate as to which mechanisms underlie the relationship between changes in miRNA expression within the peripheral immune system and cortical maturation, but based on this evidence, we hypothesize that the connection is realized via increased microglia action, which would directly impact synaptic pruning.
Future work in animals experimentally testing the effects of gene expression alterations in immune cells on synaptic pruning could further dissociate the role of these potential mechanisms in this process. Ongoing efforts to experimentally validate miRNA–mRNA databases in humans should also be incorporated into future work, as listed miRNA gene targets are often only predicted targets (Betel et al, 2008).
Cortical thinning also mediated the relationship between a previously identified miRNA classifier of conversion status (Jeffries et al, 2016), which included five miRNAs (one of which, hsa-miR-103a-3p, was also identified in the cortical thinning analysis), and conversion to psychosis, suggesting the impact of miRNA expression on psychosis onset more broadly is partially realized via effects on cortical development. This overlap is likely only partial as miRNA expression promoting psychosis may affect many systems besides cortical development; similarly, some miRNAs affecting cortical maturation may be broadly developmentally relevant and not singularly psychosis-promoting. This may also be why there was not more overlap between the miRNA classifiers for conversion and cortical thinning—these processes are only somewhat overlapping. However, even one miRNA can have a relatively large impact on the gene expression profiles and function of cells, so this overlap is not necessarily limited in impact (Lim et al, 2005).
This work has several limitations, most significantly the small sample size and lack of independent replication, indicating the current estimates should be taken as preliminary, as they are likely inflated. Unfortunately, funding restricted the sample with miRNA available to its current size; larger cohorts in the future could better test for multivariate effects of all miRNAs on cortical thinning, and more systematically across brain regions, which may yield a more comprehensive understanding of the relationship between miRNA expression and cortical thinning. In addition, while RNA sequencing offers superior resolution to other high-throughput technologies (ie, sequences are identified in reads rather than sequence-specific hybridization probes and thus do not need to be previously known), there is some concern with reproducibility. However, in this data set, a subset of markers in a subset of subjects was tested using both RNA sequencing and high-throughput quantitative PCR and similar relative expression levels were observed (Jeffries et al, 2016).
Ideally, results such as these would be externally validated in an independent replication set. However, to our knowledge, no clinical high-risk sample with analogous genetic and longitudinal imaging data exists. To internally validate this classifier, we have applied permutation testing to approximate the chance we would have identified these miRNAs spuriously. Results from these tests indicated that our model outperformed permutations 96.1% of time, yielding an empirically significant probability that our results were not due to chance (p=0.039).
The novel component of this study relates to observed associations of microRNA with the already established set of relationships among cortical thinning, inflammatory cytokines, and conversion to psychosis. However, it is important to note that the subjects in this study represent a subset of those used in our original report identifying regions showing accelerated cortical thinning among converters and demonstrating an association between peripheral blood cytokines and cortical thinning (Cannon et al, 2015). Given this, and given that the region most sensitive to differential thinning among converters was used as the target for developing the multivariate microRNA classifier in this study, it is possible that the observed associations of microRNA with the already established set of relationships among cortical thinning, inflammatory cytokines, and conversion to psychosis are incidental (ie, as in a circular by-product of being derived with reference to the same criterion—conversion to psychosis). In such a case, high-risk individuals who convert to psychosis could show differences in inflammatory signaling and rate of thinning of the cortex and, for unrelated reasons, also show differences in miRNA expression. On the basis of the present data and study design, we cannot rule out this possibility. Nonetheless, it is difficult to imagine miRNA expression having no mechanistic contribution to inflammation and cortical thinning, particularly given that many of the miRNAs selected by this approach signal within immune- and synaptic plasticity-related pathways. Clearly, a prospective longitudinal study that measures gene expression, inflammatory signaling markers, cortical thickness, and conversion to psychosis at multiple time points is needed to help clarify the mechanistic connections between these phenomena.
In the current study, we have provided preliminary evidence for the involvement of nine miRNAs in cortical development during conversion to psychosis. These miRNAs regulate intracellular signaling pathways that are central to coordinating immune response and correlate with markers of systemic inflammation, highlighting a set of potential mechanisms by which tissue-specific cell signaling may influence cortical developmental trajectories of high-risk individuals who develop psychosis.
FUNDING AND DISCLOSURE
This work was supported by National Institute of Mental Health grants (grant numbers MH082004 to CDJ and DOP, MH081902 to TDC, MH081857 to BAC, MH081988 to EFW, MH081928 to LJS, MH082022 to KSC, MH081984 to JA, P50 MH066286 to CEB, and MH066160 to SWW) given to the North American Prodrome Longitudinal Study (NAPLS). NAPLS authors were also supported by a gift to the UCLA Foundation from the International Mental Health Research Organization (IMHRO). AMC holds equity in Spring Health (doing business as Spring Care Inc.), a behavioral health startup. He is lead inventor on a provisional patent submission by Yale University. DHM is a consultant for Boehringer Ingelheim. SWW reports that during the last 36 months he has received investigator-initiated research funding support from Pfizer and sponsor-initiated research funding support from Auspex and Teva. He has consulted to Biomedisyn (unpaid) and Boehringer Ingelheim. He has also served as an unpaid consultant to DSM-5. He has been granted US patent no. 8492418 B2 for a method of treating prodromal schizophrenia with glycine agonists, is an inventor on a patent pending for a method of predicting psychosis risk using blood biomarker analysis, and has received royalties from Oxford University Press. TDC reports that he is a consultant to the Los Angeles County Department of Mental Health and to Boehringer Ingelheim Pharmaceuticals and is a co-inventor on a pending patent for a blood-based prediction algorithm for psychosis. The remaining authors declare no conflict of interest.
References
Addington J, Cadenhead KS, Cornblatt BA, Mathalon DH, McGlashan TH, Perkins DO et al (2012). North American Prodrome Longitudinal Study (NAPLS 2): overview and recruitment. Schizophr Res 142: 77–82.
van Berckel BN, Bossong MG, Boellaard R, Kloet R, Schuitemaker A, Caspers E et al (2008). Microglia activation in recent-onset schizophrenia: a Quantitative (R)-[11C]PK11195 Positron Emission Tomography Study. Biol Psychiatry 64: 820–822.
Betel D, Wilson M, Gabow A, Marks DS, Sander C (2008). The microRNA.org resource: Targets and expression. Nucleic Acids Res 36: D149–D153.
Boksa P (2012). Abnormal synaptic pruning in schizophrenia: Urban myth or reality? J Psychiatry Neurosci 37: 75–77.
Borgwardt SJ, McGuire PK, Aston J, Gschwandtner U, Pflüger MO, Stieglitz RD et al (2008). Reductions in frontal, temporal and parietal volume associated with the onset of psychosis. Schizophr Res 106: 108–114.
Boulanger LM, Shatz CJ (2004). Immune signalling in neural development, synaptic plasticity and disease. Nat Rev Neurosci 5: 521–531.
Cannon TD, Chung Y, He G, Sun D, Jacobson A, Van Erp TGM et al (2015). Progressive reduction in cortical thickness as psychosis develops: a multisite longitudinal neuroimaging study of youth at elevated clinical risk. Biol Psychiatry 77: 147–157.
Cannon TD, Kaprio J, Lonnqvist J, Huttunen M, Koskenvuo M (1998). The genetic epidemiology of schizophrenia in a Finnish twin cohort. A population-based modeling study. Arch Gen Psychiatry 55: 67–74.
Caputo V, Ciolfi A, Macri S, Pizzuti A (2015). The emerging role of MicroRNA in schizophrenia. CNS Neurol Disord Drug Targets 14: 208–221.
Doorduin J, de Vries EFJ, Willemsen ATM, de Groot JC, Dierckx RA, Klein HC (2009). Neuroinflammation in schizophrenia-related psychosis: A PET Study. J Nucl Med 50: 1801–1807.
Feinberg I (1982). Schizophrenia: caused by a fault in programmed synaptic elimination during adolescence? J Psychiatr Res 17: 319–334.
First M, Spitzer RL, Gibbon M, Williams B, Williams JBW (1995) Structured Clinical Interview for DSM-IV Axis I Disorders, Patient Edition. Biometrics Research Department, New York State Psychiatric Institute: New York.
Friedman RC, Farh KKH, Burge CB, Bartel DP (2009). Most mammalian mRNAs are conserved targets of microRNAs. Genome Res 19: 92–105.
Gallego JA, Gordon ML, Claycomb K, Bhatt M, Lencz T, Malhotra AK (2012). In vivo MicroRNA detection and quantitation in cerebrospinal fluid. J Mol Neurosci 47: 243–248.
Garay PA, McAllister AK (2010). Novel roles for immune molecules in neural development: Implications for neurodevelopmental disorders. Front Synaptic Neurosci 2: 136.
Gardiner E, Beveridge NJ, Wu JQ, Carr V, Scott RJ, Tooney PA et al (2012). Imprinted DLK1-DIO3 region of 14q32 defines a schizophrenia-associated miRNA signature in peripheral blood mononuclear cells. Mol Psychiatry 17: 827–840.
Glausier JR, Lewis DA (2013). Dendritic spine pathology in schizophrenia. Neuroscience 251: 90–107.
Gumerov V, Hegyi H (2015). MicroRNA-derived network analysis of differentially methylated genes in schizophrenia, implicating GABA receptor B1 [GABBR1] and protein kinase B [AKT1]. Biol Direct 10: 59.
Hand DJ (2006). Classifier Technology and the Illusion of Progress. Stat Sci 21: 1–14.
Hauberg ME, Roussos P, Grove J, Børglum AD, Mattheisen M (2016). Analyzing the role of MicroRNAs in schizophrenia in the context of common genetic risk variants. JAMA Psychiatry 73: 369–377.
Hu Z, Zhao J, Hu T, Luo Y, Zhu J, Li Z (2015). miR-501-3p mediates the activity-dependent regulation of the expression of AMPA receptor subunit GluA1. J Cell Biol 208: 949–959.
Jeffries CD, Perkins DO, Chandler SD, Stark T, Yeo E, Addington J et al (2016). Insights into psychosis risk from leukocyte microRNA expression. Transl Psychiatry 6: e981.
Khandaker GM, Cousins L, Deakin J, Lennox BR, Yolken R, Jones PB (2015). Inflammation and immunity in schizophrenia: Implications for pathophysiology and treatment. Lancet Psychiatry 2: 258–270.
Kim AH, Reimers M, Maher B, Williamson V, McMichael O, McClay JL et al (2010). MicroRNA expression profiling in the prefrontal cortex of individuals affected with schizophrenia and bipolar disorders. Schizophr Res 124: 183–191.
Lai CY, Yu SL, Hsieh MH, Chen CH, Chen HY, Wen CC et al (2011). MicroRNA expression aberration as potential peripheral blood biomarkers for schizophrenia. PLoS ONE 6: e21635.
Lee H, Brott BK, Kirkby La, Adelson JD, Cheng S, Feller MB et al (2014). Synapse elimination and learning rules co-regulated by MHC class I H2-Db. Nature 509: 195–200.
Lim LP, Lau NC, Garrett-Engele P, Grimson A, Schelter JM, Castle J et al (2005). Microarray analysis shows that some microRNAs downregulate large numbers of target mRNAs. Nature 433: 769–773.
Liu X, Krishnan A, Mondry A (2005). An entropy-based gene selection method for cancer classification using microarray data. BMC Bioinformatics 6: 76.
Loza MJ, McCall CE, Li L, Isaacs WB, Xu J, Chang BL (2007). Assembly of inflammation-related genes for pathway-focused genetic analysis. PLoS ONE 2: e1035.
McGlashan TH, Walsh BC, Woods SW (2010) The Psychosis-Risk Syndrome: Handbook for Diagnosis and Follow-up. Oxford University Press: Oxford.
Medzhitov R, Horng T (2009). Transcriptional control of the inflammatory response. Nat Rev Immunol 9: 692–703.
Mellios N, Sur M (2012). The emerging role of microRNAs in schizophrenia and autism spectrum disorders. Front Psychiatry 3: 39.
Miller BH, Zeier Z, Xi L, Lanz TA., Deng S, Strathmann J et al (2012). MicroRNA-132 dysregulation in schizophrenia has implications for both neurodevelopment and adult brain function. Proc Natl Acad Sci USA 109: 3125–3130.
Monji A, Kato TA, Mizoguchi Y, Horikawa H, Seki Y, Kasai M et al (2013). Neuroinflammation in schizophrenia especially focused on the role of microglia. Prog Neuropsychopharmacol Biol Psychiatry 42: 115–121.
Monzón-Sandoval J, Castillo-Morales A, Crampton S, McKelvey L, Nolan A, O’Keeffe G et al (2015). Modular and coordinated expression of immune system regulatory and signaling components in the developing and adult nervous system. Front Cell Neurosci 9: 337.
Moreau MP, Bruse SE, David-Rus R, Buyske S, Brzustowicz LM (2011). Altered MicroRNA expression profiles in postmortem brain samples from individuals with schizophrenia and bipolar disorder. Biol Psychiatry 69: 188–193.
Perkins DO, Jeffries CD, Addington J, Bearden CE, Cadenhead KS, Cannon TD et al (2015). Towards a psychosis risk blood diagnostic for persons experiencing high-risk symptoms: preliminary results from the NAPLS project. Schizophr Bull 41: 419–428.
Ponomarev ED, Veremeyko T, Weiner HL (2013). MicroRNAs are universal regulators of differentiation, activation, and polarization of microglia and macrophages in normal and diseased CNS. Glia 61: 91–103.
Preacher KJ, Hayes AF (2008). Asymptotic and resampling strategies for assessing and comparing indirect effects in multiple mediator models. Behav Res Methods 40: 879–891.
Saklatvala J, Dean J, Clark A (2003). Control of the expression of inflammatory response genes. Biochem Soc Symp 70: 95–106.
Sekar A, Bialas AR, de Rivera H, Davis A, Hammond TR, Kamitaki N et al (2016). Schizophrenia risk from complex variation of complement component 4. Nature 530: 177–183.
Sohn SJ, Li D, Lee LK, Winoto A (2005). Transcriptional regulation of tissue-specific genes by the ERK5 mitogen-activated protein kinase. Mol Cell Biol 25: 8553–8566.
Sonkoly E, Ståhle M, Pivarcsi A (2008). MicroRNAs and immunity: novel players in the regulation of normal immune function and inflammation. Semin Cancer Biol 18: 131–140.
Tingley D, Yamamoto T, Hirose K, Keele L, Imai K (2014). mediation: R package for causal mediation analysis. J Stat Softw 59: 1–38.
Topol A, Zhu S, Hartley BJ, English J, Hauberg ME, Tran N et al (2016). Dysregulation of miRNA-9 in a subset of schizophrenia patient-derived neural progenitor cells. Cell Rep 15: 1024–1036.
Warnica W, Merico D, Costain G, Alfred SE, Wei J, Marshall CR et al (2015). Copy number variable micrornas in schizophrenia and their neurodevelopmental gene targets. Biol Psychiatry 77: 158–166.
Wei H, Yuan Y, Liu S, Wang C, Yang F, Lu Z et al (2015). Detection of circulating miRNA levels in schizophrenia. Am J Psychiatry 172: 1141–1147.
Wu H, Neilson JR, Kumar P, Manocha M, Shankar P, Sharp PA et al (2007). miRNA profiling of naive, effector and memory CD8 T cells. PLoS ONE 2: e1020.
Zhang F, Xu Y, Shugart YY, Yue W, Qi G, Yuan G et al (2015). Converging evidence implicates the abnormal microRNA system in schizophrenia. Schizophr Bull 41: 728–735.
Ziats MN, Rennert OM (2014). Identification of differentially expressed microRNAs across the developing human brain. Mol Psychiatry 19: 848–852.
Acknowledgements
We acknowledge Stephanie Lane for her contributions to the CALF algorithm and package development for R and Sarah McCoy for her technical assistance in miRNA library preparation.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on the Neuropsychopharmacology website
Supplementary information
PowerPoint slides
Rights and permissions
About this article
Cite this article
Zheutlin, A., Jeffries, C., Perkins, D. et al. The Role of microRNA Expression in Cortical Development During Conversion to Psychosis. Neuropsychopharmacol 42, 2188–2195 (2017). https://doi.org/10.1038/npp.2017.34
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/npp.2017.34
This article is cited by
-
Nicotine pretreatment alleviates MK-801-induced behavioral and cognitive deficits in mice by regulating Pdlim5/CRTC1 in the PFC
Acta Pharmacologica Sinica (2023)
-
Urinary exosomal microRNAs as predictive biomarkers for persistent psychotic-like experiences
Schizophrenia (2023)
-
A greedy regression algorithm with coarse weights offers novel advantages
Scientific Reports (2022)
-
Neurodevelopment regulators miR-137 and miR-34 family as biomarkers for early and adult onset schizophrenia
npj Schizophrenia (2021)
-
An epigenetic biomarker for adult high-functioning autism spectrum disorder
Scientific Reports (2019)