Abstract
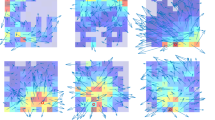
Flexible navigation demands knowledge of boundaries, routes and their relationships. Within a multi-path environment, a subpopulation of subiculum neurons robustly encoded the axis of travel. The firing of axis-tuned neurons peaked bimodally, at head orientations 180° apart. Environmental manipulations showed these neurons to be anchored to environmental boundaries but to lack axis tuning in an open arena. Axis-tuned neurons thus provide a powerful mechanism for mapping relationships between routes and the larger environmental context.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
O'Keefe, J. & Dostrovsky J. Brain Res. 34, 171–175 (1971).
McNaughton, B.L., Barnes, C.A. & O'Keefe, J. Exp. Brain Res. 52, 41–49 (1983).
Markus, E.J. et al. J. Neurosci. 15, 7079–7094 (1995).
Wood, E.R., Dudchenko, P.A., Robitsek, R.J. & Eichenbaum, H. Neuron 27, 623–633 (2000).
Frank, L.M., Brown, E.N. & Wilson, M. Neuron 27, 169–178 (2000).
O'Mara, S. J. Anat. 207, 271–282 (2005).
Witter, M.P. Behav. Brain Res. 174, 251–264 (2006).
Sharp, P.E. & Green, C. J. Neurosci. 14, 2339–2356 (1994).
Kim, S.M., Ganguli, S. & Frank, L.M. J. Neurosci. 32, 11539–11558 (2012).
Sharp, P.E. Behav. Brain Res. 85, 71–92 (1997).
Sharp, P.E. Behav. Neurosci. 113, 643–662 (1999).
Barry, C. et al. Rev. Neurosci. 17, 71–97 (2006).
Stewart, S., Jeewajee, A., Wills, T.J., Burgess, N. & Lever, C. Phil. Trans. R. Soc. Lond. B 369, 20120514 (2013).
Brotons-Mas, J.R., Montejo, N., O'Mara, S.M. & Sanchez-Vives, M.V. Eur. J. Neurosci. 32, 648–658 (2010).
Taube, J.S., Muller, R.U. & Ranck, J.B. Jr. J. Neurosci. 10, 420–435 (1990).
Derdikman, D. et al. Nat. Neurosci. 12, 1325–1332 (2009).
Nitz, D.A. Neuron 49, 747–756 (2006).
Whitlock, J.R., Pfuhl, G., Dagslott, N., Moser, M.-B. & Moser, E.I. Neuron 73, 789–802 (2012).
McNamara, T.P., Rump, B. & Werner, S. Psychon. Bull. Rev. 10, 589–595 (2003).
Berens, P. J. Stat. Softw. 31, 1–21 (2009).
Harris, K.D., Henze, D.A., Csicsvari, J., Hirase, H. & Buzsáki, G. J. Neurophysiol. 84, 401–414 (2000).
Krause, B. Nanconv.m MATLAB Central File Exchange http://www.mathworks.com/matlabcentral/fileexchange/41961-nanconv (2013, retrieved 15 January 2016).
Skaggs, W.E., McNaughton, B.L., Gothard, K.M. & Markus, E.J. An information-theoretic approach to deciphering the hippocampal code. in Advances in Neural Information Processing Systems Vol. 5 (eds. Hanson, S.J., Cowan, J.D. & Giles, C.L.) 1030–1037 (1993).
Kubie, J.L., Muller, R.U. & Bostock, E. J. Neurosci. 10, 1110–1123 (1990).
Hartley, T., Burgess, N., Lever, C., Cacucci, F. & O'Keefe, J. Hippocampus 10, 369–379 (2000).
Dempster, A.P., Laird, N.M. & Rubin, D.B. J. R. Stat. Soc. Series B Stat. Methodol. 39, 1–38 (1977).
Park, T. & Lee, T. Multichannel audio signal source separation based on an interchannel loudness vector sum. Preprint at https://arxiv.org/abs/1512.08075 (2015).
Acknowledgements
The authors wish to thank the following people for technical assistance, discussion of the data, and editing of the original manuscript: E. Tao, J. Li, E. Mukamel, L. Quinn, L. Rangel, A. Chiba, A. Alexander, L. Shelley, D. Tingley, B. Voytek, R. Gao and J. Niebaum. This work was supported by the National Science Foundation (IOS-1149718) and the Kavli Institute for Brain and Mind (#2015-055).
Author information
Authors and Affiliations
Contributions
All authors contributed significantly to the design of experiments and analyses, data collection, and manuscript organization and writing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Summary of recording site histological data.
Recordings of subiculum neurons (N=542) were obtained from a total of five four-tetrode bundles in three animals. Numbers of total recorded neurons and numbers of neurons with axis-tuned firing are included above each figure. Red arrows depict tracks left by the bundles and their approximate endpoints. Three of the recording sites were restricted to the subiculum, and two (NS15-left and the lateral bundle in NS16-right) were in a transition zone bordering the CA1 subregion. Abbreviations: RCTX (retrosplenial cortex), DG (dentate gyrus), SUB (subiculum).
Supplementary Figure 2 Waveform discrimination.
A. Histogram of isolation distance values, binned in log space, for all 542 neurons in the dataset (blue). Overlaid in orange is the histogram of isolation distance scores for the model-defined axis-tuned neurons. B. Waveform discrimination for five axis-tuned neurons recorded on the same tetrode from one recording of rat NS14. Center: Color-coded clusters of individual waveforms in a 3D plot of peak-valley voltage on wires 2, 3, and 4 (wire 1 not recorded). For each cluster, the color-coded waveform plots of all waveforms were included. Isolation distance values are reported for each cluster here as well as in C and D. Surrounding polar plots: Adjacent polar plots to each waveform plot show the directional tuning of that neuron on the track. C. Waveform discrimination of one axis-tuned neuron (red) with all other waveforms recorded (black points) shown from rat NS15. The same three plots are included here as in panel A, but the peak-valley voltage plot is only of wires 1 and 2. Wires 3 and 4 were not recorded on this tetrode. D. Waveform discrimination for three neurons recorded on a single tetrode from rat NS16. Here, two are axis-tuned and one is not.
Supplementary Figure 3 Spike characterization and open arena spatial specificity
A. Neurons (N=542) plotted based on spike width versus burst index. The distribution is consistent with prior work (Kim et al., 2012). B. Histogram of spatial information scores for the entire dataset. Similar to previous studies, the degree of spatially-specific activity among the population varied greatly. C. Shown are example neurons whose positional firing rate maps reflect the observed range of high to low specificity as measured by spatial information (Skaggs et al., 1993). Selected neurons are the median representatives from each decile of the spatial information value distribution. This is done to provide a holistic representation of the range of informativeness of SUB neurons.
Supplementary Figure 4 Open arena spatial correlates
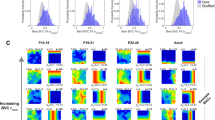
A. Four sample neurons with strong correlations to ideal place cell templates. 2D arena firing rate maps (larger images) are shown using the same color mapping as in supplemental figure 3. Max firing rate listed above and to the right of the rate map. Smaller images are the best fitting BVC (top) and place cell (bottom) templates. Pearson r values for each is overlaid on the corresponding template. In all four neurons, the place cell template r value greatly exceeds that of the BVC template, capturing the patterns seen in the actual rate map. B. Four sample neurons with strong correlations to ideal BVC templates, presented identically to those in A. C. Bottom panels: Histograms of Pearson r values for all 354 neurons with greater than 250 spikes on the arena (blue) and of just the axis-tuned neurons (orange). The left histogram shows the place cell template values while the right histogram presents the BVC template values. The red lines show the cutoff value of 0.4 chosen to characterize a neuron as well fit by the template. Middle histogram: Histogram of the difference between the best BVC and place cell templates for all neurons with either best template Pearson r value over the 0.4 cutoff (blue N = 114/354) and the axis-tuned neurons meeting the same criteria (N =12/47, orange). This orders neurons having spatially specific firing from more place-like on the left to more BVC-like on the right. Top panel: 2D arena firing rate maps from an arbitrarily-selected bin where the superior template between the place cell and BVC templates are visually identified to be inconclusive. At this juncture, approximately 40 neurons (3 axis-tuned) are more BVC-like and 75 (9 axis-tuned) are more place-like.
Supplementary Figure 5 Model schematic.
A. Example neuron training data and model fits. For all plots, directional tuning data is shown from a randomly-selected half of the track data used for training (blue rose plots). Von Mises mixture model fits of each order used (0-8) are overlaid (red ellispses). B. Directional tuning plots from the remaining half of data were used for cross-validation. Sum-squared-error (SSE) between the cross-validation data and each order model (red, same models as in A) is printed above each figure along with its SSE Ratio, the remaining error normalized by the amount of SSE of the naïve circular model (order 0). For a model to be considered a ‘best fit’, the SSE Ratio must be below 0.5 (<50% error of naïve circular model). Among model orders with SSE Ratios below 0.5, the difference between the model and preceding order model must be greater than 0.2 (20% improvement; difference value printed above plots, see also supplemental figure 5). For this neuron, the criteria lead to selection of the 2nd-order model (red box). C. To be considered as strongly axis-tuned, two more criteria must be met. First, the ratio of mean actual data firing rate at model maxima to mean actual data firing rate at model minima must exceed 2. Mean peak (maxima) values for this neuron (green lines) were 8.2X mean minima values (black lines). Finally, the spatial independence of each model’s maxima must both exceed 50%. For all track locations associated with movement in either of the two preferred directions, the neuron was considered ‘active’ if its mean rate at that position and orientation was at least 50% of the overall mean rate for that direction. In the case of this neuron, most of the points in the light green orientation lie along the light green arrows on the right panel (see also figure 1B – right panel) and the points in the dark green orientation lie along the dark green arrows. Neurons were considered spatially independent if the majority of associated locations met this criterion. For this sample neuron, the larger, light green peak had 88% spatial independence and the smaller, dark green peak had 58% spatial independence. Because it met all three criteria, this neuron is in the axis-tuned subpopulation.
Supplementary Figure 6 Model parameter flexibility.
To select the best order for the von Mises mixture model for each neuron, we trade off model complexity with fit improvement by selecting the most complicated model that yields 20% improvement in sum squared error over the preceding order model. The criterion is arbitrary but qualitatively consistent across a wide range of values. Here we plot the number of neurons in the population categorized into each order type based on a range of criterion values. Criterion values ranged from 2.5-40% by increments of 2.5% (blue lines, dark colors – light colors = low to high criterion values). We selected the 20% value for our criteria (red lines). Track model fitting data is given in the left panel. From 10-22.5%, more neurons are classified as 2nd-order than any other order. This demonstrates that this is a property of the population and not the model parameters, especially when compared to arena data (right panel), where no model parameter results in a population bias to 2nd-order (bimodal) mixture models.
Supplementary Figure 7 Track axis–tuned neurons in the open arena.
For each neuron meeting criteria for strong axis-tuned firing during the track running session, the positional firing map for the same neuron is shown for the arena foraging session. Peak (arena max) firing rates are given above each, utilizing the same color map from figures 1 and 3 and listed in the legend in the top left.
Supplementary Figure 8 Axis-tuned neurons in light versus dark.
One day of both light and dark recording was obtained which included 3 model-defined axis-tuned neurons. Each panel contains an individual axis-tuned neuron’s firing rate color-mapped as a function of track position for all time periods associated with travel >3 cm/second for both the light and dark conditions, highlighting the similarity in firing without the presence of prominent visual cues.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 (PDF 2834 kb)
Rights and permissions
About this article
Cite this article
Olson, J., Tongprasearth, K. & Nitz, D. Subiculum neurons map the current axis of travel. Nat Neurosci 20, 170–172 (2017). https://doi.org/10.1038/nn.4464
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nn.4464