Abstract
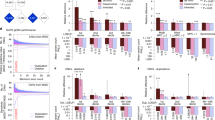
Disruptive, damaging ultra-rare variants in highly constrained genes are enriched in individuals with neurodevelopmental disorders. In the general population, this class of variants was associated with a decrease in years of education (YOE). This effect was stronger among highly brain-expressed genes and explained more YOE variance than pathogenic copy number variation but less than common variants. Disruptive, damaging ultra-rare variants in highly constrained genes influence the determinants of YOE in the general population.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
McLendon, M.K. & Perna, L.W. Ann. Am. Acad. Pol. Soc. Sci. 655, 6–15 (2014).
Haveman, R. & Wolfe, B. J. Econ. Lit. 33, 1829–1878 (1995).
Krapohl, E. et al. Proc. Natl. Acad. Sci. USA 111, 15273–15278 (2014).
Cutler, D.M. & Lleras-Muney, A. Education and health: evaluating theories and evidence. National Bureau of Economic Research Working Paper Series 12352 (2006).
Okbay, A. et al. Nature 533, 539–542 (2016).
Marioni, R.E. et al. Intelligence 44, 26–32 (2014).
Branigan, A.R., McCallum, K.J. & Freese, J. Soc. Forces 92, 109–140 (2013).
Zuk, O. et al. Proc. Natl. Acad. Sci. USA 111, E455–E464 (2014).
Lek, M. et al. Nature 536, 285–291 (2016).
Genovese, G. et al. Nat. Neurosci. http://dx.doi.org/10.1038/nn.4402 (2016).
Gilissen, C. et al. Nature 511, 344–347 (2014).
Iossifov, I. et al. Nature 515, 216–221 (2014).
Robinson, E.B. et al. Nat. Genet. 48, 552–555 (2016).
GTEx Consortium. Nat. Genet. 45, 580–585 (2013).
Joshi, P.K. et al. Nature 523, 459–462 (2015).
Stefansson, H. et al. Nature 505, 361–366 (2014).
Wu, M.C. et al. Am. J. Hum. Genet. 89, 82–93 (2011).
Davies, K. Br. J. Nurs. 15, 252–256 (2006).
Yang, J. et al. Nat. Genet. 47, 1114–1120 (2015).
Ripke, S. et al. Nat. Genet. 45, 1150–1159 (2013).
Leitsalu, L. et al. Int. J. Epidemiol. 44, 1137–1147 (2015).
Vartiainen, E. et al. Int. J. Epidemiol. 39, 504–518 (2010).
Jostins, L. et al. Nature 491, 119–124 (2012).
Schizophrenia Working Group of the Psychiatric Genomics Consortium. Nature 511, 421–427 (2014).
DePristo, M.A. et al. Nat. Genet. 43, 491–498 (2011).
Li, H. Bioinformatics 30, 2843–2851 (2014).
Cingolani, P. et al. Front. Genet. 3, 35 (2012).
Cingolani, P. et al. Fly (Austin) 6, 80–92 (2012).
Liu, X., Jian, X. & Boerwinkle, E. Hum. Mutat. 34, E2393–E2402 (2013).
Samocha, K.E. et al. Nat. Genet. 46, 944–950 (2014).
Chang, C.C. et al. Gigascience 4, 7 (2015).
Marshall, C. et al. Preprint at bioRxiv http://dx.doi.org/10.1101/040493 (2016).
Handsaker, R.E. et al. Nat. Genet. 47, 296–303 (2015).
Madsen, B.E. & Browning, S.R. PLoS Genet. 5, e1000384 (2009).
Ionita-Laza, I., Lee, S., Makarov, V., Buxbaum, J.D. & Lin, X. Am. J. Hum. Genet. 92, 841–853 (2013).
Acknowledgements
We thank R. Walters for discussions. A.G. is supported by the Knut and Alice Wallenberg Foundation (2015.0327) and the Swedish Research Council (2016-00250). M.G.N. is supported by the Royal Netherlands Academy of Science Professor Award (PAH/6635) to Dorret I. Boomsma. V.S. was supported by the Finnish Foundation for Cardiovascular Research. This study was supported by grants from the National Human Genome Research Institute (U54 HG003067 and R01 HG006855); the National Institute of Mental Health (1U01MH105666-01 and 1R01MH101244-02); the National Institute of Diabetes and Digestive and Kidney Disease (1U54DK105566-02); the Stanley Center for Psychiatric Research; the Alexander and Margaret Stewart Trust; the National Institutes of Mental Health (R01 MH077139 and RC2 MH089905); the Sylvan C. Herman Foundation; EU H2020 grants 692145, 676550 and 654248; Estonian Research Council Grant IUT20-60, NIASC, EIT–Health; NIH-BMI Grant No. 2R01DK075787-06A1; and by the EU through the European Regional Development Fund (Project No. 2014-2020.4.01.15-0012 GENTRANSMED).
Author information
Authors and Affiliations
Contributions
A.G. and G.G. designed the study, performed the analysis and wrote the manuscript. B.M.N. supervised the project. D.H., A. Byrnes, M.I.K., S.M.Z., C.W.W., M.K., M.G.N., P.N. and R.M. performed the analyses. A. Bloemendal, J.M.B., J.I.G., T.P., C.S. and R.E.H. developed and provided computational tools. D.G. provide data management support. E.B.R., A.M., V.S., J.S., S.M.P., P.S., S.K., M.J.D., S.A.M., P.F.S., A.P., T.E. and C.M.H. collected and provided the data.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 2 Association between disruptive, damaging and synonymous URVs and YOE in cases and controls of schizophrenia.
Similar associations are observed in cases and control of schizophrenia. Total individuals included N=14,133.
Supplementary Figure 3 Association between disruptive, damaging and synonymous URVs and YOE after excluding individuals with neurodevelopmental disorders.
Only Swedish non-schizophrenic participants are included in this analysis. Similar associations are observed once individuals with neurodevelopmental disorders are excluded.
Supplementary Figure 4 Association between disruptive and damaging URVs with YOE for different quantiles of brain gene-expression.
Only Swedish participants are included (N=10,651).
Supplementary Figure 5 Association between disruptive, damaging and synonymous URVs and YOE in highly brain-specific HC genes, sparsely brain-specific HC genes and all highly brain-specific genes.
Meta-analysis results (N=14,133).
Supplementary Figure 6 Association between polygenic score and YOE, in individuals carrying 0 or ≥1 URVs or pathogenic CNVs
Only Swedish participants are included (N=10,651).
Supplementary Figure 7 Q-Q plots for the gene-based burden test for association between disruptive and damaging variants and YOE.
Each point is a gene. Results are from meta-analysis across studies (N=14,133). In the upper panels we included only URVs, in the lower panels we included variants with minor allele frequency < 0.05%.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 (PDF 1104 kb)
Supplementary Methods Checklist
(PDF 397 kb)
Supplementary Table 1
Educational attainment coding for each study (XLSX 10 kb)
Supplementary Table 2
Distribution of YOE by study, year of birth and sex (XLSX 10 kb)
Supplementary Table 3
Number and distribution of URVs by study (XLSX 10 kb)
Supplementary Table 4
Large pathogenic CNVs included in the study and number of carriers (XLSX 13 kb)
Rights and permissions
About this article
Cite this article
Ganna, A., Genovese, G., Howrigan, D. et al. Ultra-rare disruptive and damaging mutations influence educational attainment in the general population. Nat Neurosci 19, 1563–1565 (2016). https://doi.org/10.1038/nn.4404
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nn.4404
This article is cited by
-
H2A monoubiquitination: insights from human genetics and animal models
Human Genetics (2023)
-
Ultra-rare and common genetic variant analysis converge to implicate negative selection and neuronal processes in the aetiology of schizophrenia
Molecular Psychiatry (2022)
-
Contribution of rare whole-genome sequencing variants to plasma protein levels and the missing heritability
Nature Communications (2022)
-
Exome sequencing in bipolar disorder identifies AKAP11 as a risk gene shared with schizophrenia
Nature Genetics (2022)
-
Genetic associations with mathematics tracking and persistence in secondary school
npj Science of Learning (2020)