Abstract
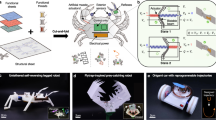
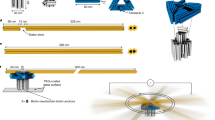
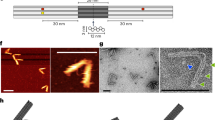
Biological systems are collections of discrete molecular objects that move around and collide with each other. Cells carry out elaborate processes by precisely controlling these collisions, but developing artificial machines that can interface with and control such interactions remains a significant challenge. DNA is a natural substrate for computing and has been used to implement a diverse set of mathematical problems1,2,3, logic circuits4,5,6 and robotics7,8,9. The molecule also interfaces naturally with living systems, and different forms of DNA-based biocomputing have already been demonstrated10,11,12,13. Here, we show that DNA origami14,15,16 can be used to fabricate nanoscale robots that are capable of dynamically interacting with each other17,18 in a living animal. The interactions generate logical outputs, which are relayed to switch molecular payloads on or off. As a proof of principle, we use the system to create architectures that emulate various logic gates (AND, OR, XOR, NAND, NOT, CNOT and a half adder). Following an ex vivo prototyping phase, we successfully used the DNA origami robots in living cockroaches (Blaberus discoidalis) to control a molecule that targets their cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Adleman, L. M. Molecular computation of solutions to combinatorial problems. Science 266, 1021–1024 (1994).
Braich, R. S., Chelyapov, N., Johnson, C., Rothemund, P. W. & Adleman, L. Solution of a 20-variable 3-SAT problem on a DNA computer. Science 296, 499–502 (2002).
Qian, L., Winfree, E. & Bruck, J. Neural network computation with DNA strand displacement cascades. Nature 475, 368–372 (2011).
Seelig, G., Soloveichik, D., Zhang, D. Y. & Winfree, E. Enzyme-free nucleic acid logic circuits. Science 314, 1585–1588 (2006).
Stojanovic, M. N., Mitchell, T. E. & Stefanovic, D. Deoxyribozyme-based logic gates. J. Am. Chem. Soc. 124, 3555–3561 (2002).
Stojanovic, M. N. et al. Deoxyribozyme-based ligase logic gates and their initial circuits. J. Am. Chem. Soc. 127, 6914–6915 (2005).
Andersen, E. S. et al. Self-assembly of a nanoscale DNA box with a controllable lid. Nature 459, 73–76 (2009).
Lund, K. et al. Molecular robots guided by prescriptive landscapes. Nature 465, 206–210 (2010).
Muscat, R. A., Bath, J. & Turberfield, A. J. A programmable molecular robot. Nano Lett. 11, 982–987 (2011).
Benenson, Y., Gil, B., Ben-Dor, U., Adar, R. & Shapiro, E. An autonomous molecular computer for logical control of gene expression. Nature 429, 423–429 (2004).
Benenson, Y. et al. Programmable and autonomous computing machine made of biomolecules. Nature 414, 430–434 (2001).
Modi, S., Nizak, C., Surana, S., Halder, S. & Krishnan, Y. Two DNA nano machines map pH changes along intersecting endocytic pathways inside the same cell. Nature Nanotech. 8, 459–467 (2013).
Rudchenko, M. et al. Autonomous molecular cascades for evaluation of cell surfaces. Nature Nanotech. 8, 580–586 (2013).
Douglas, S. M. et al. Self-assembly of DNA into nanoscale three-dimensional shapes. Nature 459, 414–418 (2009).
Douglas, S. M. et al. Rapid prototyping of 3D DNA-origami shapes with caDNAno. Nucleic Acids Res. 37, 5001–5006 (2009).
Rothemund, P. W. Folding DNA to create nanoscale shapes and patterns. Nature 440, 297–302 (2006).
Zhang, D. Y. & Seelig, G. Dynamic DNA nanotechnology using strand-displacement reactions. Nature Chem. 3, 103–113 (2011).
Zhang, D. Y. & Winfree, E. Control of DNA strand displacement kinetics using toehold exchange. J. Am. Chem. Soc. 131, 17303–17314 (2009).
Douglas, S. M., Bachelet, I. & Church, G. M. A logic-gated nanorobot for targeted transport of molecular payloads. Science 335, 831–834 (2012).
Lakin, M. R., Youssef, S., Polo, F., Emmott, S. & Phillips, A. Visual DSD: a design and analysis tool for DNA strand displacement systems. Bioinformatics 27, 3211–3213 (2011).
Lee, W. K. & Socha, J. J. Direct visualization of hemolymph flow in the heart of a grasshopper (S. americana). BMC Physiol. 9, 2 (2009).
Liu, D. et al. Resettable, multi-readout logic gates based on controllably reversible aggregation of gold nanoparticles. Angew. Chem. Int. Ed. 50, 4103–4107 (2011).
Acknowledgements
The authors thank the following colleagues for their valuable advice and comments on the manuscript: A. Adamatzky, S. Revzen, D.Y. Zhang, R. Jungmann, P. Yin, A. Marblestone, E. Shapiro, A. Munitz, A. Binshtok, L. Qian, E. Winfree and G.M. Church. The authors are particularly grateful to S.M. Douglas for valuable contributions. The authors acknowledge the members of the Bachelet lab at Bar Ilan University for support, technical help and valuable discussions. This work was supported by a European Research Council Starting grant (no. 335332) to I.B., a Kamin grant from the Israeli Ministry of Industry & Commerce to I.B. and grants from the Faculty of Life Sciences and the Institute of Nanotechnology & Advanced Materials at Bar-Ilan University.
Author information
Authors and Affiliations
Contributions
All authors designed the experiments. Y.A., E.B.I, S.I., A.A.H. and I.B. performed experiments. All authors contributed to writing the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary information
Supplementary Information (PDF 21786 kb)
Rights and permissions
About this article
Cite this article
Amir, Y., Ben-Ishay, E., Levner, D. et al. Universal computing by DNA origami robots in a living animal. Nature Nanotech 9, 353–357 (2014). https://doi.org/10.1038/nnano.2014.58
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nnano.2014.58
This article is cited by
-
Structure and dynamics of an archetypal DNA nanoarchitecture revealed via cryo-EM and molecular dynamics simulations
Nature Communications (2023)
-
A temporally resolved DNA framework state machine in living cells
Nature Machine Intelligence (2023)
-
Gene-encoding DNA origami for mammalian cell expression
Nature Communications (2023)
-
Functionalizing DNA origami to investigate and interact with biological systems
Nature Reviews Materials (2022)
-
DNA origami
Nature Reviews Methods Primers (2021)