Abstract
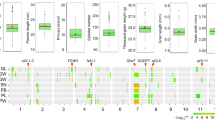
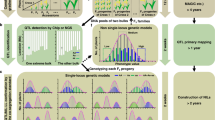
Grain size is one of the most important components of grain yield and selecting large seeds has been a main target during plant domestication. Surprisingly, the grain of African cultivated rice (Oryza glaberrima Steud.) typically is smaller than that of its progenitor, Oryza barthii. Here we report the cloning and characterization of a quantitative trait locus, GL4, controlling the grain length on chromosome 4 in African rice, which regulates longitudinal cell elongation of the outer and inner glumes. Interestingly, GL4 also controls the seed shattering phenotype like its orthologue SH4 gene in Asian rice. Our data show that a single-nucleotide polymorphism (SNP) mutation in the GL4 gene resulted in a premature stop codon and led to small seeds and loss of seed shattering during African rice domestication. These results provide new insights into diverse domestication practices in African rice, and also pave the way for enhancing crop yield to meeting the challenge of cereal demand in West Africa.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
United Nations-Department of Economic and Social Affairs. World Population Prospects, The 2015 Revision (United Nations, New York, 2015).
Martin, K. I. et al. Can sub-Saharan Africa feed itself? Proc. Natl Acad. Sci. USA 113, 14964–14969 (2016).
Sweeney, M. & McCouch, S. The complex history of the domestication of rice. Ann. Bot. 100, 951–957 (2007).
Harlan, J., De Wet, J. & Stemler, A. in Origins of African Plant Domestication (ed. Harlan, J. R. ) 3–20 (Mouton Publishers, 1976).
Wang, M. et al. The genome sequence of African rice (Oryza glaberrima) and evidence for independent domestication. Nat. Genet. 46, 982–988 (2014).
Meyer, R. S. et al. Domestication history and geographical adaptation inferred from a SNP map of African rice. Nat. Genet. 48, 1083–1088 (2016).
Linares, O. F. African rice (Oryza glaberrima): history and future potential. Proc. Natl Acad. Sci. USA 99, 16360–16365 (2002).
Rhodes, E. R., Jalloh, A. & Diouf, A. Review of Research and Policy for Climate Change Adaptation in the Agriculture Sector of West Africa (AfricaInteract/FAC, 2014).
Jones, M. P., Dingkuhn, M., Aluko, G. K. & Semon, M. Interspecific Oryza sativa L. x O. glaberrima Steud. progenies in upland rice improvement. Euphytica 92, 237–246 (1997).
Sarla, N. & Swamy, B. P. Oryza glaberrima: a source for the improvement of Oryza sativa. Curr. Sci. 89, 955–963 (2005).
Dorian, F. & Robin, A. Seed dispersal and crop domestication: shattering, germination and seasonality in evolution under cultivation. Annu. Plant Rev. 38, 238–295 (2009).
Meyer, R. S. & Purugganan, M. D. Evolution of crop species: genetics of domestication and diversification. Nat. Rev. Genet. 14, 840–852 (2013).
Gegas, V. C. et al. A genetic framework for grain size and shape variation in wheat. Plant Cell 22, 1046–1056 (2010).
Han, L. et al. Fine mapping of qGW1, a major QTL for grain weight in sorghum. Theor. Appl. Genet. 128, 1813–1825 (2015).
Zuo, J. R. & Li, J. Y. Molecular genetic dissection of quantitative trait loci regulating rice grain size. Annu. Rev. Genet. 48, 99–118 (2014).
Song, X., Huang, W., Shi, M., Zhu, M. & Lin, H. A QTL for rice grain width and weight encodes a previously unknown RING-type E3 ubiquitin ligase. Nat. Genet. 39, 623–630 (2007).
Shomura, A. et al. Deletion in a gene associated with grain size increased yields during rice domestication. Nat. Genet. 40, 1023–1028 (2008).
Li, Y. et al. Natural variation in GS5 plays an important role in regulating grain size and yield in rice. Nat. Genet. 43, 1266–1269 (2011).
Ishimaru, K. et al. Loss of function of the IAA-glucose hydrolase gene TGW6 enhances rice grain weight and increases yield. Nat. Genet. 45, 707–711 (2013).
Wang, S. et al. Control of grain size, shape and quality by OsSPL16 in rice. Nat. Genet. 44, 950–954 (2012).
Che, R. et al. Control of grain size and rice yield by GL2-mediated brassinosteroid responses. Nat. Plants 2, 15195 (2015).
Duan, P. et al. Regulation of OsGRF4 by OsmiR396 controls grain size and yield in rice. Nat. Plants 2, 15203 (2015).
Wang, Y. et al. Copy number variation at the GL7 locus contributes to grain size diversity in rice. Nat. Genet. 47, 944–948 (2015).
Wang, S. et al. The OsSPL16-GW7 regulatory module determines grain shape and simultaneously improves rice yield and grain quality. Nat. Genet. 47, 949–954 (2015).
Mao, H. et al. Linking differential domain functions of the GS3 protein to natural variation of grain size in rice. Proc. Natl Acad. Sci. USA 107, 19579–19584 (2010).
Si, L. et al. OsSPL13 control grain size in cultivated rice. Nat. Genet. 48, 447–456 (2016).
Katayama, T. C. & Sumi, A. Studies on agronomic traits of Africa rice (Oryza glaberrima steud.), 3: some grain morphological aspects of domestication and decrement. Jpn. J. Crop. Sci. 64, 807–814 (1995).
Li, C., Zhou, A. & Sang, T. Rice domestication by reducing shattering. Science 311, 1936–1939 (2006).
Lin, Z. et al. Origin of seed shattering in rice (Oryza sativa L.). Planta 226, 11–20 (2007).
Huang, R. Y. et al. Genetic bases of rice grain shape: so many genes, so little known. Trends Plant Sci. 18, 218–226 (2013).
Tang, W. et al. SNP-based analysis of genetic diversity reveals important alleles associated with seed size in rice. BMC Plant Biol. 16, 1 (2016).
Takano-Kai, N. et al. Evolutionary history of GS3, a gene conferring grain length in rice. Genetics 182, 1323–1334 (2009).
Rodenburg, J. et al. Weed competitiveness of the lowland rice varieties of NERICA in the southern Guinea savanna. Field Crop Res. 114, 411–418 (2009).
Agnoun, Y. et al. The African rice Oryza glaberrima steud: knowledge distribution and prospects. Int. J. Biol. 4, 158–180 (2012).
Balasubramanian, V., Sie, M., Hijmans, R. J. & Otsuka, K. Increasing rice production in Sub-Saharan Africa: challenges and opportunities. Adv. Agron. 94, 79–80 (2007).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler Transform. Bioinformatics 25, 1754–1760 (2009).
McKenna, A. et al. The genome analysis toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Patterson, N., Price, A. L. & Reich, D. Population structure and eigenanalysis. PLoS Genet. 2, 2074–2093 (2006).
Browning, S. R. & Browning, B. L. Rapid and accurate haplotype phasing and missing data inference for whole genome association studies by use of localized haplotype clustering. Am. J. Hum. Genet. 81, 1084–1097 (2007).
Browning, B. L. & Browning, S. R. Genotype imputation with millions of reference samples. Am. J. Hum. Genet. 98, 116–126 (2016).
Gautier, M., Klassmann, A. & Vitali, R. REHH 2.0: a reimplementation of the R package REHH to detect positive selection from haplotype structure. Mol. Ecol. Resour. 17, 78–90 (2017).
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011).
Acknowledgements
We thank the International Rice Research Institute and the United States Department of Agriculture Research Service for providing the wild rice and cultivated rice samples. This research was supported by the National Key R&D Program for Crop Breeding (2016YFD0100901), the Ministry of Agriculture of China (2016ZX08009-003) and the National Natural Science Foundation of China (Grant 31471457). We acknowledge the NSF Plant Genome Program for support to Meyer (1202803). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
Z.Z. and R.A.W. designed and supervised this study. W.W. conducted characterization of introgression line, map-based cloning, genetic transformation, gene expression analysis. W.W. and X. Liu constructed the introgression lines. M.W., R.S.M. and J.Z. performed evolutionary analysis of GL4. X. Luo bred the NIL-GL4Og. M.-N.N., L.T., J.W., H.C., C.S. and X.W. conducted the collection of rice germplasm and phenotypic data. Z.Z., R.S.M. and R.A.W. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Figures 1–14, Supplementary Table 1–4. (PDF 1564 kb)
Rights and permissions
About this article
Cite this article
Wu, W., Liu, X., Wang, M. et al. A single-nucleotide polymorphism causes smaller grain size and loss of seed shattering during African rice domestication. Nature Plants 3, 17064 (2017). https://doi.org/10.1038/nplants.2017.64
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/nplants.2017.64
This article is cited by
-
SpykProps: an imaging pipeline to quantify architecture in unilateral grass inflorescences
Plant Methods (2023)
-
Molecular mechanisms of adaptive evolution in wild animals and plants
Science China Life Sciences (2023)
-
Seed size, an imperative trait for seed vigor and drought tolerance in rice
Cereal Research Communications (2023)
-
Precisely mapping a major QTL for grain weight on chromosome 5B of the founder parent Chuanmai42 in the wheat-growing region of southwestern China
Theoretical and Applied Genetics (2023)
-
Genes and Their Molecular Functions Determining Seed Structure, Components, and Quality of Rice
Rice (2022)