Abstract
The human gut microbiome provides us with functional features that we did not have to evolve ourselves and can be viewed as a structured microbial community that operates like a microbial organ within the human host. A minor but important part of this microbiome is the ability to colonise and thrive within the mucous layer that covers the colon epithelium. These mucosal microbes intimately interact with the intestinal tissue and seem to be important modulators of human health. Embedded in the host-secreted mucous matrix, they form a ‘mucosal biofilm’ with a distinct composition and functionality. In this review, we provide evidence that six specific (micro)environmental factors near the colon mucosa shape and determine mucosal biofilm formation and stability, that is, (1) mucous rigidity, (2) gradients of fluid shear, (3) radial oxygen gradients, (4) secretions of host defense molecules, (5) the presence of a rich but challenging nutrient platform and (6) the presence of niches at the colon epithelial surface. In addition, it appears that microbes actively participate in shaping their mucosal environment. Current insights into the interaction between mucosal microbes and their environment are rather limited, and many questions regarding the contribution of mucosal biofilm functionality and stability to human health remain to be answered. Yet, given the higher potency of mucosal microbes than their luminal counterparts to interact with the host, new insights can accelerate the development of novel disease-preventive or therapeutic strategies.
Similar content being viewed by others
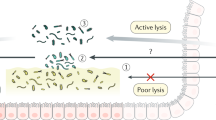
The human colon harbours a highly dense microbial community of 1011–1012 cells per gram of gut content.1,2 Although the majority of this community thrives in the lumen, few microbes can be found within the protective mucous layer that covers the epithelial cells (105–106 cells per ml of mucus).3 These microbes form the colon ‘mucosal biofilm’. Mucosal biofilms were previously defined as microbial biofilms that are unique to the mucosal environment.4 In contrast to biofilms that grow on inert surfaces, they are modulated by host inflammatory responses, and host proteins and cells contribute to the composition of the extracellular matrix.4 Healthy colon mucosal biofilms typically have low microbial densities,5 and growth of complex mushroom-shaped structures that are common for mature biofilms (indicated as stage 4 by Stoodley et al.6) have thus far not been observed. Hence, from a microbial point of view, we infer that the colon mucosal biofilm should be regarded as an immature biofilm.
The mucosal biofilm is unique in composition compared with the microbial populations that colonise the large intestinal lumen and make up faeces.7–9 At the phylum level, colon mucosal biopsies are enriched in Firmicutes (especially members of the Lachnospiraeceae and Ruminococcaceae families) compared with luminal or faecal samples.10–12 At the genus level, abundant and ubiquitous mucosal members are Bacteroides, Faecalibacterium, Roseburia, Blautia and a series of lactic acid bacteria classified as Leuconostoc, Weissella, Lactococcus and Streptococcus.13,14
Mucosal microbes intimately interact with the gut epithelium and gut-associated lymphoid tissue,15–17 and are increasingly evidenced to be important modulators of human health. On one hand, their abundance and prevalence is associated with disease. Low levels of Faecalibacterium prausnitzii have been correlated with active IBD and infectious colitis,18 quiescent ileal Crohn’s disease19 and colorectal adenomas.20 On the other hand, mucosal species (Bacteroides fragilis, Lactobacillus reuteri) have been shown to protect against experimentally induced colitis in animal models.21,22 Interestingly, members of the mucosal microbiome are found to possess distinct functionalities that are not expressed in standard in vitro set-ups. One such example is the extremely oxygen-sensitive F. prausnitzii species, which has been described to use an exogenous flavin-thiol electron shuttle to cope with elevated oxidative stress in the mucosal environment.23 The case of F. prausnitzii is one of many examples showing how physicochemical conditions in the gut mucosa determine microbial colonisation processes.
We review the current knowledge on how the specific physicochemical conditions near the colon mucosa shape the mucosal biofilm composition and functionality of a healthy, adult colon and vice versa. Data of the small intestinal mucosa are not considered, because of its different physiology and interaction with gut microbes.24 On the basis of this analysis, we argue that a better understanding of the specific interactions between mucosal microbes and their environment is essential for exploring the full potential of gut microbes to modulate human health.
The major environmental factor shaping the mucosal biofilm is mucus. Colon mucus—in contrast to the unattached and discontinuous mucous layer in the small intestine—is present as a thick and continuous layer.25,26 It forms a viscoelastic, permeable gel that lubricates and protects the colon epithelial cells against foreign particles and microbial invasion while selectively allowing for the transport of gases, ions, nutrients and proteins.27 These selective barrier properties are determined by its biochemical composition, with the most important constituents being gel-forming mucins. Mucins are densely glycosylated glycoproteins that are cross-linked and assembled via disulphide bonds, and often negatively charged with sialic acids or sulphate groups. In the colon, Muc2 mucins are the only gel-forming mucins.28 Colon Muc2 mucins form two distinct layers: a firm, cell-adherent inner mucous layer, and a loose, unattached outer mucous layer.26,29 The inner mucous layer is a thin, lamellar, highly organised layer of densely stacked Muc2 mucins that remain anchored to the epithelial cells. It has a thickness of several hundred micrometres.30 The outer mucous layer is formed from the inner layer by proteolytic cleavages allowing the Muc2 mucin to expand three to four times into a loose polymeric network removable by suction.31 The rigidity and viscosity of mucus may further fluctuate, as mucin glycosylation patterns, their charge and their degree of cross-linking are highly responsive to the exact concentration of other mucous constituents such as water, lipids, ions, DNA, proteins, cells and cellular debris.27 For example, high concentrations of calcium can entirely collapse the mucus gel by facilitating protein cross-links between mucin monomers,32 while high acidity can decrease the viscosity of mucus by reducing the negative charges on sialic acids.33
Another intrinsic feature of the colon mucous layer is its rapid turnover time. A recent mouse study has revealed that the inner mucous layer is fully renewed within 1 h,34 while the gut epithelium tissue is self-renewed within 4–5 days.35 Hence, it is clear that mucus forms a remarkable and sensitive niche for microbial life, in which microbes are challenged to adapt to the biochemical dynamics determining mucous viscoelasticity and to persist within a continuously renewed environment.
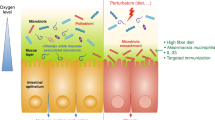
We identify six microenvironmental conditions in the mucosal biofilm that have been evidenced to impact microbial colonisation and functionality (Figure 1). First, the increased rigidity of the inner mucous layer is shown to physically interfere with the migration of microbes. Mucosal microbes mostly colonise the looser, outer mucous layer, whereas the inner, firmer layer is reported to be largely devoid of or contain very low numbers of bacteria.22,31 In rats, the inner mucous layer was furthermore shown to have a distinct bacterial community composition.22 Indeed, differences in viscosity were found to significantly select for microbes with a different morphology.36 Low viscosity selects for short coccoid rods like Bacteroides spp., a moderately viscous environment is preferred by long curved rods like Eubacterium rectale, and a high viscosity immobilizes all bacterial groups.36 Gradients in mucous rigidity may further select for microbes that developed tools to attach to and/or migrate within the mucous layer. These tools are typically extracellular protein polymers like flagella, pili and fimbriae.37 Pili and fimbriae are mostly used for adherence to epithelial cells and mucus, while flagella additionally enable microbes to swim and swarm across the mucus. These structures have extensively been studied in human intestinal pathogens and are important for colonisation of the host tissue and virulence.38 However, they may also be important for mucous colonisation by beneficial microbes, as adherent pili and fimbriae have been identified in L. rhamnosus GG39 and Bifidobacterium breve.40 Moreover, a variety of Lactobacillus spp. was found to possess mucus-binding proteins that enable adhesion to a wide range of mucous ligands.37 Finally, it was suggested that commensal mucus-binding Bacteroides spp. utilise extracellular proteins that bind starch and other carbohydrates on the mucin glycoproteins.41
Interestingly, there is increasing evidence that gut microbes may actively participate in regulating mucous viscosity. A recent study with two colonies of genetically identical mice showed that the presence of a specific gut microbiome may determine the strength of the mucous barrier.42 The first mouse colony had an inner mucous layer that was impenetrable by bacteria or beads of the same size, whereas the inner mucous layer of the second colony was penetrable by bacteria and beads. The mucous phenotypes depended on the gut microbiome because they were transmissible by transfer of caecal microbiota to germ-free mice. At the biochemical level, evidence for a microbial impact on mucous viscosity and penetrability is provided by study of the human commensal F. prausnitzii. F. prausnitzii growth in the oxygenated mucosal biofilm was found to require free pools of thiols, such as cysteine and glutathione.23 These are redox mediators and are important for maintaining mucous fluidity while they can modulate reduction of the disulphide bonds that assemble the mucin glycoproteins into polymeric networks.43 Hence, F. prausnitzii growth and mucous rigidity rely on the same biochemical products, and might be closely connected.
Second, differences in rigidity between inner and outer mucous layer expose the mucosal microbes to gradients of fluid shear. Microbes that reside between the microvilli of the epithelial cells are exposed to a low fluid shear environment44 and experience much less physical perturbation when compared with the microbes in the outer, more turbulent mucous layer. Low fluid shear was shown to completely reprogramme the gene expression, physiology and stress resistance of several (opportunistic) pathogens, including Salmonella enterica serovar Typhimurium (for which virulence was also altered), Escherichia coli, Pseudomonas aeruginosa and Staphylococcus aureus.44–46 The latter two organisms also showed alterations in biofilm formation.46,47 In the case of S. aureus, low fluid shear conditions that altered biofilm architecture made the bacterium more resistant to antibiotic stress, lowered its resistance to oxidative stress and decreased carotenoid production.46 Although the impact of low fluid shear on commensal gut microbes has not been described so far, it seems plausible that mucous fluidity and shear are important determinants of mucosal biofilm structure and functionality.
Third, microbial life in the mucosal biofilm experiences gradients in oxygen, which leaks from the epithelial cells and dilutes into the mucous layer. Albenberg et al. recently measured a steep and radial oxygen gradient in the mice caecum going from 40 mm Hg in the intestinal tissue to deeply anaerobic conditions (<1 mm Hg) in the bulk of the lumen.48 For the first time, these authors demonstrated the potential of oxygen to shape the (mucosal) gut microbiome. They increased the oxygenation of mouse intestinal tissues by hyperbaric oxygen therapy, and found shifts in their faecal microbial composition. Moreover, in humans, the authors found that rectal mucosal biopsies and swabs contained a higher relative abundance of oxygen-tolerant Proteobacteria and Actinobacteria when compared with paired stool samples.
Additional evidence for oxygen gradients to shape mucosal biofilm formation is provided by the fact that prominent strictly anaerobic mucosal microbes seem to benefit from low oxygen concentrations. As mentioned previously in this text, F. prausnitzii species were found to tolerate low concentrations of oxygen by using an extracellular electron shuttle of flavins and thiols.23 This way of electron shuttling to oxygen results in a competitive growth advantage and may explain why F. prausnitzii is found in vivo and in vitro in the outer mucous layer and lumen/mucus interphase.19,49,50 Another mechanism by which gut microbes may benefit from low oxygen levels near the gut mucosa was found in Bacteroides fragilis. This species encodes cytochrome bd oxidase, an essential enzyme for oxygen consumption that was shown to stimulate its growth in the presence of nanomolar concentrations of oxygen.51 This feature explains why strictly anaerobic Bacteroides spp. are able to colonise the mouse intestine without pre-colonisation by facultative anaerobes that reduce oxygen creating a strictly anaerobic environment.52 Cytochrome oxidases seem to be widespread in the genome of (gut) bacterial species belonging to different phyla, among which are found many human intestinal pathogens.51,53 Hence, it is likely that a variety of microbes and pathogens are benefiting from low oxygen levels near the colon epithelial surface.53
Finally, increased oxygen availability upon migration to the colonic mucosal surface was found to enhance the adhesive and/or invasive properties of human enteric pathogens that infect colon epithelial cells, including Salmonella enterica,54,55 Shigella flexneri56 and enterohaemorrhagic Escherichia coli.57 Radial oxygen gradients in the colon mucous layer may thus also have a role in disease and the disruption of the normal microbiota.
A fourth factor shaping colon mucosal colonisation patterns is the secretion of host defense molecules such as antimicrobial peptides (AMPs) and secretory IgA, by the colon epithelial cells. In comparison with the small intestinal epithelium, colon epithelial cells are secreting much lower amounts of AMPs and sIgA than small intestinal epithelium.25 This is due to their different physiology: AMP-secreting Paneth cells are abundant in the small intestine but rare in the colon, and sIgA-regulation by aggregated lymphoid nodules (Peyer’s patches) in the small intestine is taken over by smaller, isolated lymphoid follicles in the colon.25 Yet, the antimicrobial activity in the colon mucosal biofilm should not be underestimated. It has been shown that secretions of AMPs remain trapped in the mucus and only marginally leach to the lumen.58 In addition, AMPs can also be produced by mouse Goblet cells, which are mucus-secreting enterocytes that are highly abundant in the colon.59 Although the presence of these antimicrobial components in the mucous layer impacts the colonisation ability of several microorganisms, mucosal microbes themselves are capable to regulate their secretion and thereby modify the mucosal microbiome. Bacteroides thetaiotaomicron has been described to shape innate immunity by modulating antimicrobial angiogenin4 production by mouse Paneth cells.60 Another example is the inducation of RegIIIγ production in the mouse colon, not ileum, by the mucinolytic Akkermansia muciniphila. As RegIIIγ exerts direct antimicrobial activity against gram-positive microbes in the intestine,61 A. muciniphila is thus able to favour its own colonisation and growth. Hence, we infer that colon mucosal biofilm formation is shaped by the build-up of host defense molecules in the thick mucous layer, with antimicrobial activities comparable to (or even greater than) those in the small intestine.
A fifth factor that determines whether microbes are able to persist within the mucous layer is the presence of a rich but challenging nutrient platform. Mucosal microbes may feed or cross-feed on ‘easy’ substrates that are leaching from the lumen in the (outer) mucous layer or that are produced in the biofilm. However, it appears that the colon mucosal nutrient platform is dominated by less degradable host-derived secretions. Indeed, the composition of the mucosal biofilm was evidenced to be enriched in asaccharolytic microbes that primarily metabolise peptones and amino acids, which are probably derived from the proteinaceous substrates from mucus and shedded epithelium.48 In the mouse caecum, stable isotope probing with threonine revealed that two bacterial species, A. muciniphila and Bacteroides acidifaciens, are key host-protein foragers in vivo.62 A. muciniphila and Bacteroides spp. are dominant members of the human colon microbiome8,63 that have been shown to degrade mucin glycoproteins.64,65 Bacteroides spp. and A. muciniphila have evolved rather different strategies to forage on mucin. Although Bacteroides spp. have broad polysaccharide degradation potential66 and only turn to mucin glycans when dietary glycans are depleted,67 A. muciniphila has specialized to utilise mucins as a sole carbon and nitrogen source.63,64 For B. thetaiotaomicron, foraging on mucin glycans was found to be important for successful colonisation of the mouse gut, because deletion of the involved gene cassettes was shown to compromise its transmission from gnotobiotic mother to offspring.68 Hence, its capacity to thrive on mucin glycans and live in close proximity to the intestinal epithelial surface seems to be an evolutionary trait allowing for its stable and continuous colonisation of the gut. For A. muciniphila, foraging on mucin seems not to confine the species to the mucosal biofilm niche. In our dynamic in vitro model of the proximal and distal colon microbiome (SHIME),69,70 A. muciniphila was found to abundantly colonise the distal colon compartment in the absence of a mucosal biofilm. Moreover, supplementation of mucin to this model resulted in a strong and quick increase in luminal A. muciniphila qPCR counts (4.5 log increase in 2 days) (unpublished data). In addition, A. muciniphila seems not to bind to human colon mucus.71 Considering A. muciniphila’s unique specialisation to grow on mucin, we infer that its in vivo niche is associated with mucus regardless of their embedment within the colon mucosal biofilm or not.
Although some mucosal species directly benefit from host-secreted mucin, their activity was found to depend on the presence of other community members.62,72 Indeed, complete degradation of the heterogeneous and complex structure of mucin glycoproteins requires a diverse set of enzymes,73 and the in vitro mucin degradation potential of microbes is phylogenetically widespread.74,75 On the other hand, specialist mucin degraders may stimulate the growth of other mucosal microbes. For example, Bacteroides spp. are known to release sulphate from mucin glycans, which makes them more accessible for further degradation by other microbes76 and may stimulate the growth of sulphate-reducing bacteria.77
Finally, microbes may enhance their persistence in the colon mucosal biofilm by specializing to colonise niches at the colon epithelial surface. An example of such a niche is the intestinal crypt. Crypts are vulnerable sites of the gut epithelium as they contain stem cells and are the site of epithelial proliferation and restitution. In healthy rodents, crypt-associated microbes have been detected in the caecum and proximal colon, but not in the distal colon and rectum.36,78,79 Likewise, healthy human rectal crypts were found to be devoid of bacteria.80,81 Crypt-associated microbes in the human proximal colon have not been studied so far owing to the difficulties with obtaining correct and unbiased samples. The crypt micro-environment is proposed to be characterized by a low mucous viscosity,36 a high partial oxygen pressure that selects for aerobic species in mice79 and the presence of specific types and concentrations of host glycans mediating saturable crypt occupancy by Bacteroides spp.17 Interestingly, glycan-mediated crypt occupancy by Bacteroides was found to be responsible for its stable colonisation of the gut, even in the event of enteric infection or following antibiotic treatment.17 The authors therefore proposed that crypt-associated microbiota might represent bacterial reservoirs of founder cells that repopulate the gut following disruption by infections or antibiotic consumption.17
Altogether, it has become evident that the colon mucosa is a unique and highly structured gut environment. The combination of radial gradients in rigidity, fluid shear and oxygen concentrations, and the presence of antimicrobial host secretions and a distinct nutrient pool creates a variety of microbial microenvironments that result in a specific biofilm stratification or architecture that impact mucosal biofilm composition and functionality. On the other hand, microbes seem to actively participate in shaping the mucosal environment, for example, by degrading mucus and affecting mucous viscosity or triggering the production of AMPs.
Although the composition of the mucosal biofilm seems to correlate with health status, insight into the intrinsic relation between mucosal microbes and their environment could hold exciting perspectives for monitoring and modulating human health. Particular interest thereby goes to the interrelated and simultaneous impact of the different microenvironmental factors on mucosal biofilm functioning, in both health and disease. Overall, such research efforts should help in identifying (i) what mucosal species and/or microbial functionalities are crucial for maintaining a healthy mucosal barrier, (ii) how the immune system copes with these mucosal microbes and what role mucolytic activity from specific gut microbes has in modulating human health.
References
Stephen, A. M. & Cummings, J. H. Microbial contribution to human fecal mass. J. Med. Microbiol. 13, 45–56 (1980).
Whitman, W. B., Coleman, D. C. & Wiebe, W. J. Prokaryotes: the unseen majority. Proc. Natl Acad. Sci. USA 95, 6578–6583 (1998).
Van den Abbeele, P., Van de Wiele, T., Verstraete, W. & Possemiers, S. The host selects mucosal and luminal associations of coevolved gut microorganisms: a novel concept. FEMS Microbiol. Rev. 35, 681–704 (2011).
Post, J. C., Stoodley, P., Hall-Stoodley, L. & Ehrlich, G. The role of biofilms in otolaryngologic infections. Curr. Opin. Otolaryngol. Head Neck Surg. 12, 185–190 (2004).
Swidsinski, A., Weber, J., Loening-Baucke, V., Hale, L. P. & Lochs, H. Spatial organization and composition of the mucosal flora in patients with inflammatory bowel disease. J. Clin. Microbiol. 43, 3380–3389 (2005).
Stoodley, P., Sauer, K., Davies, D. G. & Costerton, J. W. Biofilms as complex differentiated communities. Annu. Rev. Microbiol. 56, 187–209 (2002).
Zoetendal, E. G. et al. Mucosa-associated bacteria in the human gastrointestinal tract are uniformly distributed along the colon and differ from the community recovered from feces. Appl. Environ. Microbiol. 68, 3401–3407 (2002).
Eckburg, P. B. et al. Diversity of the human intestinal microbial flora. Science 308, 1635–1638 (2005).
Lepage, P. et al. Biodiversity of the mucosa-associated microbiota is stable along the distal digestive tract in healthy individuals and patients with IBD. Inflamm. Bowel Dis. 11, 473–480 (2005).
Wang, Y. et al. Laser capture microdissection and metagenomic analysis of intact mucosa-associated microbial communities of human colon. Appl. Microbiol. Biotechnol. 88, 1333–1342 (2010).
Willing, B. P. et al. A pyrosequencing study in twins shows that gastrointestinal microbial profiles vary with inflammatory bowel disease phenotypes. Gastroenterology 139, 1844–U1105 (2010).
Nava, G. M., Friedrichsen, H. J. & Stappenbeck, T. S. Spatial organization of intestinal microbiota in the mouse ascending colon. ISME J. 5, 627–638 (2011).
Hong, P.-Y., Croix, J. A., Greenberg, E., Gaskins, H. R. & Mackie, R. I. Pyrosequencing-based analysis of the mucosal microbiota in healthy individuals reveals ubiquitous bacterial groups and micro-heterogeneity. PLoS ONE 6, e25042 (2011).
Nava, G. M. & Stappenbeck, T. S. Diversity of the autochthonous colonic microbiota. Gut Microbes 2, 99–104 (2011).
Lebeer, S. et al. Functional analysis of Lactobacillus rhamnosus GG pili in relation to adhesion and immunomodulatory interactions with intestinal epithelial cells. Appl. Environ. Microbiol. 78, 185–193 (2012).
Atarashi, K. et al. Induction of colonic regulatory T cells by indigenous clostridium species. Science 331, 337–341 (2011).
Lee, S. M. et al. Bacterial colonization factors control specificity and stability of the gut microbiota. Nature 501, 426–429 (2013).
Sokol, H. et al. Low counts of Faecalibacterium prausnitzii in colitis microbiota. Inflamm. Bowel Dis. 15, 1183–1189 (2009).
Willing, B. et al. Twin studies reveal specific imbalances in the mucosa-associated microbiota of patients with ileal Crohn's disease. Inflamm. Bowel Dis. 15, 653–660 (2009).
Shen, X. J. et al. Molecular characterization of mucosal adherent bacteria and associations with colorectal adenomas. Gut Microbes 1, 138–147 (2010).
Mazmanian, S. K., Round, J. L. & Kasper, D. L. A microbial symbiosis factor prevents intestinal inflammatory disease. Nature 453, 620–625 (2008).
Dicksved, J. et al. Lactobacillus reuteri maintains a functional mucosal barrier during DSS treatment despite mucus layer dysfunction. PLoS ONE 7, e46399 (2012).
Khan, M. T. et al. The gut anaerobe Faecalibacterium prausnitzii uses an extracellular electron shuttle to grow at oxic-anoxic interphases. ISME J. 6, 1578–1585 (2012).
Wells, J. M., Rossi, O., Meijerink, M. & van Baarlen, P. Epithelial crosstalk at the microbiota-mucosal interface. Proc. Natl Acad Sci USA 108, 4607–4614 (2011).
Brown, E. M., Sadarangani, M. & Finlay, B. B. The role of the immune system in governing host-microbe interactions in the intestine. Nat. Immunol. 14, 660–667 (2013).
Johansson, M. E. V., Sjovall, H. & Hansson, G. C. The gastrointestinal mucus system in health and disease. Nat. Rev. Gastroenterol. Hepatol. 10, 352–361 (2013).
Lai, S. K., Wang, Y. -Y., Wirtz, D. & Hanes, J. Micro- and macrorheology of mucus. Adv. Drug Deliv. Rev. 61, 86–100 (2009).
Kim, Y. S. & Ho, S. B. Intestinal goblet cells and mucins in health and disease:recent insights and progress. Curr. Gastroenterol. Rep. 12, 319–330 (2010).
Johansson, M. E. V., Larsson, J. M. H. & Hansson, G. C. The two mucus layers of colon are organized by the MUC2 mucin, whereas the outer layer is a legislator of host-microbial interactions. Proc. Natl Acad. Sci. USA 108, 4659–4665 (2011).
Swidsinski, A. et al. Comparative study of the intestinal mucus barrier in normal and inflamed colon. Gut 56, 343–350 (2007).
Johansson, M. E. V. et al. The inner of the two Muc2 mucin-dependent mucus layers in colon is devoid of bacteria. Proc. Natl Acad. Sci. USA 105, 15064–15069 (2008).
Raynal, B. D. E., Hardingham, T. E., Sheehan, J. K. & Thornton, D. J. Calcium-dependent protein interactions in MUC5B provide reversible cross-links in salivary mucus. J. Biol. Chem. 278, 28703–28710 (2003).
Canani, R. B. & Terrin, G. Gastric acidity inhibitors and the risk of intestinal infections. Curr. Opin. Gastroenterol. 26, 31–35 (2010).
Johansson, M. E. V. Fast renewal of the distal colonic mucus layers by the surface goblet cells as measured by in vivo labeling of mucin glycoproteins. PLoS ONE 7, e41009 (2012).
van der Flier, L. G. & Clevers, H. Stem cells, self-renewal, and differentiation in the intestinal epithelium. Ann. Rev. Physiol. 71, 241–260 (2009).
Swidsinski, A. et al. Viscosity gradient within the mucus layer determines the mucosal barrier function and the spatial organization of the intestinal microbiota. Inflamm. Bowel Dis. 13, 963–970 (2007).
Juge, N. Microbial adhesins to gastrointestinal mucus. Trends Microbiol. 20, 30–39 (2012).
Jonson, A. B., Normark, S. & Rhen, M. Fimbriae, pili, flagella and bacterial virulence. Contrib. Microbiol. 12, 67–89 (2005).
von Ossowski, I. et al. Mucosal adhesion properties of the probiotic Lactobacillus rhamnosus GG SpaCBA and SpaFED pilin subunits. Appl. Environ. Microbiol. 76, 2049–2057 (2010).
Motherway, M. O. C. et al. Functional genome analysis of Bifidobacterium breve UCC2003 reveals type IVb tight adherence (Tad) pili as an essential and conserved host-colonization factor. Proc. Natl Acad. Sci. USA 108, 11217–11222 (2011).
Huang, J. Y., Lee, S. M. & Mazmanian, S. K. The human commensal Bacteroides fragilis binds intestinal mucin. Anaerobe 17, 137–141 (2011).
Jakobsson, H. E. et al. The composition of the gut microbiota shapes the colon mucus barrier. EMBO Rep. 16, 164–177 (2015).
Circu, M. L. & Aw, T. Y. Redox biology of the intestine. Free Radic. Res. 45, 1245–1266 (2011).
Nickerson, C. A., Ott, C. M., Wilson, J. W., Ramamurthy, R. & Pierson, D. L. Microbial responses to microgravity and other low-shear environments. Microbiol. Mol. Biol. Rev. 68, 345–361 (2004).
Crabbe, A. et al. Response of Pseudomonas aeruginosa PAO1 to low shear modelled microgravity involves AlgU regulation. Environ. Microbiol. 12, 1545–1564 (2010).
Castro, S. L., Nelman-Gonzalez, M., Nickerson, C. A. & Ott, C. M. Induction of attachment-independent biofilm formation and repression of hfq expression by low-fluid-shear culture of Staphylococcus aureus . Appl. Environ. Microbiol. 77, 6368–6378 (2011).
Crabbe, A. et al. Use of the rotating wall vessel technology to study the effect of shear stress on growth behaviour of Pseudomonas aeruginosa PA01. Environ. Microbiol. 10, 2098–2110 (2008).
Albenberg L. et al. Correlation between intraluminal oxygen gradient and radial partitioning of intestinal microbiota. Gastroenterology 2014; 147: 1055–1063.
Swidsinski, A., Loening-Baucke, V., Verstraelen, H., Osowska, S. & Doerffel, Y. Biostructure of fecal microbiota in healthy subjects and patients with chronic idiopathic diarrhea. Gastroenterology 135, 568–579 (2008).
Marzorati, M. et al. The HMI (TM) module: a new tool to study the host-microbiota interaction in the human gastrointestinal tract in vitro . BMC Microbiol. 14, 133 (2014).
Baughn, A. D. & Malamy, M. H. The strict anaerobe Bacteroides fragilis grows in and benefits from nanomolar concentrations of oxygen. Nature 427, 441–444 (2004).
Mazmanian, S. K., Liu, C. H., Tzianabos, A. O. & Kasper, D. L. An immunomodulatory molecule of symbiotic bacteria directs maturation of the host immune system. Cell 122, 107–118 (2005).
Morris, R. L. & Schmidt, T. M. Shallow breathing: bacterial life at low O-2. Nat. Rev. Microbiol. 11, 205–212 (2013).
Lee, C. A. & Falkow, S. The ability of salmonella to enter mammalian-cells is affected by bacterial-growth state. Proc. Natl Acad. Sci. USA 87, 4304–4308 (1990).
Jones, B. D. & Falkow, S. Identification and characterization of a salmonella-typhimurium oxygen-regulated gene required for bacterial internalization. Infect. Immun. 62, 3745–3752 (1994).
Marteyn, B. et al. Modulation of Shigella virulence in response to available oxygen in vivo . Nature 465, 355–358 (2010).
Schuller, S. & Phillips, A. D. Microaerobic conditions enhance type III secretion and adherence of enterohaemorrhagic Escherichia coli to polarized human intestinal epithelial cells. Environ. Microbiol. 12, 2426–2435 (2010).
Meyer-Hoffert, U. et al. Secreted enteric antimicrobial activity localises to the mucus surface layer. Gut 57, 764–771 (2008).
Forman, R. A. et al. The goblet cell is the cellular source of the anti-microbial angiogenin 4 in the large intestine post Trichuris muris infection. PLoS ONE 7, e42248 (2012).
Hooper, L. V., Stappenbeck, T. S., Hong, C. V. & Gordon, J. I. Angiogenins: a new class of microbicidal proteins involved in innate immunity. Nat. Immunol. 4, 269–273 (2003).
Everard, A. et al. Cross-talk between Akkermansia muciniphila and intestinal epithelium controls diet-induced obesity. Proc. Natl Acad. Sci. USA 110, 9066–9071 (2013).
Berry, D. et al. Host-compound foraging by intestinal microbiota revealed by single-cell stable isotope probing. Proc. Natl Acad. Sci. USA 110, 4720–4725 (2013).
Derrien, M., Collado, M. C., Ben-Amor, K., Salminen, S. & de Vos, W. M. The mucin degrader Akkermansia muciniphila is an abundant resident of the human intestinal tract. Appl. Environ. Microbiol. 74, 1646–1648 (2008).
Derrien, M., Vaughan, E. E., Plugge, C. M. & de Vos, W. M. Akkermansia muciniphila gen. nov., sp nov., a human intestinal mucin-degrading bacterium. Int. J. Syst. Evol. Microbiol. 54, 1469–1476 (2004).
Sonnenburg, J. L. et al. Glycan foraging in vivo by an intestine-adapted bacterial symbiont. Science 307, 1955–1959 (2005).
Xu, J. et al. Evolution of symbiotic bacteria in the distal human intestine. PLoS Biol. 5, 1574–1586 (2007).
Lynch, J. B. & Sonnenburg, J. L. Prioritization of a plant polysaccharide over a mucus carbohydrate is enforced by a Bacteroides hybrid two-component system. Mol. Microbiol. 85, 478–491 (2012).
Martens, E. C., Chiang, H. C. & Gordon, J. I. Mucosal glycan foraging enhances fitness and transmission of a saccharolytic human gut bacterial symbiont. Cell Host Microbe 4, 447–457 (2008).
Molly, K., Vandewoestyne, M., Desmet, I. & Verstraete, W. Validation of the simulator of the human intestinal microbial ecosystem (shime) reactor using microorganism-associated activities. Microb. Ecol. Health Dis. 7, 191–200 (1994).
Van den Abbeele, P. et al. Microbial community development in a dynamic gut model is reproducible, colon region specific, and selective for Bacteroidetes and Clostridium cluster IX. Appl. Environ. Microbiol. 76, 5237–5246 (2010).
Reunanen, J. et al. Akkermansia muciniphila adheres to enterocytes and strengthens the integrity of the epithelial cell layer. Appl. Environ. Microbiol. 81, 3655–3662 (2015).
Png, C. W. et al. Mucolytic bacteria with increased prevalence in IBD mucosa augment in vitro utilization of mucin by other bacteria. Am. J. Gastroenterol. 105, 2420–2428 (2010).
Tailford, L. E., Crost, E. H., Kavanaugh, D. & Juge, N. Mucin glycan foraging in the human gut microbiome. Front. Genet. 6, 81 (2015).
McGuckin, M. A., Linden, S. K., Sutton, P. & Florin, T. H. Mucin dynamics and enteric pathogens. Nat. Rev. Microbiol. 9, 265–278 (2011).
Derrien, M. et al. Mucin-bacterial interactions in the human oral cavity and digestive tract. Gut Microbes 1, 254–268 (2010).
Benjdia, A., Martens, E. C., Gordon, J. I. & Berteau, O. Sulfatases and a radical S-adenosyl-L-methionine (AdoMet) enzyme are key for mucosal foraging and fitness of the prominent human gut symbiont, Bacteroides thetaiotaomicron . J. Biol. Chem. 286, 25973–25982 (2011).
Willis, C. L., Cummings, J. H., Neale, G. & Gibson, G. R. In vitro effects of mucin fermentation on the growth of human colonic sulphate-reducing bacteria. Anaerobe 2, 117–122 (1996).
Yamamoto, K. et al. Histoplanimetrical study on the spatial relationship of distribution of indigenous bacteria with mucosal lymphatic follicles in alimentary tract of rat. J. Vet. Med. Sci. 71, 621–630 (2009).
Pedron, T. et al. A crypt-specific core microbiota resides in the mouse colon. MBio 3 (2012).
Sokol, H. et al. Crypt abscess-associated microbiota in inflammatory bowel disease and acute self-limited colitis. World J. Gastroenterol. 16, 583–587 (2010).
Macfarlane, S., Furrie, E., Cummings, J. H. & Macfarlane, G. T. Chemotaxonomic analysis of bacterial populations colonizing the rectal mucosa in patients with ulcerative colitis. Clin. Infect. Dis. 38, 1690–1699 (2004).
Acknowledgements
We gratefully acknowledge Tim Lacoere for making the graphical summary (Figure 1).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
De Weirdt, R., Van de Wiele, T. Micromanagement in the gut: microenvironmental factors govern colon mucosal biofilm structure and functionality. npj Biofilms Microbiomes 1, 15026 (2015). https://doi.org/10.1038/npjbiofilms.2015.26
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/npjbiofilms.2015.26
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.