Abstract
Neuronal activity-induced gene expression modulates the function and plasticity of the nervous system. It is unknown whether and to what extent neuronal activity may trigger changes in chromatin accessibility, a major mode of epigenetic regulation of gene expression. Here we compared chromatin accessibility landscapes of adult mouse dentate granule neurons in vivo before and after synchronous neuronal activation using an assay for transposase-accessible chromatin using sequencing (ATAC-seq). We found genome-wide changes 1 h after activation, with enrichment of gained-open sites at active enhancer regions and at binding sites for AP1-complex components, including c-Fos. Some changes remained stable for at least 24 h. Functional analysis further implicates a critical role of c-Fos in initiating, but not maintaining, neuronal activity-induced chromatin opening. Our results reveal dynamic changes of chromatin accessibility in adult mammalian brains and suggest an epigenetic mechanism by which transient neuronal activation leads to dynamic changes in gene expression via modifying chromatin accessibility.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Crick, F. Memory and molecular turnover. Nature 312, 101 (1984).
Gräff, J., Kim, D., Dobbin, M.M. & Tsai, L.H. Epigenetic regulation of gene expression in physiological and pathological brain processes. Physiol. Rev. 91, 603–649 (2011).
West, A.E. & Greenberg, M.E. Neuronal activity-regulated gene transcription in synapse development and cognitive function. Cold Spring Harb. Perspect. Biol. http://dx.doi.org/10.1101/cshperspect.a005744 (2011).
Sweatt, J.D. The emerging field of neuroepigenetics. Neuron 80, 624–632 (2013).
Descalzi, G. et al. Epigenetic mechanisms of chronic pain. Trends Neurosci. 38, 237–246 (2015).
Lattal, K.M. & Wood, M.A. Epigenetics and persistent memory: implications for reconsolidation and silent extinction beyond the zero. Nat. Neurosci. 16, 124–129 (2013).
Meaney, M.J. & Ferguson-Smith, A.C. Epigenetic regulation of the neural transcriptome: the meaning of the marks. Nat. Neurosci. 13, 1313–1318 (2010).
Borrelli, E., Nestler, E.J., Allis, C.D. & Sassone-Corsi, P. Decoding the epigenetic language of neuronal plasticity. Neuron 60, 961–974 (2008).
Guo, J.U., Su, Y., Zhong, C., Ming, G.L. & Song, H. Emerging roles of TET proteins and 5-hydroxymethylcytosines in active DNA demethylation and beyond. Cell Cycle 10, 2662–2668 (2011).
Cholewa-Waclaw, J. et al. The role of epigenetic mechanisms in the regulation of gene expression in the nervous system. J. Neurosci. 36, 11427–11434 (2016).
Weng, Y.L., An, R., Shin, J., Song, H. & Ming, G.L. DNA modifications and neurological disorders. Neurotherapeutics 10, 556–567 (2013).
Jaenisch, R. & Bird, A. Epigenetic regulation of gene expression: how the genome integrates intrinsic and environmental signals. Nat. Genet. 33 (Suppl.), 245–254 (2003).
Vierstra, J. et al. Mouse regulatory DNA landscapes reveal global principles of cis-regulatory evolution. Science 346, 1007–1012 (2014).
Schmid, R.S. et al. Core pathway mutations induce de-differentiation of murine astrocytes into glioblastoma stem cells that are sensitive to radiation but resistant to temozolomide. Neuro-oncol. 18, 962–973 (2016).
Mo, A. et al. Epigenomic signatures of neuronal diversity in the mammalian brain. Neuron 86, 1369–1384 (2015).
Shin, J., Ming, G.L. & Song, H. Decoding neural transcriptomes and epigenomes via high-throughput sequencing. Nat. Neurosci. 17, 1463–1475 (2014).
Buenrostro, J.D., Giresi, P.G., Zaba, L.C., Chang, H.Y. & Greenleaf, W.J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Ma, D.K. et al. Neuronal activity-induced Gadd45b promotes epigenetic DNA demethylation and adult neurogenesis. Science 323, 1074–1077 (2009).
Guo, J.U. et al. Neuronal activity modifies the DNA methylation landscape in the adult brain. Nat. Neurosci. 14, 1345–1351 (2011).
Guo, J.U., Su, Y., Zhong, C., Ming, G.L. & Song, H. Hydroxylation of 5-methylcytosine by TET1 promotes active DNA demethylation in the adult brain. Cell 145, 423–434 (2011).
Lisanby, S.H. Electroconvulsive therapy for depression. N. Engl. J. Med. 357, 1939–1945 (2007).
Neunuebel, J.P. & Knierim, J.J. Spatial firing correlates of physiologically distinct cell types of the rat dentate gyrus. J. Neurosci. 32, 3848–3858 (2012).
Thurman, R.E. et al. The accessible chromatin landscape of the human genome. Nature 489, 75–82 (2012).
Frank, C.L. et al. Regulation of chromatin accessibility and Zic binding at enhancers in the developing cerebellum. Nat. Neurosci. 18, 647–656 (2015).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Davie, K. et al. Discovery of transcription factors and regulatory regions driving in vivo tumor development by ATAC-seq and FAIRE-seq open chromatin profiling. PLoS Genet. 11, e1004994 (2015).
Gjoneska, E. et al. Conserved epigenomic signals in mice and humans reveal immune basis of Alzheimer's disease. Nature 518, 365–369 (2015).
Shen, Y. et al. A map of the cis-regulatory sequences in the mouse genome. Nature 488, 116–120 (2012).
Pintchovski, S.A., Peebles, C.L., Kim, H.J., Verdin, E. & Finkbeiner, S. The serum response factor and a putative novel transcription factor regulate expression of the immediate-early gene Arc/Arg3.1 in neurons. J. Neurosci. 29, 1525–1537 (2009).
Kim, T.K. et al. Widespread transcription at neuronal activity-regulated enhancers. Nature 465, 182–187 (2010).
Kawashima, T. et al. Synaptic activity-responsive element in the Arc/Arg3.1 promoter essential for synapse-to-nucleus signaling in activated neurons. Proc. Natl. Acad. Sci. USA 106, 316–321 (2009).
Malik, A.N. et al. Genome-wide identification and characterization of functional neuronal activity-dependent enhancers. Nat. Neurosci. 17, 1330–1339 (2014).
Shlyueva, D., Stampfel, G. & Stark, A. Transcriptional enhancers: from properties to genome-wide predictions. Nat. Rev. Genet. 15, 272–286 (2014).
Young, M.D. et al. ChIP-seq analysis reveals distinct H3K27me3 profiles that correlate with transcriptional activity. Nucleic Acids Res. 39, 7415–7427 (2011).
Barski, A. et al. High-resolution profiling of histone methylations in the human genome. Cell 129, 823–837 (2007).
Ernst, J. & Kellis, M. Discovery and characterization of chromatin states for systematic annotation of the human genome. Nat. Biotechnol. 28, 817–825 (2010).
Machanick, P. & Bailey, T.L. MEME-ChIP: motif analysis of large DNA datasets. Bioinformatics 27, 1696–1697 (2011).
Curran, T. & Morgan, J.I. Fos: an immediate-early transcription factor in neurons. J. Neurobiol. 26, 403–412 (1995).
Nedivi, E., Hevroni, D., Naot, D., Israeli, D. & Citri, Y. Numerous candidate plasticity-related genes revealed by differential cDNA cloning. Nature 363, 718–722 (1993).
Jang, M.H. et al. Secreted frizzled-related protein 3 regulates activity-dependent adult hippocampal neurogenesis. Cell Stem Cell 12, 215–223 (2013).
De Rubeis, S. et al. Synaptic, transcriptional and chromatin genes disrupted in autism. Nature 515, 209–215 (2014).
Felling, R.J. & Song, H. Epigenetic mechanisms of neuroplasticity and the implications for stroke recovery. Exp. Neurol. 268, 37–45 (2015).
Ma, W., Noble, W.S. & Bailey, T.L. Motif-based analysis of large nucleotide data sets using MEME-ChIP. Nat. Protoc. 9, 1428–1450 (2014).
Ge, S. et al. GABA regulates synaptic integration of newly generated neurons in the adult brain. Nature 439, 589–593 (2006).
Duan, X. et al. Disrupted-In-Schizophrenia 1 regulates integration of newly generated neurons in the adult brain. Cell 130, 1146–1158 (2007).
Langmead, B. & Salzberg, S.L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Shen, L. et al. diffReps: detecting differential chromatin modification sites from ChIP-seq data with biological replicates. PLoS One 8, e65598 (2013).
Heger, A., Webber, C., Goodson, M., Ponting, C.P. & Lunter, G. GAT: a simulation framework for testing the association of genomic intervals. Bioinformatics 29, 2046–2048 (2013).
Robinson, J.T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Shen, L., Shao, N., Liu, X. & Nestler, E. ngs.plot: Quick mining and visualization of next-generation sequencing data by integrating genomic databases. BMC Genomics 15, 284 (2014).
Trapnell, C., Pachter, L. & Salzberg, S.L. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25, 1105–1111 (2009).
Trapnell, C. et al. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 28, 511–515 (2010).
Guo, J.U. et al. Distribution, recognition and regulation of non-CpG methylation in the adult mammalian brain. Nat. Neurosci. 17, 215–222 (2014).
Giresi, P.G. & Lieb, J.D. Isolation of active regulatory elements from eukaryotic chromatin using FAIRE (formaldehyde assisted isolation of regulatory elements). Methods 48, 233–239 (2009).
Simon, J.M., Giresi, P.G., Davis, I.J. & Lieb, J.D. Using formaldehyde-assisted isolation of regulatory elements (FAIRE) to isolate active regulatory DNA. Nat. Protoc. 7, 256–267 (2012).
Ernst, J. & Kellis, M. ChromHMM: automating chromatin-state discovery and characterization. Nat. Methods 9, 215–216 (2012).
Acknowledgements
We thank the members of the Song and Ming laboratories for discussions, K. Christian for comments and Y. Cai and L. Liu for technical support. This work was supported by NIH (R37NS047344 and P01NS097206 to H.S., R35NS097370 and R01MH105128 to G.-l.M.), SFARI (Award 240011 to H.S.), The Dr. Miriam and Sheldon G. Adelson Medical Research Foundation (to G.-l.M.) and The Brain and Behavior Research Foundation (to Y.S.).
Author information
Authors and Affiliations
Contributions
Y.S. and H.S. designed the project. C.Z. prepared the AAV and performed viral injections; Y.S., J.S., S.W., P.R., J.L. and D.K. contributed to data collection, analyses and interpretation. Y.S., J.S., G.-l.M. and H.S. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
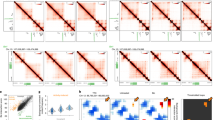
Supplementary Figure 1 Comparison of open chromatin regions between dentate granule cells and other tissues and neural cell types.
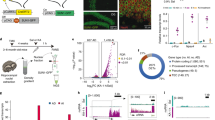
(a) Pearson correlation heatmap among open chromatin profiles of different cell types and tissues (See Supplementary Table 2 for sources of data used). (b) UCSC genome browser visualization of open chromatin profiling coverage at the Prox1 locus for the adult mouse dentate gyrus (red) and other subtypes of neuronal cells and tissues (gray). Bottom panel shows the UCSC genome browser visualization of RNA-seq coverage for dentate gyrus (blue).
Supplementary Figure 2 Characterization of open chromatin regions identified by ATAC-seq.
(a) Distribution of open chromatin regions in different genomic regions before (E0; n = 89, 946) and 1 h (E1; n = 114,959) after neuronal activation (b) ATAC-seq reads at E0 across all annotated genes are stratified by their mRNA levels (left panel) or plotted on gene bodies with their expression levels ranked in descending order for the heatmap view (right panel).
Supplementary Figure 3 Characterization of the chromatin state of regions with neuronal activity-induced chromatin-accessibility changes at E1 compared to E0.
(a) UCSC genome browser visualization of open chromatin profiling coverage at E0 and E1 and H3K4me1, H3K27Ac and H3K4me3 ChIP-seq coverage at the Arc and cFos loci for the adult mouse dentate gyrus. Red bar indicates the gained-open sites at E1. Blue bar indicates the previously functionally identified enhancer regions1,2,3. (b) Aggregate plots of histone modifications and CTCF ChIP-seq signals centered at gained-open (left panel) and gained-closed (right panel) sites at E1. The histone ChIP-seq data used for plots are from a previously published dataset from the adult mouse hippocampus4 and a CTCF ChIP-seq dataset from the adult mouse cortex5. The significance of overlap between peaks was calculated using a hypergeometric model. (c) Shown are the emission matrix, state annotation and enrichment of biological features from ChromHMM6. Refer to caption of Fig. 2c.
Supplementary Figure 4 Characterization of H3K4me1, H3K4me3, H3K27Ac and H3K27me3 changes before and after stimulation at regions with neuronal activity-induced chromatin-accessibility changes at E1 compared to E0.
Shown are aggregate plots of H3K4me1, H3K4me3, H3K27Ac and H3K27me3 signals before and after KCl stimulation centered at gained-open (up) and gained-closed (bottom) sites at E1. The histone ChIP-seq data used for plots are from a previously published dataset7 (n = 2-3).
Supplementary Figure 5 Motif prediction of neuronal activity-induced gained-open and gained-closed sites at E1 compared to E0 by two different algorithms, HOMER and MEME-ChIP.
(a) De novo motif identified by MEME-ChIP analysis of neuronal activity-induced gained-open regions at E1 resembles AP-1 complex, including cFos, JunB, Jun and JunD. Refer to Fig. 3a. (b) De novo motif identified by MEME-ChIP analysis of neuronal activation-induced gained-closed regions at E1 does not recover motifs similar to known transcription factor binding motifs (P = 4.5e -51). (c) De novo motif identified by HOMER from neuronal activity-induced gained-open and gain-closed sites at E1. ChIP-seq data for cFos, FosB, JunB and CREB used for plots were from a published dataset7
Supplementary Figure 6 Characterization of c-Fos expression in the adult mouse dentate gyrus.
(a, b) Expression of endogenous cFos in response to transient synchronous neuronal stimulation. Shown are summaries of time-course analyses of mRNA (a) and protein (b) expression of cFos in the adult dentate gyrus after electroconvulsive stimulation. Values represent mean ± s.e.m. (n = 3 for a, n = 2 for b; *P < 0.05; **P < 0.01; ANOVA). Full-length western blots are presented in Supplementary Fig. 10. (c) Schematic diagrams of AAV constructs used to express shRNA or exogenous protein in the adult mouse dentate gyrus (top panel) and the experimental paradigm (bottom panel). (d, e) Q-PCR analysis of expression of cFos in the adult dentate gyrus expressing shRNA-Ctrl and shRNA-cFos (d) or cFos and/or EYFP (e). Values represent mean ± s.e.m. (n = 3 mice; *P < 0.05; permutation test).
Supplementary Figure 7 Effect of cFos knockdown in neuronal activity-induced chromatin opening and gene expression.
(a) Confirmation of cFos knockdown efficacy upon AAV-mediated expression of shRNA-cFos in samples for RNA-seq. Values represent mean + s.e.m. (n = 3; *P < 0.05; permutation test). (b) Principal components analysis of ATAC-seq data under different experimental conditions. (c, d) Venn diagrams of differential peaks (c) and genes (d) in the adult dentate gyrus expressing shRNA-Ctrl and shRNA-cFos in response to neuronal activation. (e) Venn diagrams of gained-open regions induced by cFos overexpression and neuronal activation at E1 without exogenous cFos expression.
Supplementary Figure 8 Characterization of neuronal activation-induced chromatin-accessibility changes at E1, E4 and E24.
(a) Comparison of open chromatin profiles of the dentate gyrus of adult mice at different time points after synchronous neuronal activation. Differential regions are shown in a volcano plot. Significantly gained-open sites are shown in red and gained-closed sites are shown in blue (T-test; P < 1e -5; Fold changes > 2). (b, c) Distribution of differential peaks after neuronal activation in different genomic regions. Distal binding sites are defined as > 1 kb from an NCBI annotated RefSeq TSS (b).
Supplementary Figure 9 ChIP-qPCR analysis at the sustained gained-open regions at E4 for Kcnj6, Nr3c1, Kcnv2 and Nrxn3 in the adult dentate gyrus at E0, E1 and E4.
Data represent normalized percentage input (n = 3 mice in each group; #P > 0.05; *P < 0.05; permutation test).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–10 and Supplementary Table 2 (PDF 1598 kb)
Supplementary Table 1
Summary of open chromatin regions at E0, E1, E4 and E24. (XLSX 35678 kb)
Supplementary Table 3
Summary of dentate granule neuron-specific open chromatin regions and their associated genes compared to other neuronal subtype cells. (XLSX 1613 kb)
Supplementary Table 4
Summary of differential chromatin opening regions after synchronous neuronal activation from ATAC-seq analysis (XLSX 1629 kb)
Supplementary Table 5
Summary of RNA-seq analyses. (XLSX 1639 kb)
Supplementary Table 6
List of gained-open peaks with cFos binding site at E1 compared to E0. (XLSX 421 kb)
Supplementary Table 7
Summary of differential peaks under cFos knockdown and overexpression conditions from ATAC-seq analysis. (XLSX 1148 kb)
Rights and permissions
About this article
Cite this article
Su, Y., Shin, J., Zhong, C. et al. Neuronal activity modifies the chromatin accessibility landscape in the adult brain. Nat Neurosci 20, 476–483 (2017). https://doi.org/10.1038/nn.4494
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nn.4494
This article is cited by
-
Human neuronal maturation comes of age: cellular mechanisms and species differences
Nature Reviews Neuroscience (2024)
-
Identification of Regulatory Elements in Primary Sensory Neurons Involved in Trauma-Induced Neuropathic Pain
Molecular Neurobiology (2024)
-
Identification and characterization of transcribed enhancers during cerebellar development through enhancer RNA analysis
BMC Genomics (2023)
-
Associations between DNA methylation and gene regulation depend on chromatin accessibility during transgenerational plasticity
BMC Biology (2023)
-
Spatiotemporal proteomic atlas of multiple brain regions across early fetal to neonatal stages in cynomolgus monkey
Nature Communications (2023)