Abstract
An integrative database, Stanley Neuropathology Consortium Integrative Database (SNCID) (http://sncid.stanleyresearch.org), has been developed to facilitate psychiatric research. The SNCID includes 1749 neuropathological markers measured in 12 different brain regions in 60 human subjects (15 each schizophrenia, bipolar disorder, depression, and unaffected controls). Genome-wide expression microarray datasets from three independent studies are also included. Statistical analysis tools such as variance analysis, correlation analysis, and functional annotation tools have been integrated into the database. In this report, we first replicate an earlier correlation analysis between genome-wide expression profiles and an abnormal cytoarchitectural marker using the SNCID. We then show the potential for identifying neuropathological markers that are abnormal in subjects with psychiatric disorders. We also identify biological pathways associated with several abnormal neuropathological markers, including those in the dopamine, glutamate, Reelin, and γ-aminobutyric acid (GABA)ergic systems. Data exploration using the SNCID may provide insights into the biological pathways associated with the neurotransmitter abnormalities identified in subjects with major psychiatric disorders.
Similar content being viewed by others
INTRODUCTION
Data sharing has become more important than ever in the biomedical sciences with the advance of high-throughput technology (Ball et al, 2004) and web-based databases are one of the most efficient available resources to share datasets. Publicly available databases have been providing users with an opportunity to freely access datasets as well as find consensus regarding major questions in the field (Dennis et al, 2003; Zhang et al, 2005, 2007). In addition, a database that integrates multiple datasets from different types of experiments with statistical tools allows researchers to efficiently reanalyze datasets and often leads to novel findings (Gordon et al, 2005; Wang et al, 2003). For instance, a web-based integrative database, WebQTL, which includes genetic and phenotypic data from model animals and mapping tools (Wang et al, 2003), has been exploited in many original genetic studies (Abdeltawab et al, 2008; Bystrykh et al, 2005; Carneiro et al, 2009).
Data sharing using a web-based database is particularly essential in psychiatric studies with human postmortem tissues because of the limited sample availability. The Stanley Medical Research Institute (SMRI) has been supporting psychiatric studies by providing human postmortem tissue and has been collecting the research data derived from these samples since 1995. The Stanley Neuropathology Consortium (SNC), the first sample collection prepared and distributed, includes 15 well-matched cases in each of four groups: schizophrenia, bipolar disorder, major depression, and unaffected controls (Torrey et al, 2000). Moreover, the SNC samples were used in six independent microarray expression studies (Higgs et al, 2006). Diagnostic groups in the SNC are matched by descriptive variables such as age, gender, race, postmortem interval (PMI), mRNA quality, brain pH, and hemisphere.
To facilitate psychiatric and basic neuroscience research, a novel web-based database, the Stanley Neuropathology Consortium Integrative Database (SNCID; http://sncid.stanleyresearch.org), has been developed. The database currently integrates 1749 datasets from neuropathology studies, genome-wide expression microarray datasets, and statistical tools. In this report, we show several potential applications of this new integrative database. We first replicate an earlier correlation analysis between genome-wide expression profiles and an abnormal cytoarchitectural marker described in the SNC. We then identify several abnormal neurochemical markers in the diagnostic groups as well as the gene-expression profile and biological processes associated with these abnormal markers.
MATERIALS AND METHODS
Datasets Integrated in the SNCID
The SNCID currently includes a total of 1749 neuropathology markers measured in 12 different brain regions of the SNC (Supplementary Table S1). We classified the markers into four categories, such as RNA (n=719), Protein (n=315), Cell (which includes cytoarchitectural studies; n=332), and Other (n=383). The SNCID also contains genome-wide expression microarray datasets from three independent studies that used frontal cortex or cerebellum tissue from the SNC. As research is still being conducted with SNC tissue, and datasets are continually being returned to SMRI, the SNCID will continue to be updated.
Statistical Analysis Tools
All statistical tools in the database were developed based on R-packages (http://www.r-project.org/). The basic descriptive statistics for a marker are shown with a histogram by selecting analysis tool in the marker table. Users can choose the statistical method to use for further analysis based on the data distribution. The SNCID provides parametric and non-parametric statistical tools for each analysis. In this demonstration, we used non-parametric methods for all statistical analyses to avoid variations because of unit differences and/or distribution pattern differences. Results from variance analysis are illustrated with box plots to illustrate the comparison between the psychiatric disorder groups and the unaffected controls.
Potential confounding variables for each marker can be explored by using the confounding-variables tool, which include five continuous variables (age, PMI, brain pH, antipsychotic treatment, and duration of illness) and four categorical variables (sex, alcohol, drug abuse, and smoking). In addition, the effect of two psychiatric phenotypes (suicide vs non-sucide and psychotic features vs non-psychotic features) on a marker of interest can be assessed by using the tool. Statistical results are illustrated with a scatter plot or box plot for each variable. Genome-wide expression profiles associated with a marker of interest can also be explored by using the correlation analysis tool. Similar correlation analyses have been widely used to explore the relationship between two markers, between a marker and a demographic variable and between a marker and the expression level of a gene (Dracheva et al, 2006; Guidotti et al, 2000; Hashimoto et al, 2003). Each expression microarray dataset in the SNCID is normalized by two different algorithms: the Affymetrix Microarray Suite software, version 5.0 (MAS5), and robust multi-array averaging. In this study, we used the MAS5 normalized microarray datasets for genome-wide correlation analysis.
Link to Other Databases
The marker name in the query table is hyperlinked to detailed information in the NCBI Entrez gene db (http://www.ncbi.nlm.nih.gov/sites/entrez?db=gene) (Maglott et al, 2005). The study information provided by the original researcher is hyperlinked to ‘info.’ Publications resulting from the dataset are hyperlinked to ‘ref’ in the marker query table (Pubmed; http://www.ncbi.nlm.nih.gov/pubmed/). An interface links the SNCID to the DAVID database (http://david.abcc.ncifcrf.gov/home.jsp) (Dennis et al, 2003) so that probe sets that are significantly correlated with a particular marker can be functionally annotated. The enrichment analyses were conducted using the pre-built Affymetrix chip background and the functional annotations for the biological processes used Gene Ontology (GO) Consortium (http://www.geneontology.org) at all levels. p-values less than 0.05 were considered significant.
Repository Database
The SNCID contains a repository database that includes the zipped data files submitted by the original researchers. We strongly recommend that users access the raw data for further analysis after downloading from this repository database, to test complicated statistical models. To further explore the relationship of a marker (BDNF mRNA) to a specific psychiatric disorder we conducted a correlation analysis (StatView, SAS, Cary, NC) between BDNF mRNA and two clinical variables (duration of illness and antipsychotic treatment) using only cases with the disorders. The analysis was performed after downloading the raw data and demographic variables from the database.
RESULTS
Replication Analysis of Potential Molecular Mechanisms Underlying the Perineuronal Oligodendrocyte Deficits in the Frontal Cortex in Psychiatric Disorders
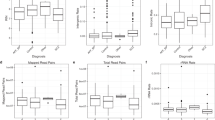
By integrating datasets from neuropathology and microarray studies with correlation analysis tools, users are able to explore genes and biological processes that may be associated with a cytoarchitectural marker at the genome-wide level. By using the SNCID, we attempted to replicate our previous results from a genome-wide analysis that identified the potential molecular mechanisms underlying the perineuronal oligodendrocyte abnormality in the prefrontal cortex (PFC) of subjects with mental disorders (Kim and Webster, 2008). Descriptive statistical tools showed the data distribution for the dataset (Figure 1a), and variance analysis tools showed that all three diagnostic groups had a significant decrease in the number of perineuronal oligodendrocytes in the PFC as compared with unaffected controls (Figure 1b). The analysis tool showed a significant decrease in the number of perineuronal oligodendrocytes in suicide completers as compared with non-completers and also in cases with psychosis as compared with those without psychosis (Figure 1c and d).
Data distribution and variance analysis for the number of perineuronal oligodendrocytes in the frontal cortex. Basic descriptive statistics and histogram for the marker (a). Box plots represent the median and distribution of the marker for each diagnostic group (b) for suicide vs non-suicide (c) and for psychotic features vs non-psychotic features (d).
A correlation analysis was then conducted with all other markers measured in the frontal cortex (Supplementary Figure S1). The correlation analysis yielded a total of 101 markers (including the marker itself) significantly correlated with the number of perineuronal oligodendrocytes in the frontal cortex (p<0.05). Several oligodendrocyte-related RNA measures derived from independent studies showed significant correlations with the cytoarchitectural marker, including the mRNA levels for the schizophrenia susceptibility gene ERBB3.
Using a genome-wide correlation analysis and functional annotation, we explored the genes and biological processes associated with the cytoarchitectural marker. Correlation analysis showed that 434 probe sets from the expression microarray data were significantly correlated with the number of perineuronal oligodendrocytes (p-value <0.001; Figure 2a). Several biological processes (GO, all levels), including amino acid metabolic processes, mitochondria-related electron transport, and protein transport, were significantly overrepresented (p<0.05; Figure 2b). A total of 3149 probe sets were correlated with the marker if we relaxed the significance level to p<0.01. Apoptosis and vesicle-mediated transport were also overrepresented in the genes correlated with the marker (Supplementary Table S2). These findings are all consistent with our earlier results (Kim and Webster, 2008).
The total number of probe sets correlated with the number of perineuronal oligodendrocytes and the significantly associated biological processes. (a) Screen shot captures the number of probe sets correlated with the perineuronal oligodendrocytes by p-values from genome-wide correlation analysis and (b) biological processes (gene ontology, all levels) overrepresented in the probe sets (n=243) significantly correlated with the marker (p<0.001). The functional annotation was done using an interface in the SNCID that links the SNCID (http://sncid.stanleyresearch.org) and DAVID (http://david.abcc.ncifcrf.gov/).
Potential Neurochemical Markers for Psychiatric Disorders
By using the datasets from the neuropathology studies and the variance analysis tool, the SNCID facilitates the exploration of potential neurochemical markers that may be associated with psychiatric disorders. We examined the RNA levels of 13 genes for which data are available in three or more different brain regions. RNA levels for BDNF, NTRK2, NTRK3, GAD1, GRIA1, GRIA2, GRIA3, GRIA4, GRIN1, GRIN2A, GRIN2B, GRIN2C, and GRIN2D were measured in three or more different postmortem brain regions in the SNCID (Table 1; Supplementary Table S3). BDNF mRNA levels were most significantly altered in the temporal cortex (p<0.05, uncorrected for multiple testing), with significant decreases in all three diagnostic groups as compared with the controls. BDNF mRNA levels were also decreased in the frontal cortex and the hippocampus in the bipolar disorder and schizophrenia groups as compared with the controls. There was no significant alteration of BDNF mRNA levels in the entorhinal cortex or in the white matter of the frontal cortex. The BDNF receptor NTRK2 mRNA levels were significantly decreased in at least two layers of the entorhinal cortex in all three diagnostic groups.
Levels of GAD1 mRNA, a marker of the γ-aminobutyric acid (GABA)ergic system, were measured in six different brain regions. GAD1 mRNA levels were abnormal in five of the six layers of the orbitofrontal cortex and in three of the five subregions of the hippocampus in at least one diagnostic group. GAD1 mRNA levels were also decreased in both subregions of the striatum in at least one diagnostic group. There were no abnormalities in GAD1 mRNA levels in the superior temporal cortex or the frontal cortex in any of the diagnostic groups.
RNA levels for nine genes related to the glutamatergic system were measured in three or more brain regions. Of the four alpha-amino-3-hydroxy-5-methyl-4-isoxazole propionate (AMPA) receptors included in the database, GRIA1 mRNA levels showed the most significant changes between the diagnostic groups and the unaffected controls. GRIA1 was abnormally expressed in 5 of the 10 different subregions in which it was measured and primarily in bipolar disorder and schizophrenia. GRIA2, GRIA3, and GRIA4 were all abnormally expressed in two or fewer subregions and primarily in bipolar disorder. RNA levels for five N-methyl-D-aspartate (NMDA) receptors, GRIN1, GRIN2A, GRIN2B, GRIN2C, and GRIN2D, were measured in three or more brain regions. Very few abnormalities were detected in NMDA expression in any diagnostic group.
Statistical tools in the SNCID enable users to identify potential confounding variables for each marker. We analyzed the correlation between antipsychotic treatment and the neurochemical markers (Supplementary Table S3). For example, though BDNF mRNA levels were significantly correlated with antipsychotic treatment in four of the six layers in the frontal cortex, there were no significant correlations in temporal cortex. Moreover, there were no significant correlations between GAD 1 mRNA levels and antipsychotic treatment in any brain areas.
Examining a particular marker across multiple brain regions in the same cohort and finding that it is differentially affected across areas and in different diagnostic groups would indicate that it is not just a nonspecific by-product of severe mental illness. For example, as described above, BDNF mRNA is decreased in all three diagnostic groups in the temporal cortex, but only in schizophrenia and bipolar disorder in the frontal cortex and hippocampus and is not affected in any disorder group in the entorhinal cortex. Moreover, the decreases are specific to particular layers in the cortices (Supplementary Table S3). Thus, it is unlikely that a decrease in BDNF mRNA is just a by-product of having severe mental illness. However, additional analysis with disorder specific clinical variables in the 45 disorder cases showed that though there is no relationship between BDNF mRNA levels in any area with duration of illness, there is a correlation with antipsychotic treatment, but only in the frontal cortex and the subiculum. Moreover, BDNF mRNA levels are decreased in the psychotic vs the non-psychotic subjects in frontal cortex but not in other areas. Thus, the analyses indicate that BDNF mRNA levels in the frontal cortex are specific to those subjects with psychosis. Whether the decrease in frontal cortex is a marker for psychosis or a by-product of taking antipsychotic medicine would have to be explored with additional (including animal model) studies. The analysis illustrates the utility of the SNCID to explore the disorder specificity of abnormalities in particular markers.
Biological Processes Associated with Dopamine or Glutamate Abnormalities in the Frontal Cortex
The major neurotransmitters, dopamine and glutamate, have both been implicated in the neuropathology of psychiatric disorders (Carlsson, 1988; Coyle, 1996; Meador-Woodruff et al, 2001). For example, dopaminergic neurons in the mesencephalic tegmentum project into the PFC, which is involved in sensory, cognitive, and affective processes (Knable and Weinberger, 1997). Moreover, antipsychotic drugs not only block dopamine receptors (Seeman and Van Tol, 1994) but induce dopamine release in the rodent PFC (Moghaddam and Bunney, 1990). However, there is little understanding of the biological processes associated with the neurotransmitter abnormalities that exist in the frontal cortex of patients with psychiatric disorders. Consequently, we explored the biological processes that may be correlated with dopamine and glutamate in the frontal cortex of subjects with psychiatric disorders. We found that dopamine levels were significantly increased in the frontal cortex of subjects with schizophrenia as compared with unaffected controls (Supplementary Figure S2a). We also found that glutamate levels were significantly increased in cases with major depression and bipolar disorder as compared with unaffected controls (Supplementary Figure S2b). In addition, the analysis tool showed that glutamate levels were increased in suicide completers as compared with non-suicide cases (p=0.05). A total of 93 and 71 markers measured in the frontal cortex were significantly correlated with dopamine and glutamate levels, respectively (p<0.05). COMT protein, a dopamine-degrading enzyme, was negatively correlated with dopamine levels (ρ=−0.336, p=0.01). RNA levels of GFAP, a marker of astrocytes, were measured in two independent studies, and both datasets were positively correlated with glutamate levels (all p<0.001). A total of 43 probe sets from the microarray dataset were significantly correlated with dopamine levels, and nine biological processes were overrepresented in the probe sets (Table 2). Processes related to response to stress, cellular metabolism, and organ development were associated with dopamine levels in the frontal cortex (p<0.05). As very few genes were highly correlated with glutamate levels (p<0.001), we relaxed the significance level to p<0.01 and showed 151 probe sets correlated with glutamate levels. Cell growth, cell death, and cell-cycle regulation were the main biological processes significantly associated with glutamate levels in the frontal cortex (Table 3).
Comparative Analysis of the Biological Processes Associated with GABAergic Abnormalities in both the Frontal Cortex and the Cerebellum
Integration of multiple datasets for a marker measured in several different brain regions, together with microarray data derived from the same brain regions, provides researchers with an opportunity to conduct comparative analysis of differential abnormalities between individual brain regions. We compared the biological processes associated with GABAergic abnormalities that occur in the frontal cortex as compared with the cerebellum. In the frontal cortex, GAD67 proteins were significantly reduced in subjects with bipolar disorder and schizophrenia as compared with unaffected controls. In contrast, there was no significant difference in GAD65 protein levels in any of the diagnostic groups compared with controls. The correlation analysis tool yielded a total of 71 and 53 probe sets from the microarray dataset that were significantly correlated with GAD67 and GAD65 protein levels in the frontal cortex, respectively (p<0.01). Although RNA metabolism was the process significantly correlated with GAD67 protein levels (Figure 3a), it was tissue development that was significantly associated with GAD65 protein levels (Figure 3b).
Biological processes (gene ontology, all levels) overrepresented in the probe sets significantly correlated with GAD67 (a) and GAD65 protein levels (b) in the frontal cortex. The functional annotation was done using an interface in the SNCID that links the SNCID (http://sncid.stanleyresearch.org) and DAVID (http://david.abcc.ncifcrf.gov/).
In contrast to the frontal cortex, in the cerebellum both GAD67 and GAD65 protein levels were significantly reduced in all three diagnostic groups as compared with unaffected controls. A total of 59 and 51 probe sets were significantly correlated with GAD67 and GAD65 protein levels in the cerebellum, respectively (p<0.01). Cell motility and tissue development were significantly associated with both GAD67 and GAD65 protein levels in the cerebellum (Supplementary Figure S3a, b).
Genes and Neurochemical Markers Associated with Reelin and Parvalbumin
Reelin and parvalbumin were the markers most significantly altered in the psychiatric disorders in our earlier meta-analysis (Torrey et al, 2005). Correlation analyses were conducted to identify neurochemical markers, demographic variables, and pathways associated with these markers. Reelin mRNA (RELN) was significantly decreased in the frontal cortex and cerebellum of the schizophrenia and bipolar disorder groups as compared with the normal control group (all p<0.01). In contrast, these markers did not change significantly in the occipital cortex of any disorder group. A total of 126 markers (including the marker itself) correlated significantly with RELN in the frontal cortex (p<0.05). Interestingly, mRNA levels of OLIG2 (an oligodendrocyte marker) and the density of oligodendrocytes and parvalbumin-positive neurons were both highly significantly correlated with RELN (p<0.001) (Supplementary Figure S4). A genome-wide correlation analysis was then conducted to explore the potential mechanisms underlying the reduced expression of RELN. A total of 76 probe sets were significantly correlated with RELN levels in the frontal cortex (p< 0.01) and a total of 17 biological processes such as transcription/regulation of viral genome replication and immune system processes were overrepresented in the probe sets (Supplementary Table S4). This suggests that epigenetic controls such as an immune response by viral infection may underlie the decrease in RELN that occurs in the frontal cortex of psychiatric disorders. This is also consistent with the earlier finding that prenatal viral infections lead to a reduction in reelin expression in the cortex of neonatal mice (Fatemi et al, 1999).
The density of parvalbumin-positive GABAgergic neurons was significantly reduced in the CA2 subfield of the hippocampus in schizophrenia subjects (p<0.0001). A total of 35 RNA markers significantly correlated with the density of parvalbumin-positive neurons in the hippocampus (Supplementary Table S5). Several glutamate receptors in the same brain region such as GRIA1, GRIA2, GRIA3, and GRIN2B were positively correlated with the marker, suggesting a functional relationship between the reduced density of parvalbumin containing GABAergic neurons and AMPA and/or NMDA receptors in the hippocampus of schizophrenia patients.
DISCUSSION
The development of an integrative database will facilitate psychiatric studies by encouraging data sharing and providing user-friendly data-mining tools. Here, we give a basic description of the database in its initial stages and show several examples of potential applications. We successfully replicated a previous correlation analysis between genome-wide expression profiles and an abnormal cytoarchitectural marker (decreased number of perineuronal oligodendrocytes in schizophrenia) described in the SNC. We then explored the database for RNA markers that may be abnormally expressed in psychiatric disorders. We found that many RNA markers showed regional specific differences between the diagnostic groups and unaffected controls. In fact, there is no RNA marker that was abnormally expressed in all brain areas in which it was measured.
Although deficits in several neurotransmitter systems have been widely reported in psychiatric disorders, the biological pathways associated with the deficits are largely unexplored. In this study, we explored the genes and biological processes associated with the neurotransmitter abnormalities in the psychiatric disorders, using the SNCID at the genome-wide level. Regulation of metabolic processes was most significantly associated with dopamine levels in the frontal cortex. This is consistent with earlier findings of global metabolic abnormalities in schizophrenia (Prabakaran et al, 2004) as well as in bipolar disorder (Iwamoto et al, 2005). Moreover, our results suggest that the dopamine abnormality may contribute to the metabolic perturbations of major psychiatric disorders.
L-glutamate is one of the main excitatory neurotransmitters in the CNS (Robinson and Coyle, 1987) and mediates synaptic transmission, plasticity, and neurotoxicity (Ozawa et al, 1998). Our analysis showed that cell growth and apoptosis are the main biological pathways associated with glutamate levels in the frontal cortex. Moreover, cytoarchitectural abnormalities in the PFC are one of the most significant reproducible findings in schizophrenia (Harrison, 1999), bipolar disorder, (Rajkowska, 2002), and depression (Rajkowska, 2003). Earlier postmortem studies have shown reduced cell density of GABAergic interneurons (Reynolds et al, 2001), reduced density of oligodendrodendrocytes (Uranova et al, 2004), reduced numbers of perineuronal oligodendrocytes (Vostrikov et al, 2007), and decreased cell size of pyramidal neurons (Vostrikov et al, 2007) in the PFC of subjects with major psychiatric disorders. Our previous genome-wide correlation analysis also showed apoptosis to be one of the potential mechanisms that may underlie the decrease in the number of perineuronal oligodendrocytes in the PFC of subjects with major mental disorders (Kim and Webster, 2008) and this finding was replicated in this demonstration using the SNCID. Collectively, the results suggest that apoptosis, perhaps associated with cytotoxicity mediated by a glutamate abnormality, may be one of the causes of reduced numbers of oligodendrocytes in the PFC of major psychiatric disorders.
The SNCID allows researchers to examine molecules associated with neurotransmitter abnormalities in psychiatric disorders by exploring genes and biological pathways associated with, for example, key enzymes involved in the synthesis of the neurotransmitters. We compared the biological processes associated with GAD67 and GAD65 protein levels, two key enzymes for GABA synthesis, in the frontal cortex and the cerebellum. The biological processes associated with GAD65 levels are very similar in both the frontal cortex and the cerebellum. Tissue development is the main biological process associated with GAD65 levels in both brain areas. However, the pathways associated with GAD67 are different in the two brain regions. Although tissue development is also the process significantly correlated with GAD67 protein levels in the cerebellum, in the frontal cortex it is RNA metabolism that is most significantly associated with GAD67 protein levels. Thus, the processes associated with the GABA abnormalities mediated by GAD67 in the two brain regions are different and may reflect the differential involvement of the two areas in the pathogenesis of psychiatric disorders. Moreover, the different intracellular localization of GAD67 and GAD65 indicates that the two enzymes have distinct roles in the GABAergic neurons (Martin and Rimvall, 1993) and may reflect the different processes we find associated with them in the frontal cortex. However, the cellular localization and functional roles of the two enzymes are likely to differ in the frontal cortex and the cerebellum, which is a relatively understudied brain area in psychiatric research.
Although the SNCID provides efficient data-exploring tools for major psychiatric disorders, researchers must understand that there are several limitations in the current version. First, because datasets deposited in the SNCID were generated in multiple laboratories all over the world, there will be technical variations in the datasets. However, we have observed significant correlations between the markers that should be correlated for biological reasons even though they were generated from independent studies. For instance, RNA markers for oligodendrocytes that were generated by independent researchers using different techniques were significantly correlated with the number of perineuronal oligodendrocytes in the frontal cortex. Moreover, the RNA studies were done in frozen tissue from the opposite hemisphere to the cytoarchitectural study that counted the oligodendrocytes in the fixed hemisphere. In addition, protein levels of COMT, a major enzyme for dopamine degradation, were negatively correlated with dopamine levels. This supports the feasibility of an integrative analysis with datasets deposited in the SNCID. However, we also found an absence of correlation between the datasets for the same marker that was measured in the same brain region but by independent laboratories. For example, there are two independent datasets for GAD67 and GAD65 protein that were measured in the cerebellum; however there is no significant correlation between the two datasets. Thus, researchers should be cautious when interpreting the results and should find consensus if there are multiple datasets available in the database. Second, the current version of the database provides omnibus ANOVA and correlation analysis without covariance to maximize performance. Thus, the results derived from the SNCID are exploratory, and we strongly recommended that users follow up any interesting findings by downloading the raw datasets from the repository sub-database and examining them with more sophisticated statistical models. As data continue to be generated on the 60 cases in the SNCID, we will add it to the database, and eventually additional modules containing SNP array data, microRNA array data, and methylation array data will be integrated into the system. We believe this integrative database will give researchers a unique opportunity to explore the abnormal neuropathological markers that occur in the major psychiatric disorders and will provide the data and tools necessary to explore the genes and biological processes associated with those abnormal markers.
References
Abdeltawab NF, Aziz RK, Kansal R, Rowe SL, Su Y, Gardner L et al (2008). An unbiased systems genetics approach to mapping genetic loci modulating susceptibility to severe streptococcal sepsis. PLoS Pathog 4: e1000042.
Ball CA, Sherlock G, Brazma A (2004). Funding high-throughput data sharing. Nat Biotechnol 22: 1179–1183.
Bystrykh L, Weersing E, Dontje B, Sutton S, Pletcher MT, Wiltshire T et al (2005). Uncovering regulatory pathways that affect hematopoietic stem cell function using ‘genetical genomics’. Nat Genet 37: 225–232.
Carlsson A (1988). The current status of the dopamine hypothesis of schizophrenia. Neuropsychopharmacology 1: 179–186.
Carneiro AM, Airey DC, Thompson B, Zhu CB, Lu L, Chesler EJ et al (2009). Functional coding variation in recombinant inbred mouse lines reveals multiple serotonin transporter-associated phenotypes. Proc Natl Acad Sci USA 106: 2047–2052.
Coyle JT (1996). The glutamatergic dysfunction hypothesis for schizophrenia. Harv Rev Psychiatry 3: 241–253.
Dennis Jr G, Sherman BT, Hosack DA, Yang J, Gao W, Lane HC et al (2003). DAVID: Database for Annotation, Visualization, and Integrated Discovery. Genome Biol 4: P3.
Dracheva S, Davis KL, Chin B, Woo DA, Schmeidler J, Haroutunian V (2006). Myelin-associated mRNA and protein expression deficits in the anterior cingulate cortex and hippocampus in elderly schizophrenia patients. Neurobiol Dis 21: 531–540.
Fatemi SH, Emamian ES, Kist D, Sidwell RW, Nakajima K, Akhter P et al (1999). Defective corticogenesis and reduction in Reelin immunoreactivity in cortex and hippocampus of prenatally infected neonatal mice. Mol Psychiatry 4: 145–154.
Gordon E, Cooper N, Rennie C, Hermens D, Williams LM (2005). Integrative neuroscience: the role of a standardized database. Clin EEG Neurosci 36: 64–75.
Guidotti A, Auta J, Davis JM, Di-Giorgi-Gerevini V, Dwivedi Y, Grayson DR et al (2000). Decrease in reelin and glutamic acid decarboxylase67 (GAD67) expression in schizophrenia and bipolar disorder: a postmortem brain study. Arch Gen Psychiatry 57: 1061–1069.
Harrison PJ (1999). The neuropathology of schizophrenia. A critical review of the data and their interpretation. Brain 122 (Pt 4): 593–624.
Hashimoto T, Volk DW, Eggan SM, Mirnics K, Pierri JN, Sun Z et al (2003). Gene expression deficits in a subclass of GABA neurons in the prefrontal cortex of subjects with schizophrenia. J Neurosci 23: 6315–6326.
Higgs BW, Elashoff M, Richman S, Barci B (2006). An online database for brain disease research. BMC Genomics 7: 70.
Iwamoto K, Bundo M, Kato T (2005). Altered expression of mitochondria-related genes in postmortem brains of patients with bipolar disorder or schizophrenia, as revealed by large-scale DNA microarray analysis. Hum Mol Genet 14: 241–253.
Kim S, Webster MJ (2008). Correlation analysis between genome-wide expression profiles and cytoarchitectural abnormalities in the prefrontal cortex of psychiatric disorders. Mol Psychiatry advance, print copy in press (originally published online September 2, 2008 (DOI:10.1038/mp 2008.99).
Knable MB, Weinberger DR (1997). Dopamine, the prefrontal cortex and schizophrenia. J Psychopharmacol 11: 123–131.
Maglott D, Ostell J, Pruitt KD, Tatusova T (2005). Entrez Gene: gene-centered information at NCBI. Nucleic Acids Res 33 (Database issue): D54–D58.
Martin DL, Rimvall K (1993). Regulation of gamma-aminobutyric acid synthesis in the brain. J Neurochem 60: 395–407.
Meador-Woodruff JH, Hogg Jr AJ, Smith RE (2001). Striatal ionotropic glutamate receptor expression in schizophrenia, bipolar disorder, and major depressive disorder. Brain Res Bull 55: 631–640.
Moghaddam B, Bunney BS (1990). Acute effects of typical and atypical antipsychotic drugs on the release of dopamine from prefrontal cortex, nucleus accumbens, and striatum of the rat: an in vivo microdialysis study. J Neurochem 54: 1755–1760.
Ozawa S, Kamiya H, Tsuzuki K (1998). Glutamate receptors in the mammalian central nervous system. Prog Neurobiol 54: 581–618.
Prabakaran S, Swatton JE, Ryan MM, Huffaker SJ, Huang JT, Griffin JL et al (2004). Mitochondrial dysfunction in schizophrenia: evidence for compromised brain metabolism and oxidative stress. Mol Psychiatry 9: 684–697, 643.
Rajkowska G (2002). Cell pathology in bipolar disorder. Bipolar Disord 4: 105–116.
Rajkowska G (2003). Depression: what we can learn from postmortem studies. Neuroscientist 9: 273–284.
Reynolds GP, Zhang ZJ, Beasley CL (2001). Neurochemical correlates of cortical GABAergic deficits in schizophrenia: selective losses of calcium binding protein immunoreactivity. Brain Res Bull 55: 579–584.
Robinson MB, Coyle JT (1987). Glutamate and related acidic excitatory neurotransmitters: from basic science to clinical application. FASEB J 1: 446–455.
Seeman P, Van Tol HH (1994). Dopamine receptor pharmacology. Trends Pharmacol Sci 15: 264–270.
Torrey EF, Barci BM, Webster MJ, Bartko JJ, Meador-Woodruff JH, Knable MB (2005). Neurochemical markers for schizophrenia, bipolar disorder, and major depression in postmortem brains. Biol Psychiatry 57: 252–260.
Torrey EF, Webster MJ, Knable M, Johnston N, Yolken RH (2000). The Stanley Foundation brain collection and Neuropathology Consortium. Schizophr Res 44: 151–155.
Uranova NA, Vostrikov VM, Orlovskaya DD, Rachmanova VI (2004). Oligodendroglial density in the prefrontal cortex in schizophrenia and mood disorders: a study from the Stanley Neuropathology Consortium. Schizophr Res 67: 269–275.
Vostrikov VM, Uranova NA, Orlovskaya DD (2007). Deficit of perineuronal oligodendrocytes in the prefrontal cortex in schizophrenia and mood disorders. Schizophr Res 94: 273–280.
Wang J, Williams RW, Manly KF (2003). WebQTL: web-based complex trait analysis. Neuroinformatics 1: 299–308.
Zhang B, Kirov S, Snoddy J (2005). WebGestalt: an integrated system for exploring gene sets in various biological contexts. Nucleic Acids Res 33 (Web Server issue): W741–W748.
Zhang W, Zhang Y, Zheng H, Zhang C, Xiong W, Olyarchuk JG et al (2007). SynDB: a Synapse protein DataBase based on synapse ontology. Nucleic Acids Res 35 (Database issue): D737–D741.
Acknowledgements
We thank all the investigators who generated the original data in the SNCID, and their many collaborators, who made this database possible. We also thank all the technicians in the SMRI brain laboratory who prepared the brain tissues and extracted the RNA and DNA from the tissues. We specially thank the Keymind Company for their technical assistance with the database, in particular Marvin Suo.
Author information
Authors and Affiliations
Corresponding author
Additional information
DISCLOSURE
The authors declare no conflict of interest.
Supplementary Information accompanies the paper on the Neuropsychopharmacology website (http://www.nature.com/npp)
Supplementary information
Rights and permissions
About this article
Cite this article
Kim, S., Webster, M. The Stanley Neuropathology Consortium Integrative Database: a Novel, Web-Based Tool for Exploring Neuropathological Markers in Psychiatric Disorders and the Biological Processes Associated with Abnormalities of Those Markers. Neuropsychopharmacol 35, 473–482 (2010). https://doi.org/10.1038/npp.2009.151
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/npp.2009.151
Keywords
This article is cited by
-
Overexpression of transmembrane TNFα in brain endothelial cells induces schizophrenia-relevant behaviors
Molecular Psychiatry (2023)
-
Novel open reading frames in human accelerated regions and transposable elements reveal new leads to understand schizophrenia and bipolar disorder
Molecular Psychiatry (2022)
-
The Stanley Neuropathology Consortium Integrative Database (SNCID) for Psychiatric Disorders
Neuroscience Bulletin (2019)
-
Comprehensive integrative analyses identify GLT8D1 and CSNK2B as schizophrenia risk genes
Nature Communications (2018)
-
The Gene Encoding Protocadherin 9 (PCDH9), a Novel Risk Factor for Major Depressive Disorder
Neuropsychopharmacology (2018)