Abstract
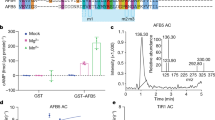
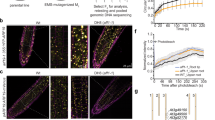
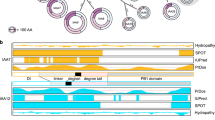
Plant genomes encode large numbers of F-box proteins (FBPs), the substrate recognition subunit of SKP1–CULLIN–F-box (SCF) ubiquitin ligases. There are ∼700 FBPs in Arabidopsis, most of which are uncharacterized. TIR1 is among the best-studied plant FBPs and functions as a receptor for the plant hormone auxin. Here we use a yeast two-hybrid system to identify novel TIR1 mutants with altered properties. The analysis of these mutants reveals that TIR1 associates with the CULLIN1 (CUL1) subunit of the SCF through the N-terminal H1 helix of the F-box domain. Mutations that untether TIR1 from CUL1 stabilize the FBP and cause auxin resistance and associated growth defects, probably by protecting TIR1 substrates from degradation. Based on these results we propose that TIR1 is subject to autocatalytic degradation when assembled into an SCF. Further, our results suggest a general method for determining the physiological function of uncharacterized FBPs. Finally, we show that a key amino acid variation in the F-box domain of auxin signalling F-box (AFB1), a closely related FBP, reduces its ability to form an SCF, resulting in an increase in AFB1 levels.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Deshaies, R. J. & Joazeiro, C. A. RING domain E3 ubiquitin ligases. Annu. Rev. Biochem. 78, 399–434 (2009).
Hua, Z. & Vierstra, R. D. The cullin-RING ubiquitin-protein ligases. Annu. Rev. Plant Biol. 62, 299–334 (2011).
Gagne, J. M., Downes, B. P., Shiu, S. H., Durski, A. M. & Vierstra, R. D. The F-box subunit of the SCF E3 complex is encoded by a diverse superfamily of genes in Arabidopsis. Proc. Natl Acad. Sci. USA 99, 11519–11524 (2002).
Olmedo, G. et al. ETHYLENE-INSENSITIVE5 encodes a 5′-->3′ exoribonuclease required for regulation of the EIN3-targeting F-box proteins EBF1/2. Proc. Natl Acad. Sci. USA 103, 13286–13293 (2006).
Potuschak, T. et al. The exoribonuclease XRN4 is a component of the ethylene response pathway in Arabidopsis. Plant Cell 18, 3047–3057 (2006).
Navarro, L. et al. A plant miRNA contributes to antibacterial resistance by repressing auxin signaling. Science 312, 436–439 (2006).
Si-Ammour, A. et al. miR393 and secondary siRNAs regulate expression of the TIR1/AFB2 auxin receptor clade and auxin-related development of Arabidopsis leaves. Plant Physiol. 157, 683–691 (2011).
Bosu, D. R. & Kipreos, E. T. Cullin-RING ubiquitin ligases: global regulation and activation cycles. Cell Div. 3, 7 (2008).
Galan, J. M. & Peter, M. Ubiquitin-dependent degradation of multiple F-box proteins by an autocatalytic mechanism. Proc. Natl Acad. Sci. USA 96, 9124–9129 (1999).
Luke-Glaser, S. et al. CIF-1, a shared subunit of the COP9/signalosome and eukaryotic initiation factor 3 complexes, regulates MEL-26 levels in the Caenorhabditis elegans embryo. Mol. Cell. Biol. 27, 4526–4540 (2007).
Schmidt, M. W., McQuary, P. R., Wee, S., Hofmann, K. & Wolf, D. A. F-box-directed CRL complex assembly and regulation by the CSN and CAND1. Mol. Cell 35, 586–597 (2009).
Wirbelauer, C. et al. The F-box protein Skp2 is a ubiquitylation target of a Cul1-based core ubiquitin ligase complex: evidence for a role of Cul1 in the suppression of Skp2 expression in quiescent fibroblasts. EMBO J. 19, 5362–5375 (2000).
Zhou, P. & Howley, P. M. Ubiquitination and degradation of the substrate recognition subunits of SCF ubiquitin-protein ligases. Mol. Cell 2, 571–580 (1998).
Wang, R. & Estelle, M. Diversity and specificity: auxin perception and signaling through the TIR1/AFB pathway. Curr. Opin. Plant Biol. 21C, 51–58 (2014).
Tan, X. et al. Mechanism of auxin perception by the TIR1 ubiquitin ligase. Nature 446, 640–645 (2007).
Dharmasiri, N. et al. Plant development is regulated by a family of auxin receptor F box proteins. Dev. Cell 9, 109–119 (2005).
Greenham, K. et al. The AFB4 auxin receptor is a negative regulator of auxin signaling in seedlings. Curr. Biol. 21, 520–525 (2011).
Calderon Villalobos, L. I. et al. A combinatorial TIR1/AFB-Aux/IAA co-receptor system for differential sensing of auxin. Nature Chem. Biol. 8, 477–485 (2012).
Prigge, M. J., Lavy, M., Ashton, N. W. & Estelle, M. Physcomitrella patens auxin-resistant mutants affect conserved elements of an auxin-signaling pathway. Curr. Biol. 20, 1907–1912 (2010).
Yu, H. et al. Mutations in the TIR1 auxin receptor that increase affinity for auxin/indole-3-acetic acid proteins result in auxin hypersensitivity. Plant Physiol. 162, 295–303 (2013).
Havens, K. A. et al. A synthetic approach reveals extensive tunability of auxin signaling. Plant Physiol. 160, 135–142 (2012).
Nishimura, K., Fukagawa, T., Takisawa, H., Kakimoto, T. & Kanemaki, M. An auxin-based degron system for the rapid depletion of proteins in nonplant cells. Nature Methods 6, 917–922 (2009).
Parry, G. et al. Complex regulation of the TIR1/AFB family of auxin receptors. Proc. Natl Acad. Sci. USA 106, 22540–22545 (2009).
Stuttmann, J., Parker, J. E. & Noel, L. D. Novel aspects of COP9 signalosome functions revealed through analysis of hypomorphic csn mutants. Plant Signal. Behav. 4, 896–898 (2009).
Zheng, N. et al. Structure of the Cul1-Rbx1-Skp1-F boxSkp2 SCF ubiquitin ligase complex. Nature 416, 703–709 (2002).
Ruegger, M. et al. The TIR1 protein of Arabidopsis functions in auxin response and is related to human SKP2 and yeast grr1p. Genes Dev. 12, 198–207 (1998).
Lincoln, C., Britton, J. H. & Estelle, M. Growth and development of the axr1 mutants of Arabidopsis. Plant Cell 2, 1071–1108 (1990).
Rouse, D., Mackay, P., Stirnberg, P., Estelle, M. & Leyser, O. Changes in auxin response from mutations in an AUX/IAA gene. Science 279, 1371–1373 (1998).
Wilson, A. K., Pickett, F. B., Turner, J. C. & Estelle, M. A dominant mutation in Arabidopsis confers resistance to auxin, ethylene and abscisic acid. Mol. Gen. Genet. 222, 377–383 (1990).
Rogg, L. E., Lasswell, J. & Bartel, B. A gain-of-function mutation in IAA28 suppresses lateral root development. Plant Cell 13, 465–480 (2001).
Tian, Q. & Reed, J. W. Control of auxin-regulated root development by the Arabidopsis thaliana SHY2/IAA3 gene. Development 126, 711–721 (1999).
Skaar, J. R., Pagan, J. K. & Pagano, M. Mechanisms and function of substrate recruitment by F-box proteins. Nature Rev. Mol. Cell Biol. 14, 369–381 (2013).
Yan, J. et al. The Arabidopsis F-box protein CORONATINE INSENSITIVE1 is stabilized by SCFCOI1 and degraded via the 26S proteasome pathway. Plant Cell 25, 486–498 (2013).
Jurado, S. et al. The Arabidopsis cell cycle F-box protein SKP2A binds to auxin. Plant Cell 22, 3891–3904 (2010).
Bargmann, B. O. & Birnbaum, K. D. Positive fluorescent selection permits precise, rapid, and in-depth overexpression analysis in plant protoplasts. Plant Physiology 149, 1231–1239 (2009).
Karimi, M., Inze, D. & Depicker, A. GATEWAY vectors for Agrobacterium-mediated plant transformation. Trends Plant Sci. 7, 193–195 (2002).
Acknowledgements
We gratefully acknowledge Eric Klavins for use of the cytometer and Shelly Jang for assistance with calculation of k5 values. This work was supported by the National Institutes of Health (grant no. GM43644, M.E.), the Howard Hughes Medical Institute (M.E.), the Gordon and Betty Moore Foundation (M.E.), and the Paul G. Allen Family Foundation (J.N.).
Author information
Authors and Affiliations
Contributions
H.Y., Y.Z., B.M., B.O.R.B., R.W., J.N. and M.E. conceived and designed experiments. H.Y., Y.Z., B.M., B.O.R.B. and R.W. performed the experiments. M.P. did the phylogenetic analysis. H.Y., Y.Z., B.M., M.P. and M.E. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Yu, H., Zhang, Y., Moss, B. et al. Untethering the TIR1 auxin receptor from the SCF complex increases its stability and inhibits auxin response. Nature Plants 1, 14030 (2015). https://doi.org/10.1038/nplants.2014.30
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/nplants.2014.30
This article is cited by
-
SmTIR1 positively regulates the growth and root development of Salvia miltiorrhiza
Plant Cell, Tissue and Organ Culture (PCTOC) (2024)
-
Insights of auxin signaling F-box genes in wheat (Triticum aestivum L.) and their dynamic expression during the leaf rust infection
Protoplasma (2023)
-
CUL3 E3 ligases in plant development and environmental response
Nature Plants (2021)
-
Chemical hijacking of auxin signaling with an engineered auxin–TIR1 pair
Nature Chemical Biology (2018)
-
Variation in auxin sensing guides AUX/IAA transcriptional repressor ubiquitylation and destruction
Nature Communications (2017)