Abstract
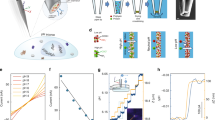
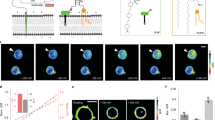
DNA is a versatile scaffold for molecular sensing in living cells, and various cellular applications of DNA nanodevices have been demonstrated. However, the simultaneous use of different DNA nanodevices within the same living cell remains a challenge. Here, we show that two distinct DNA nanomachines can be used simultaneously to map pH gradients along two different but intersecting cellular entry pathways. The two nanomachines, which are molecularly programmed to enter cells via different pathways, can map pH changes within well-defined subcellular environments along both pathways inside the same cell. We applied these nanomachines to probe the pH of early endosomes and the trans-Golgi network, in real time. When delivered either sequentially or simultaneously, both nanomachines localized into and independently captured the pH of the organelles for which they were designed. The successful functioning of DNA nanodevices within living systems has important implications for sensing and therapies in a diverse range of contexts.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Seeman, N. C. DNA in a material world. Nature 421, 427–431 (2003).
Bath, J. & Turberfield, A. J. DNA nanomachines. Nature Nanotech. 2, 275–284 (2007).
Krishnan, Y. & Bathe, M. Designer nucleic acids to probe and program the cell. Trends Cell Biol. 22, 624–633 (2012).
McMahon, D. Chemical messengers in development: a hypothesis. Science 185, 1012–1021 (1974).
Modi, S. et. al. A DNA nanomachine that maps spatial and temporal pH changes inside living cells. Nature Nanotech. 4, 325–330 (2009).
Surana, S., Bhat, J. M., Koushika, S. P. & Krishnan, Y. An autonomous DNA nanomachine maps spatiotemporal pH changes in a multicellular living organism. Nature Commun. 2, 340 (2011).
Douglas, S. M., Bachelet, I. & Church, G. M. A logic-gated nanorobot for targeted transport of molecular payloads. Science 335, 831–834 (2012).
Lee, H. et al. Molecularly self-assembled nucleic acid nanoparticles for targeted in vivo siRNA delivery. Nature Nanotech. 7, 389–393 (2012).
Bhatia, D., Surana, S., Chakraborty, S., Koushika, S. P. & Krishnan, Y. A synthetic icosahedral DNA-based host–cargo complex for functional in vivo imaging. Nature Commun. 2, 339 (2011).
Mallet, W. G. & Maxfield, F. R. Chimeric forms of furin and TGN38 are transported with the plasma membrane in the trans-Golgi network via distinct endosomal pathways. J. Cell Biol. 146, 345–359 (1999).
Chia, P. Z. C., Gasnereau, I., Lieu, Z. Z. & Gleeson, P. A. Rab9-dependent retrograde transport and endosomal sorting of the endopeptidase furin. J. Cell Sci. 124, 2401–2413 (2011).
Presley, J. F. et al. The End2 mutation in CHO cells slows the exit of transferrin receptors from the recycling compartment but bulk membrane recycling is unaffected. J. Cell Biol. 122, 1231–1241 (1993).
McCafferty, J., Griffiths, A. D., Winter, G. & Chiswell, D. J. Phage antibodies: filamentous phage displaying antibody variable domains. Nature 348, 552–554 (1990).
Geisow, M. J., D'Arcy Hart, P. & Young, M. R. Temporal changes of lysosome and phagosome pH during phagolysosome formation in macrophages: studies by fluorescence spectroscopy. J. Cell Biol. 89, 645–652 (1981).
Llopis, J., McCaffery, J. M., Miyawaki, A., Farquhar, M. G. & Tsien, R. Y. Measurement of cytosolic, mitochondrial, and golgi pH in single living cells with green fluorescent proteins. Proc. Natl Acad. Sci. USA 95, 6803–6808 (1998).
Kim, J. H. et al. Noninvasive measurement of the pH of the endoplasmic reticulum at rest and during calcium release. Proc. Natl Acad. Sci. USA 95, 2997–3002 (1998).
Molloy, S. S., Thomas, L., VanSlyke, J. K., Stenberg, P. E. & Thomas, G. Intracellular trafficking and activation of the furin proprotein convertase: localization to the TGN and recycling from the cell surface. EMBO J. 13, 18–33 (1994).
Molloy, S. S., Anderson, E. D., Jean, F. & Thomas, G. Bi-cycling the furin pathway: from TGN localization to pathogen activation and embryogenesis. Trends Cell Biol. 9, 28–35 (1999).
Presley, J. F., Mayor, S., McGraw, T. E., Dunn, K. W. & Maxfield, F. R. Bafilomycin A1 treatment retards transferrin receptor recycling more than bulk membrane recycling. J. Biol. Chem. 272, 13929–13936 (1997).
Yamashiro, D. J. & Maxfield, F. R. Acidification of morphologically distinct endosomes in mutant and wild-type Chinese hamster ovary cells. J. Cell Biol. 105, 2723–2733 (1987).
Maeda, Y., Ide, T., Koike, M., Uchiyama, Y. & Kinoshita, T. GPHR is a novel anion channel critical for acidification and functions of the Golgi apparatus. Nature Cell Biol. 10, 1135–1145 (2008).
Derivery, E. et al. The Arp2/3 Activator WASH controls the fission of endosomes through a large multiprotein complex. Dev. Cell 17, 712–723 (2009).
Mesaki, K., Tanabe, K., Obayashi, M., Oe, N. & Takei, K. Fission of tubular endosomes triggers endosomal acidification and movement. PLoS ONE 6, e19764 (2011).
Macia, E. et al. Dynasore, a cell-permeable inhibitor of dynamin. Dev. Cell 10, 839–850 (2006).
Reaves, B. & Banting, G. Perturbation of the morphology of the trans-Golgi network following Brefeldin A treatment: redistribution of a TGN-specific integral membrane protein, TGN38. J. Cell Biol. 116, 85–94 (1992).
Sciaky, N. et al. Golgi tubule traffic and the effects of brefeldin A visualized in living cells. J. Cell Biol. 139, 1137–1155 (1997).
Strous, G. J. et al. Brefeldin A induces a microtubule-dependent fusion of galactosyltransferase-containing vesicles with the rough endoplasmic reticulum. Biol. Cell 71, 25–31 (1991).
Waguri, S. et al. Visualization of TGN to endosome trafficking through fluorescently labeled MPR and AP-1 in living cells. Mol. Biol. Cell 14, 142–155 (2003).
Maeda, Y., Beznoussenko, G. V., Van Lint, J., Mironov, A. A. & Malhotra, V. Recruitment of protein kinase D to the trans-Golgi network via the first cysteine-rich domain. EMBO J. 20, 5982–5990 (2001).
Acknowledgements
The authors thank S. Mayor, D. Lilley, A. Sarin, G.V. Shivashankar and W. Shih for critical input, and the Central Imaging and Flow Facility at NCBS for imaging. The authors also thank S. Mayor for scFv libraries and the IA2.2 cell line, and J. Bonifacino and M. Marks for the Tac-furin chimera plasmids. S.M., S.S. and S.H. thank the CSIR for research fellowships. C.N. thanks NCBS for generous support of this collaboration. Y.K. thanks the Wellcome Trust–DBT India Alliance and the Innovative Young Biotechnologist Award for funding.
Author information
Authors and Affiliations
Contributions
S.M. and Y.K. conceived and designed the experiments. C.N. contributed phage display expertise. S.M. performed the in vitro and in cellulo experiments. S.S. optimized the IFu used herein, and S.H. addressed scFv–furin stability. S.M. and Y.K. analysed the data. S.M., S.S. and Y.K. co-wrote the paper and all authors commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary information
Supplementary information (PDF 5833 kb)
Supplementary movie S1
Supplementary movie S1 (MP4 842 kb)
Supplementary movie S2
Supplementary movie S2 (MP4 1010 kb)
Supplementary movie S3
Supplementary movie S3 (MP4 1009 kb)
Supplementary movie S4
Supplementary movie S4 (MP4 2045 kb)
Supplementary movie S5
Supplementary movie S5 (MP4 2059 kb)
Rights and permissions
About this article
Cite this article
Modi, S., Nizak, C., Surana, S. et al. Two DNA nanomachines map pH changes along intersecting endocytic pathways inside the same cell. Nature Nanotech 8, 459–467 (2013). https://doi.org/10.1038/nnano.2013.92
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nnano.2013.92