Abstract
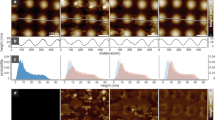
High-speed atomic force microscopy (HS-AFM) can be used to visualize function-related conformational changes of single soluble proteins. Similar studies of single membrane proteins are, however, hampered by a lack of suitable flat, non-interacting membrane supports and by high protein mobility. Here we show that streptavidin crystals grown on mica-supported lipid bilayers can be used as porous supports for membranes containing biotinylated lipids. Using SecYEG (protein translocation channel) and GlpF (aquaglyceroporin), we demonstrate that the platform can be used to tune the lateral mobility of transmembrane proteins to any value within the dynamic range accessible to HS-AFM imaging through glutaraldehyde-cross-linking of the streptavidin. This allows HS-AFM to study the conformation or docking of spatially confined proteins, which we illustrate by imaging GlpF at sub-molecular resolution and by observing the motor protein SecA binding to SecYEG.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Cho, N.-J., Frank, C. W., Kasemo, B. & Höök, F. Quartz crystal microbalance with dissipation monitoring of supported lipid bilayers on various substrates. Nat. Protoc. 5, 1096–1106 (2010).
Patching, S. G. Surface plasmon resonance spectroscopy for characterisation of membrane protein–ligand interactions and its potential for drug discovery. Biochim. Biophys. Acta 1838, 43–55 (2014).
Ando, T. et al. A high-speed atomic force microscope for studying biological macromolecules. Proc. Natl Acad. Sci. USA 98, 12468–12472 (2001).
Giocondi, M.-C., Seantier, B., Dosset, P., Milhiet, P.-E. & Le Grimellec, C. Characterizing the interactions between GPI-anchored alkaline phosphatases and membrane domains by AFM. Pflugers Arch. 456, 179–188 (2008).
Yilmaz, N. et al. Real-time visualization of assembling of a sphingomyelin-specific toxin on planar lipid membranes. Biophys. J. 105, 1397–1405 (2013).
Preiner, J. et al. IgGs are made for walking on bacterial and viral surfaces. Nat. Commun. 5, 4593 (2014).
Casuso, I. et al. Characterization of the motion of membrane proteins using high-speed atomic force microscopy. Nat. Nanotech. 7, 525–529 (2012).
Casuso, I., Sens, P., Rico, F. & Scheuring, S. Experimental evidence for membrane-mediated protein–protein interaction. Biophys. J. 99, L47–L49 (2010).
Johnson, C. P. et al. Structural studies of the neural-cell-adhesion molecule by X-ray and neutron reflectivity. Biochemistry 44, 546–554 (2005).
Miller, C. E., Majewski, J., Gog, T. & Kuhl, T. L. Characterization of biological thin films at the solid–liquid interface by X-ray reflectivity. Phys. Rev. Lett. 94, 238104 (2005).
Nováková, E., Giewekemeyer, K. & Salditt, T. Structure of two-component lipid membranes on solid support: An x-ray reflectivity study. Phys. Rev. E 74, 051911 (2006).
Przybylo, M. et al. Lipid diffusion in giant unilamellar vesicles is more than 2 times faster than in supported phospholipid bilayers under identical conditions. Langmuir 22, 9096–9099 (2006).
Preiner, J. et al. High-speed AFM images of thermal motion provide stiffness map of interfacial membrane protein moieties. Nano Lett. 15, 759–763 (2014).
Horner, A. et al. The mobility of single-file water molecules is governed by the number of H-bonds they may form with channel-lining residues. Sci. Adv. 1, e1400083 (2015).
Yamashita, H. et al. Dynamics of bacteriorhodopsin 2D crystal observed by high-speed atomic force microscopy. J. Struct. Biol. 167, 153–158 (2009).
Yamamoto, D. et al. Chapter twenty-High-Speed atomic force microscopy techniques for observing dynamic biomolecular processes. Methods Enzymol. 475, 541–564 (2010).
Chan, Y.-H. M. & Boxer, S. G. Model membrane systems and their applications. Curr. Opin. Chem. Biol. 11, 581–587 (2007).
Castellana, E. T. & Cremer, P. S. Solid supported lipid bilayers: From biophysical studies to sensor design. Surf. Sci. Rep. 61, 429–444 (2006).
Park, E. et al. Structure of the SecY channel during initiation of protein translocation. Nature 506, 102–106 (2014).
Antonenko, Y. N., Horner, A. & Pohl, P. Electrostatically induced recruitment of membrane peptides into clusters requires ligand binding at both interfaces. PloS ONE 7, e52839 (2012).
Tanaka, M. & Sackmann, E. Polymer-supported membranes as models of the cell surface. Nature 437, 656–663 (2005).
Gonçalves, R. P. et al. Two-chamber AFM: probing membrane proteins separating two aqueous compartments. Nat. Methods 3, 1007–1012 (2006).
Frauenfeld, J. et al. Cryo-EM structure of the ribosome–SecYE complex in the membrane environment. Nat. Struct. Mol. Biol. 18, 614–621 (2011).
Fu, D. et al. Structure of a glycerol-conducting channel and the basis for its selectivity. Science 290, 481–486 (2000).
Saparov, S. M., Tsunoda, S. P. & Pohl, P. Proton exclusion by an aquaglyceroprotein: a voltage clamp study. Biol. Cell 97, 545–550 (2005).
Reviakine, I. & Brisson, A. Streptavidin 2D crystals on supported phospholipid bilayers: toward constructing anchored phospholipid bilayers. Langmuir 17, 8293–8299 (2001).
Yamamoto, D., Nagura, N., Omote, S., Taniguchi, M. & Ando, T. Streptavidin 2D crystal substrates for visualizing biomolecular processes by atomic force microscopy. Biophys. J. 97, 2358–2367 (2009).
Calvert, T. L. & Leckband, D. Two-dimensional protein crystallization at solid-liquid interfaces. Langmuir 13, 6737–6745 (1997).
Schütz, G. J., Schindler, H. & Schmidt, T. Single-molecule microscopy on model membranes reveals anomalous diffusion. Biophys. J. 73, 1073–1080 (1997).
Le Trong, I. et al. Streptavidin and its biotin complex at atomic resolution. Acta Crystallogr. D 67, 813–821 (2011).
Knyazev, D. G. et al. The bacterial translocon SecYEG opens upon ribosome binding. J. Biol. Chem. 288, 17941–17946 (2013).
Breyton, C., Haase, W., Rapoport, T. A., Kühlbrandt, W. & Collinson, I. Three-dimensional structure of the bacterial protein-translocation complex SecYEG. Nature 418, 662–665 (2002).
Zimmer, J., Nam, Y. & Rapoport, T. A. Structure of a complex of the ATPase SecA and the protein-translocation channel. Nature 455, 936–943 (2008).
de Keyzer, J., van der Does, C., Kloosterman, T. G. & Driessen, A. J. M. Direct demonstration of ATP-dependent release of SecA from a translocating preprotein by surface plasmon resonance. J. Biol. Chem. 278, 29581–29586 (2003).
Lill, R., Dowhan, W. & Wickner, W. The ATPase activity of SecA is regulated by acidic phospholipids, SecY, and the leader and mature domains of precursor proteins. Cell 60, 271–280 (1990).
Wallin, E. & Heijne, G. V. Genome-wide analysis of integral membrane proteins from eubacterial, archaean, and eukaryotic organisms. Protein Sci. 7, 1029–1038 (1998).
Kodera, N., Yamamoto, D., Ishikawa, R. & Ando, T. Video imaging of walking myosin V by high-speed atomic force microscopy. Nature 468, 72–76 (2010).
Uchihashi, T., Iino, R., Ando, T. & Noji, H. High-speed atomic force microscopy reveals rotary catalysis of rotorless F1-ATPase. Science 333, 755–758 (2011).
Schwenen, L. L. et al. Resolving single membrane fusion events on planar pore-spanning membranes. Sci. Rep. 5, 12006 (2015).
Knyazev, D. G., Winter, L., Bauer, B. W., Siligan, C. & Pohl, P. Ion conductivity of the bacterial translocation channel SecYEG engaged in translocation. J. Biol. Chem. 289, 24611–24616 (2014).
Sbalzarini, I. F. & Koumoutsakos, P. Feature point tracking and trajectory analysis for video imaging in cell biology. J. Struct. Biol. 151, 182–195 (2005).
Tseng, Q. et al. Spatial organization of the extracellular matrix regulates cell–cell junction positioning. Proc. Natl Acad. Sci. USA 109, 1506–1511 (2012).
Bowman, A. W. & Azzalini, A. Applied Smoothing Techniques for Data Analysis (Clarendon, 2004).
Rankl, C. et al. Multiple receptors involved in human rhinovirus attachment to live cells. Proc. Natl Acad. Sci. USA 105, 17778–17783 (2008).
Axelrod, D., Koppel, D. E., Schlessinger, J., Elson, E. & Webb, W. W. Mobility measurement by analysis of fluorescence photobleaching recovery kinetics. Biophys. J. 16, 1055–1069 (1976).
Klotzsch, E. et al. Superresolution microscopy reveals spatial separation of UCP4 and F0F1-ATP synthase in neuronal mitochondria. Proc. Natl Acad. Sci. USA 112, 130–135 (2015).
Smith, C. S., Joseph, N., Rieger, B. & Lidke, K. A. Fast, single-molecule localization that achieves theoretically minimum uncertainty. Nat. Methods 7, 373–375 (2010).
Gao, Y. & Kilfoil, M. L. Accurate detection and complete tracking of large populations of features in three dimensions. Opt. Express 17, 4685–4704 (2009).
Wieser, S. & Schütz, G. J. Tracking single molecules in the live cell plasma membrane—do's and don't’s. Methods 46, 131–140 (2008).
Harpaz, Y., Gerstein, M. & Chothia, C. Volume changes on protein folding. Structure 2, 641–649 (1994).
Acknowledgements
This work was supported by the Austrian Science Fund (FWF, P25844 to J.P.), the European Fund for Regional Development (EFRE, Regio 13) and the Federal State of Upper Austria. The authors thank H. Gruber for helpful discussion and Q. Beatty for editorial help.
Author information
Authors and Affiliations
Contributions
A.K. and J.P. performed HS-AFM experiments and performed data analysis. B.N., B.P., and E.K. performed fluorescence experiments and did data analysis. A.K., A.H., D.G.K., R.K., K.W., L.W., C.S., N.O, and J.P. developed sample preparation techniques. J.P. and A.K. designed the experiments. A.K., J.P. and P.P. prepared the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary information
Supplementary information (PDF 917 kb)
Supplementary information
Supplementary Movie 1 (MP4 677 kb)
Supplementary information
Supplementary Movie 2 (MP4 1685 kb)
Supplementary information
Supplementary Movie 3 (MP4 699 kb)
Supplementary information
Supplementary Movie 4 (MP4 1747 kb)
Supplementary information
Supplementary Movie 5 (MP4 438 kb)
Supplementary information
Supplementary Movie 6 (MP4 608 kb)
Rights and permissions
About this article
Cite this article
Karner, A., Nimmervoll, B., Plochberger, B. et al. Tuning membrane protein mobility by confinement into nanodomains. Nature Nanotech 12, 260–266 (2017). https://doi.org/10.1038/nnano.2016.236
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nnano.2016.236
This article is cited by
-
Native-like membrane models of E. coli polar lipid extract shed light on the importance of lipid composition complexity
BMC Biology (2021)
-
Localized detection of ions and biomolecules with a force-controlled scanning nanopore microscope
Nature Nanotechnology (2019)
-
High-speed AFM height spectroscopy reveals µs-dynamics of unlabeled biomolecules
Nature Communications (2018)
-
Atomic force microscopy-based characterization and design of biointerfaces
Nature Reviews Materials (2017)
-
HDL particles incorporate into lipid bilayers – a combined AFM and single molecule fluorescence microscopy study
Scientific Reports (2017)