Abstract
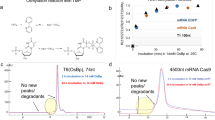
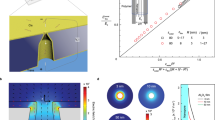
It has been proposed that single molecules of DNA could be sequenced by measuring the physical properties of the bases as they pass through a nanopore1,2. Theoretical calculations suggest that electron tunnelling can identify bases in single-stranded DNA without enzymatic processing3,4,5, and it was recently experimentally shown that tunnelling can sense individual nucleotides6 and nucleosides7. Here, we report that tunnelling electrodes functionalized with recognition reagents can identify a single base flanked by other bases in short DNA oligomers. The residence time of a single base in a recognition junction is on the order of a second, but pulling the DNA through the junction with a force of tens of piconewtons would yield reading speeds of tens of bases per second.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Zwolak, M. & Di Ventra, M. Physical approaches to DNA sequencing and detection. Rev. Mod. Phys. 80, 141–165 (2008).
Branton, D. et al. Nanopore sequencing. Nature Biotechnol. 26, 1146–1153 (2008).
Lagerqvist, J., Zwolak, M. & Di Ventra, M. Influence of the environment and probes on rapid DNA sequencing via transverse electronic transport. Biophys. J. 93, 2384–2390 (2007).
Zwolak, M. & Di Ventra, M. Electronic signature of DNA nucleotides via transverse transport. Nano Lett. 5, 421–424 (2005).
Krstic, P. S., Wells, J. C., Fuentes-Cabrera, M., Xu, D. & Lee, J. W. Toward electronic conductance characterization of DNA nucleotide bases. Solid State Phenom. 121–123, 1387–1390 (2007).
Tsutsui, M., Taniguchi, M., Yokota, K. & Kawai, T. Identification of single nucleotide via tunnelling current. Nature Nanotech. 5, 286–290 (2010).
Chang, S. et al. Electronic signature of all four DNA nucleosides in a tunneling gap. Nano Lett. 10, 1070–1075 (2010).
Clarke, J. et al. Continuous base identification for single-molecule nanopore DNA sequencing. Nature Nanotech. 4, 265–270 (2009).
Lindsay, S. et al. Recognition tunneling. Nanotechnology 21, 262001 (2010).
Gross, L., Mohn, F., Moll, N., Liljeroth, P. & Meyer, G. The chemical structure of a molecule resolved by atomic force microscopy. Science 325, 1110–1114 (2009).
Vaught, A., Jing, T. W. & Lindsay, S. M. Non-exponential tunneling in water near an electrode. Chem. Phys. Lett. 236, 306–310 (1995).
Lindsay, S. M. & Ratner, M. A. Molecular transport junctions: clearing mists. Adv. Mater. 19, 23–31 (2007).
Fuhrmann, A., Anselmetti, D., Ros, R., Getfert, S. & Reimann, P. Refined procedure of evaluating experimantal single-molecule force spectroscopy data. Phys. Rev. E 77, 031912 (2008).
Getfert, S. & Reimann, P. Optimal evaluation of single-molecule force spectroscopy experiments. Phys. Rev. E 76, 052901 (2007).
Ramachandran, G. K. et al. A bond-fluctuation mechanism for stochastic switching in wired molecules. Science 300, 1413–1415 (2003).
Goldsmith, B. R., Coroneus, J. G., Kane, A. A., Weiss, G. A. & Collins, P. G. Monitoring single-molecule reactivity on a carbon nanotube. Nano Lett. 8, 189–194 (2008).
Keyser, U. F. et al. Direct force measurements on DNA in a solid-state nanopore. Nature Phys. 2, 473–477 (2006).
Visoly-Fisher, I. et al. Conductance of a biomolecular wire. Proc. Natl Acad. Sci. USA 103, 8686–8690 (2006).
Ashcroft, B. et al. An AFM/rotaxane molecular reading head for sequence-dependent DNA structure. Small 4, 1468–1475 (2008).
Fuhrmann, A. Force Spectroscopy from Single Molecules to Whole Cells: Refined Procedures of Data Analysis. PhD thesis, Arizona State Univ. (2010).
Acknowledgements
The authors acknowledge useful discussions with O. Sankey, P. Krstic and B. Gyarfus. P. Collins made helpful comments on an earlier version of this manuscript. H. Liu composed the graphic for Fig. 1a. This work was supported by a grant from the Sequencing Technology Program of the National Human Genome Research Institute (HG004378). R.R. and A.F. were supported by a grant from the National Cancer Institute (U54CA143682).
Author information
Authors and Affiliations
Contributions
S.H., S.C. and J.H. carried out tunnelling measurements and characterized the samples. P.Z., F.L. and Sq. L. designed, synthesized and characterized reagents. M.T. prepared tunnelling probes. A.F. and R.R. carried out force spectroscopy. S.L. designed experiments, analysed data and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
S.L., P.Z. and J.H. are named as inventors in patent applications.
Supplementary information
Supplementary information
Supplementary information (PDF 2027 kb)
Rights and permissions
About this article
Cite this article
Huang, S., He, J., Chang, S. et al. Identifying single bases in a DNA oligomer with electron tunnelling. Nature Nanotech 5, 868–873 (2010). https://doi.org/10.1038/nnano.2010.213
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nnano.2010.213
This article is cited by
-
Dynamical control of nanoscale light-matter interactions in low-dimensional quantum materials
Light: Science & Applications (2024)
-
Advances in single-molecule junctions as tools for chemical and biochemical analysis
Nature Chemistry (2023)
-
Local cation-tuned reversible single-molecule switch in electric double layer
Nature Communications (2023)
-
Graphene–molecule–graphene single-molecule junctions to detect electronic reactions at the molecular scale
Nature Protocols (2023)
-
Programmable nano-reactors for stochastic sensing
Nature Communications (2021)