Abstract
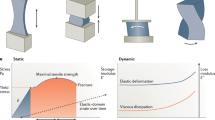
Elastomeric proteins are molecular springs that confer excellent mechanical properties1,2,3,4,5 to many biological tissues and biomaterials. Depending on the role performed by the tissue or biomaterial, elastomeric proteins can behave as molecular springs1,2,6,7 or shock absorbers3,4,5,8,9,10. Here we combine single-molecule atomic force microscopy and protein engineering techniques to create elastomeric proteins that can switch between two distinct types of mechanical behaviour in response to the binding of a molecular regulator. The proteins are mechanically labile by design and behave as entropic springs with an elasticity that is governed by their configurational entropy. However, when a molecular regulator binds to the protein, it switches into a mechanically stable state and can act as a shock absorber. These engineered proteins effectively mimic and combine the two extreme forms of elastic behaviour found in natural elastomeric proteins, and thus represent a new type of smart nanomaterial that will find potential applications in nanomechanics and material sciences.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Tatham, A. S. & Shewry, P. R. Elastomeric proteins: biological roles, structures and mechanisms. Trends Biochem. Sci. 25, 567–571 (2000).
Gosline, J. et al. Elastic proteins: biological roles and mechanical properties. Phil. Trans. R. Soc. Lond. Ser. B 357, 121–132 (2002).
Smith, B. L. et al. Molecular mechanistic origin of the toughness of natural adhesives, fibres and composites. Nature 399, 761–763 (1999).
Oberhauser, A. F., Marszalek, P. E., Erickson, H. P. & Fernandez, J. M. The molecular elasticity of the extracellular matrix protein tenascin. Nature 393, 181–185 (1998).
Labeit, S. & Kolmerer, B. Titins: giant proteins in charge of muscle ultrastructure and elasticity. Science 270, 293–296 (1995).
Elvin, C. M. et al. Synthesis and properties of crosslinked recombinant pro-resilin. Nature 437, 999–1002 (2005).
Urry, D. W. et al. Elastin: a representative ideal protein elastomer. Phil. Trans. R. Soc. Lond. B. 357, 169–184 (2002).
Li, H. et al. Reverse engineering of the giant muscle protein titin. Nature 418, 998–1002 (2002).
Lee, G. et al. Nanospring behaviour of ankyrin repeats. Nature 440, 246–249 (2006).
Bullard, B. et al. The molecular elasticity of the insect flight muscle proteins projectin and kettin. Proc. Natl Acad. Sci. USA 103, 4451–4456 (2006).
Carrion-Vazquez, M. et al. Mechanical design of proteins studied by single-molecule force spectroscopy and protein engineering. Prog. Biophys. Mol. Biol. 74, 63–91 (2000).
Lu, H. & Schulten, K. The key event in force-induced unfolding of Titin's immunoglobulin domains. Biophys. J. 79, 51–65 (2000).
Paci, E. & Karplus, M. Unfolding proteins by external forces and temperature: the importance of topology and energetics. Proc. Natl Acad. Sci. USA 97, 6521–6526 (2000).
Klimov, D. K. & Thirumalai, D. Native topology determines force-induced unfolding pathways in globular proteins. Proc. Natl Acad. Sci. USA 97, 7254–7259 (2000).
Bustanji, Y. & Samori, B. The mechanical properties of human angiostatin can be modulated by means of its disulfide bonds: A single-molecule force-spectroscopy study. Angew. Chem. Int. Ed. Engl. 41, 1546–1548 (2002).
Ainavarapu, S. R., Li, L., Badilla, C. L. & Fernandez, J. M. Ligand binding modulates the mechanical stability of dihydrofolate reductase. Biophys. J. 89, 3337–3344 (2005).
Cao, Y. & Li, H. Polyprotein of GB1 is an ideal artificial elastomeric protein. Nature Mater. 6, 109–114 (2007).
Dietz, H. & Rief, M. Exploring the energy landscape of GFP by single-molecule mechanical experiments. Proc. Natl Acad. Sci. USA 101, 16192–16197 (2004).
Gronenborn, A. M. et al. A novel, highly stable fold of the immunoglobulin binding domain of streptococcal protein G. Science 253, 657–661 (1991).
Sauer-Eriksson, A. E., Kleywegt, G. J., Uhlen, M. & Jones, T. A. Crystal structure of the C2 fragment of streptococcal protein G in complex with the Fc domain of human IgG. Structure 3, 265–278 (1995).
Cao, Y., Balamurali, M. M., Sharma, D. & Li, H. A functional single-molecule binding assay via force spectroscopy. Proc. Natl Acad. Sci. USA 104, 15677–15681 (2007).
Cao, Y., Yoo, T., Zhuang, S. & Li, H. Protein-protein interaction regulates proteins' mechanical stability. J. Mol. Biol. 378, 1132–1141 (2008).
Li, P. C. & Makarov, D. E. Ubiquitin-like protein domains show high resistance to mechanical unfolding similar to that of the 127 domain in titin: Evidence from simulations. J.Phys. Chem. B 108, 745–749 (2004).
Brockwell, D. J. et al. Mechanically unfolding the small, topologically simple protein L. Biophys. J. 89, 506–519 (2005).
Wood, S. J., Wetzel, R., Martin, J. D. & Hurle, M. R. Prolines and amyloidogenicity in fragments of the Alzheimer's peptide beta/A4. Biochemistry 34, 724–730 (1995).
Carrion-Vazquez, M. et al. Mechanical and chemical unfolding of a single protein: a comparison. Proc. Natl Acad. Sci. USA 96, 3694–3699 (1999).
Li, H., Oberhauser, A. F., Fowler, S. B., Clarke, J. & Fernandez, J. M. Atomic force microscopy reveals the mechanical design of a modular protein. Proc. Natl Acad. Sci. USA 97, 6527–6531 (2000).
Li, H. et al. Multiple conformations of PEVK proteins detected by single-molecule techniques. Proc. Natl Acad. Sci. USA 98, 10682–10686 (2001).
Sagawa, T. et al. Conformational changes in the antibody constant domains upon hapten-binding. Mol. Immunol. 42, 9–18 (2005).
Acknowledgements
This work was supported by the Natural Sciences and Engineering Research Council of Canada, Canada Research Chairs program and Canada Foundation for Innovation. H.L. is a Michael Smith Foundation for Health Research Career Investigator.
Author information
Authors and Affiliations
Contributions
Y.C. and H.L. conceived and designed the experiments. Y.C. performed the experiments and analysed the data. Y.C. and H.L. wrote the paper.
Corresponding author
Supplementary information
Rights and permissions
About this article
Cite this article
Cao, Y., Li, H. Engineered elastomeric proteins with dual elasticity can be controlled by a molecular regulator. Nature Nanotech 3, 512–516 (2008). https://doi.org/10.1038/nnano.2008.168
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nnano.2008.168
This article is cited by
-
Protein nanomechanics in biological context
Biophysical Reviews (2021)
-
Forced protein unfolding leads to highly elastic and tough protein hydrogels
Nature Communications (2013)