Abstract
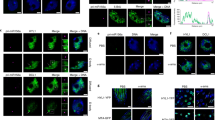
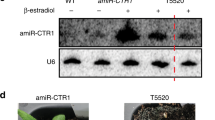
MicroRNAs (miRNAs) are a class of small non-coding RNAs that play important regulatory roles in gene expression in plants and animals. The biogenesis of miRNAs involves the transcription of primary miRNAs (pri-miRNAs) by RNA polymerase II (RNAPII) and subsequent processing by Dicer or Dicer-like (DCL) proteins. Here we show that the Elongator complex is involved in miRNA biogenesis in Arabidopsis. Disruption of Elongator reduces RNAPII occupancy at miRNA loci and pri-miRNA transcription. We also show that Elongator interacts with the DCL1-containing Dicing complex and lack of Elongator impairs DCL1 localization in the nuclear Dicing body. Finally, we show that pri-miRNA transcripts as well as DCL1 associate with the chromatin of miRNA genes and the chromatin association of DCL1 is compromised in the absence of Elongator. Our results suggest that Elongator functions in both transcription and processing of pri-miRNAs and probably couples these two processes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Rogers, K. & Chen, X. Biogenesis, turnover, and mode of action of plant microRNAs. Plant Cell 25, 2383–2399 (2013).
Voinnet, O. Origin, biogenesis, and activity of plant microRNAs. Cell 136, 669–687 (2009).
Bartel, D. P. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116, 281–297 (2004).
Kim, V. N. MicroRNA biogenesis: coordinated cropping and dicing. Nat. Rev. Mol. Cell. Biol. 6, 376–385 (2005).
Kurihara, Y. & Watanabe, Y. Arabidopsis micro-RNA biogenesis through Dicer-like 1 protein functions. Proc. Natl Acad. Sci. USA 101, 12753–12758 (2004).
Park, M. Y., Wu, G., Gonzalez-Sulser, A., Vaucheret, H. & Poethig, R. S. Nuclear processing and export of microRNAs in Arabidopsis. Proc. Natl Acad. Sci. USA 102, 3691–3696 (2005).
Dong, Z., Han, M. H. & Fedoroff, N. The RNA-binding proteins HYL1 and SE promote accurate in vitro processing of pri-miRNA by DCL1. Proc. Natl Acad. Sci. USA 105, 9970–9975 (2008).
Liu, Q., Shi, L. & Fang, Y. Dicing bodies. Plant Physiol. 158, 61–66 (2012).
Yu, B. et al. The FHA domain proteins DAWDLE in Arabidopsis and SNIP1 in humans act in small RNA biogenesis. Proc. Natl Acad. Sci. USA 105, 10073–10078 (2008).
Ren, G. et al. Regulation of miRNA abundance by RNA binding protein TOUGH in Arabidopsis. Proc. Natl Acad. Sci. USA 109, 12817–12821 (2012).
Laubinger, S. et al. Dual roles of the nuclear cap-binding complex and SERRATE in pre-mRNA splicing and microRNA processing in Arabidopsis thaliana. Proc. Natl Acad. Sci. USA 105, 8795–8800 (2008).
Kim, S. et al. Two cap-binding proteins CBP20 and CBP80 are involved in processing primary MicroRNAs. Plant Cell Physiol. 49, 1634–1644 (2008).
Gregory, B. D. et al. A link between RNA metabolism and silencing affecting Arabidopsis development. Dev. Cell 14, 854–866 (2008).
Wu, X. et al. A role for the RNA-binding protein MOS2 in microRNA maturation in Arabidopsis. Cell Res. 23, 645–657 (2013).
Wang, L. et al. NOT2 proteins promote polymerase II-dependent transcription and interact with multiple MicroRNA biogenesis factors in Arabidopsis. Plant Cell 25, 715–727 (2013).
Zhang, S., Xie, M., Ren, G. & Yu, B. CDC5, a DNA binding protein, positively regulates posttranscriptional processing and/or transcription of primary microRNA transcripts. Proc. Natl Acad. Sci. USA 110, 17588–17593 (2013).
Zhang, S., Liu, Y. & Yu, B. PRL1, an RNA-binding protein, positively regulates the accumulation of miRNAs and siRNAs in Arabidopsis. PLoS Genet. 10, e1004841 (2014).
Baumberger, N. & Baulcombe, D. C. Arabidopsis ARGONAUTE1 is an RNA Slicer that selectively recruits microRNAs and short interfering RNAs. Proc. Natl Acad. Sci. USA 102, 11928–11933 (2005).
Qi, Y., Denli, A. M. & Hannon, G. J. Biochemical specialization within Arabidopsis RNA silencing pathways. Mol. Cell 19, 421–428 (2005).
Lauressergues, D. et al. Primary transcripts of microRNAs encode regulatory peptides. Nature 520, 90–93 (2015).
Pawlicki, J. M. & Steitz, J. A. Primary microRNA transcript retention at sites of transcription leads to enhanced microRNA production. J. Cell Biol. 182, 61–76 (2008).
Kim, Y. K. & Kim, V. N. Processing of intronic microRNAs. EMBO J. 26, 775–783 (2007).
Morlando, M. et al. Primary microRNA transcripts are processed co-transcriptionally. Nature Struct. Mol. Biol. 15, 902–909 (2008).
Otero, G. et al. Elongator, a multisubunit component of a novel RNA polymerase II holoenzyme for transcriptional elongation. Mol. Cell 3, 109–118 (1999).
Wittschieben, B. O. et al. A novel histone acetyltransferase is an integral subunit of elongating RNA polymerase II holoenzyme. Mol. Cell 4, 123–128 (1999).
Rahl, P. B., Chen, C. Z. & Collins, R. N. Elp1p, the yeast homolog of the FD disease syndrome protein, negatively regulates exocytosis independently of transcriptional elongation. Mol. Cell 17, 841–853 (2005).
Huang, B., Johansson, M. J. & Bystrom, A. S. An early step in wobble uridine tRNA modification requires the Elongator complex. RNA 11, 424–436 (2005).
Krogan, N. J. & Greenblatt, J. F. Characterization of a six-subunit holo-elongator complex required for the regulated expression of a group of genes in Saccharomyces cerevisiae. Mol. Cell Biol. 21, 8203–8212 (2001).
DeFraia, C. & Mou, Z. The role of the Elongator complex in plants. Plant Signal. Behav. 6, 19–22 (2011).
Nelissen, H. et al. Plant Elongator regulates auxin-related genes during RNA polymerase II transcription elongation. Proc. Natl Acad. Sci. USA 107, 1678–1683 (2010).
Wang, W. et al. An importin β protein negatively regulates MicroRNA activity in Arabidopsis. Plant Cell 23, 3565–3576 (2011).
Nelissen, H. et al. The elongata mutants identify a functional Elongator complex in plants with a role in cell proliferation during organ growth. Proc. Natl Acad. Sci. USA 102, 7754–7759 (2005).
Zhou, X., Hua, D., Chen, Z., Zhou, Z. & Gong, Z. Elongator mediates ABA responses, oxidative stress resistance and anthocyanin biosynthesis in Arabidopsis. Plant J. 60, 79–90 (2009).
Sunkar, R., Kapoor, A. & Zhu, J. K. Posttranscriptional induction of two Cu/Zn superoxide dismutase genes in Arabidopsis is mediated by downregulation of miR398 and important for oxidative stress tolerance. Plant Cell 18, 2051–2065 (2006).
Allen, R. S. et al. Genetic analysis reveals functional redundancy and the major target genes of the Arabidopsis miR159 family. Proc. Natl Acad. Sci. USA 104, 16371–16376 (2007).
Fang, Y. & Spector, D. L. Identification of nuclear dicing bodies containing proteins for microRNA biogenesis in living Arabidopsis plants. Curr. Biol. 17, 818–823 (2007).
Song, L., Han, M. H., Lesicka, J. & Fedoroff, N. Arabidopsis primary microRNA processing proteins HYL1 and DCL1 define a nuclear body distinct from the Cajal body. Proc. Natl Acad. Sci. USA 104, 5437–5442 (2007).
Hamilton, A., Voinnet, O., Chappell, L. & Baulcombe, D. Two classes of short interfering RNA in RNA silencing. EMBO J. 21, 4671–4679 (2002).
Sobell, H. M. Actinomycin and DNA transcription. Proc. Natl Acad. Sci. USA 82, 5328–5331 (1985).
Zheng, B. et al. Intergenic transcription by RNA polymerase II coordinates Pol IV and Pol V in siRNA-directed transcriptional gene silencing in Arabidopsis. Genes Dev. 23, 2850–2860 (2009).
Tagami, Y., Motose, H. & Watanabe, Y. A dominant mutation in DCL1 suppresses the hyl1 mutant phenotype by promoting the processing of miRNA. RNA 15, 450–458 (2009).
Hawkes, N. A. et al. Purification and characterization of the human elongator complex. J. Biol. Chem. 277, 3047–3052 (2002).
Ben Chaabane, S. et al. STA1, an Arabidopsis pre-mRNA processing factor 6 homolog, is a new player involved in miRNA biogenesis. Nucleic Acids Res. 41, 1984–1997 (2013).
Esberg, A., Huang, B., Johansson, M. J. & Bystrom, A. S. Elevated levels of two tRNA species bypass the requirement for elongator complex in transcription and exocytosis. Mol. Cell 24, 139–148 (2006).
Creppe, C. & Buschbeck, M. Elongator: an ancestral complex driving transcription and migration through protein acetylation. J. Biomed. Biotechnol. 2011, 924898 (2011).
Svejstrup, J. Q. Elongator complex: how many roles does it play? Curr. Opin. Cell Biol. 19, 331–336 (2007).
Wang, Y. et al. The Arabidopsis elongator complex subunit2 epigenetically regulates plant immune responses. Plant Cell 25, 762–776 (2013).
Kruk, J. A., Dutta, A., Fu, J., Gilmour, D. S. & Reese, J. C. The multifunctional Ccr4-Not complex directly promotes transcription elongation. Genes Dev. 25, 581–593 (2011).
Dugas, D. V. & Bartel, B. Sucrose induction of Arabidopsis miR398 represses two Cu/Zn superoxide dismutases. Plant Mol. Biol. 67, 403–417 (2008).
Schneeberger, K. et al. SHOREmap: simultaneous mapping and mutation identification by deep sequencing. Nature Methods 6, 550–551 (2009).
Earley, K. W. et al. Gateway-compatible vectors for plant functional genomics and proteomics. Plant J. 45, 616–629 (2006).
Manavella, P. A. et al. Fast-forward genetics identifies plant CPL phosphatases as regulators of miRNA processing factor HYL1. Cell 151, 859–870 (2012).
Kim, Y. J. et al. The role of Mediator in small and long noncoding RNA production in Arabidopsis thaliana. EMBO J. 30, 814–822 (2011).
Zhang, H. et al. An Rrp6-like protein positively regulates noncoding RNA levels and DNA methylation in Arabidopsis. Mol. Cell 54, 418–430 (2014).
Acknowledgements
We are grateful to Z. Gong, X. Cao, Y. Fang, A. Millar and B. Bartel for sharing Arabidopsis mutants and transgenic lines. This work was supported by grants from the National Science Foundation of China (grant nos. 31225015, 31330042 and 31421001) to Y.Q.
Author information
Authors and Affiliations
Contributions
X.F. and Y.Q. conceived this project, designed experiments and analysed data. X.F., Y.C. and Y.L. performed the experiments. Y.Q. and X.F. wrote the manuscript. All authors discussed the results and made comments on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Fang, X., Cui, Y., Li, Y. et al. Transcription and processing of primary microRNAs are coupled by Elongator complex in Arabidopsis. Nature Plants 1, 15075 (2015). https://doi.org/10.1038/nplants.2015.75
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/nplants.2015.75
This article is cited by
-
Accumulation of DNA damage alters microRNA gene transcription in Arabidopsis thaliana
BMC Plant Biology (2022)
-
MicroRNA biogenesis and activity in plant cell dedifferentiation stimulated by cell wall removal
BMC Plant Biology (2022)
-
R-loops at microRNA encoding loci promote co-transcriptional processing of pri-miRNAs in plants
Nature Plants (2022)
-
Arabidopsis RBV is a conserved WD40 repeat protein that promotes microRNA biogenesis and ARGONAUTE1 loading
Nature Communications (2022)
-
The R-loop influences miRNA birth place
Nature Plants (2022)