Abstract
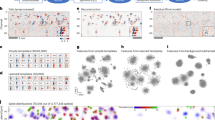
Developments in microfabrication technology have enabled the production of neural electrode arrays with hundreds of closely spaced recording sites, and electrodes with thousands of sites are under development. These probes in principle allow the simultaneous recording of very large numbers of neurons. However, use of this technology requires the development of techniques for decoding the spike times of the recorded neurons from the raw data captured from the probes. Here we present a set of tools to solve this problem, implemented in a suite of practical, user-friendly, open-source software. We validate these methods on data from the cortex, hippocampus and thalamus of rat, mouse, macaque and marmoset, demonstrating error rates as low as 5%.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Buzsáki, G. Large-scale recording of neuronal ensembles. Nat. Neurosci. 7, 446–451 (2004).
Wise, K.D. & Najafi, K. Microfabrication techniques for integrated sensors and microsystems. Science 254, 1335–1342 (1991).
Csicsvari, J. et al. Massively parallel recording of unit and local field potentials with silicon-based electrodes. J. Neurophysiol. 90, 1314–1323 (2003).
McNaughton, B.L., O'Keefe, J. & Barnes, C.A. The stereotrode: a new technique for simultaneous isolation of several single units in the central nervous system from multiple unit records. J. Neurosci. Methods 8, 391–397 (1983).
Gray, C.M., Maldonado, P.E., Wilson, M. & McNaughton, B. Tetrodes markedly improve the reliability and yield of multiple single-unit isolation from multi-unit recordings in cat striate cortex. J. Neurosci. Methods 63, 43–54 (1995).
Wilson, M.A. & McNaughton, B.L. Dynamics of the hippocampal ensemble code for space. Science 261, 1055–1058 (1993).
Recce, M. & O'Keefe, J. The tetrode: a new technique for multi-unit extracellular recording. Soc. Neurosci. Abstr. 15, 1250 (1989).
Harris, K.D., Henze, D.A., Csicsvari, J., Hirase, H. & Buzsáki, G. Accuracy of tetrode spike separation as determined by simultaneous intracellular and extracellular measurements. J. Neurophysiol. 84, 401–414 (2000).
Henze, D.A. et al. Intracellular features predicted by extracellular recordings in the hippocampus in vivo. J. Neurophysiol. 84, 390–400 (2000).
Gold, C., Henze, D.A., Koch, C. & Buzsáki, G. On the origin of the extracellular action potential waveform: A modeling study. J. Neurophysiol. 95, 3113–3128 (2006).
Einevoll, G.T., Franke, F., Hagen, E., Pouzat, C. & Harris, K.D. Towards reliable spike-train recordings from thousands of neurons with multielectrodes. Curr. Opin. Neurobiol. 22, 11–17 (2012).
Lewicki, M.S. A review of methods for spike sorting: the detection and classification of neural action potentials. Network 9, R53–R78 (1998).
Hazan, L., Zugaro, M. & Buzsáki, G. Klusters, NeuroScope, NDManager: a free software suite for neurophysiological data processing and visualization. J. Neurosci. Methods 155, 207–216 (2006).
Briggman, K.L., Helmstaedter, M. & Denk, W. Wiring specificity in the direction-selectivity circuit of the retina. Nature 471, 183–188 (2011).
Berényi, A. et al. Large-scale, high-density (up to 512 channels) recording of local circuits in behaving animals. J. Neurophysiol. 111, 1132–1149 (2014).
Du, J., Blanche, T.J., Harrison, R.R., Lester, H.A. & Masmanidis, S.C. Multiplexed, high density electrophysiology with nanofabricated neural probes. PLoS One 6, e26204 (2011).
Bouveyron, C. & Brunet-Saumard, C. Model-based clustering of high-dimensional data: a review. Comput. Stat. Data Anal. 71, 52–78 (2014).
Ekanadham, C., Tranchina, D. & Simoncelli, E.P. A unified framework and method for automatic neural spike identification. J. Neurosci. Methods 222, 47–55 (2014).
Carlson, D.E. et al. Multichannel electrophysiological spike sorting via joint dictionary learning and mixture modeling. IEEE Trans. Biomed. Eng. 61, 41–54 (2014).
Calabrese, A. & Paninski, L. Kalman filter mixture model for spike sorting of non-stationary data. J. Neurosci. Methods 196, 159–169 (2011).
Franke, F., Natora, M., Boucsein, C., Munk, M.H. & Obermayer, K. An online spike detection and spike classification algorithm capable of instantaneous resolution of overlapping spikes. J. Comput. Neurosci. 29, 127–148 (2010).
Quiroga, R.Q., Nadasdy, Z. & Ben-Shaul, Y. Unsupervised spike detection and sorting with wavelets and superparamagnetic clustering. Neural Comput. 16, 1661–1687 (2004).
Swindale, N.V. & Spacek, M.A. Spike sorting for polytrodes: a divide and conquer approach. Front. Syst. Neurosci. 8, 6 (2014).
Swindale, N.V. & Spacek, M.A. Spike detection methods for polytrodes and high density microelectrode arrays. J. Comput. Neurosci. 38, 249–261 (2015).
Buzsáki, G. & Kandel, A. Somadendritic backpropagation of action potentials in cortical pyramidal cells of the awake rat. J. Neurophysiol. 79, 1587–1591 (1998).
Logothetis, N.K., Kayser, C. & Oeltermann, A. In vivo measurement of cortical impedance spectrum in monkeys: implications for signal propagation. Neuron 55, 809–823 (2007).
Harris, K.D., Hirase, H., Leinekugel, X., Henze, D.A. & Buzsáki, G. Temporal interaction between single spikes and complex spike bursts in hippocampal pyramidal cells. Neuron 32, 141–149 (2001).
Quirk, M.C., Blum, K.I. & Wilson, M.A. Experience-dependent changes in extracellular spike amplitude may reflect regulation of dendritic action potential back-propagation in rat hippocampal pyramidal cells. J. Neurosci. 21, 240–248 (2001).
Quirk, M.C. & Wilson, M.A. Interaction between spike waveform classification and temporal sequence detection. J. Neurosci. Methods 94, 41–52 (1999).
Kadir, S.N., Goodman, D.F. & Harris, K.D. High-dimensional cluster analysis with the masked EM algorithm. Neural Comput. 26, 2379–2394 (2014).
Fowlkes, E.B. & Mallows, C.L. A method for comparing 2 hierarchical clusterings. J. Am. Stat. Assoc. 78, 553–569 (1983).
Schmitzer-Torbert, N., Jackson, J., Henze, D., Harris, K. & Redish, A.D. Quantitative measures of cluster quality for use in extracellular recordings. Neuroscience 131, 1–11 (2005).
Hill, D.N., Mehta, S.B. & Kleinfeld, D. Quality metrics to accompany spike sorting of extracellular signals. J. Neurosci. 31, 8699–8705 (2011).
Owens, J.D. et al. GPU computing. Proc. IEEE 96, 879–899 (2008).
Freeman, J. et al. Mapping brain activity at scale with cluster computing. Nat. Methods 11, 941–950 (2014).
Comaniciu, D. & Meer, P. Mean shift: a robust approach toward feature space analysis. IEEE Trans. Pattern Anal. Mach. Intell. 24, 603–619 (2002).
Rodriguez, A. & Laio, A. Machine learning. Clustering by fast search and find of density peaks. Science 344, 1492–1496 (2014).
Marre, O. et al. Mapping a complete neural population in the retina. J. Neurosci. 32, 14859–14873 (2012).
Pillow, J.W., Shlens, J., Chichilnisky, E.J. & Simoncelli, E.P. A model-based spike sorting algorithm for removing correlation artifacts in multi-neuron recordings. PLoS One 8, e62123 (2013).
Saleem, A.B., Ayaz, A., Jeffery, K.J., Harris, K.D. & Carandini, M. Integration of visual motion and locomotion in mouse visual cortex. Nat. Neurosci. 16, 1864–1869 (2013).
Ayaz, A., Saleem, A.B., Schölvinck, M.L. & Carandini, M. Locomotion controls spatial integration in mouse visual cortex. Curr. Biol. 23, 890–894 (2013).
Ecker, A.S. et al. State dependence of noise correlations in macaque primary visual cortex. Neuron 82, 235–248 (2014).
Ecker, A.S. et al. Decorrelated neuronal firing in cortical microcircuits. Science 327, 584–587 (2010).
Zeater, N., Cheong, S.K., Solomon, S.G., Dreher, B. & Martin, P.R. Binocular visual responses in the primate lateral geniculate nucleus. Curr. Biol. 25, 3190–3195 (2015).
The HDF Group. Hierarchical Data Format, version 5. http://www.hdfgroup.org/HDF5/ (2014).
Rossant, C. & Harris, K.D. Hardware-accelerated interactive data visualization for neuroscience in Python. Front. Neuroinform. 7, 36 (2013).
Shreiner, D., Sellers, G., Kessenich, J.M., Licea-Kane, B. & Khronos OpenGL ARB Working Group. OpenGL Programming Guide: The Official Guide to Learning OpenGL, version 4.3. 8th edn. (Addison Wesley, 2013).
Swayne, D.F., Cook, D. & Buja, A. XGobi: interactive dynamic data visualization in the X Window System. J. Comput. Graph. Stat. 7, 113–130 (1998).
Acknowledgements
We thank the 200+ members of the klustaviewas@groups.google.com mailing list for their feedback, bug reports and suggestions. This work was supported by EPSRC (K015141, I005102, K.D.H.) and the Wellcome Trust (95668, 95669, 100154, K.D.H., M.C.). M.C. is supported by the GlaxoSmithKline/Fight for Sight chair in Visual Neuroscience.
Author information
Authors and Affiliations
Contributions
C.R., D.F.M.G., S.N.K. and J.S. wrote SpikeDetekt. K.D.H., S.N.K. and D.F.M.G. designed the masked EM algorithm and wrote KlustaKwik. C.R. and M.L.D.H. wrote KlustaViewa. C.R. wrote Galry. S.N.K. analyzed a lgorithm performance. Rat data were recorded by A.G., M.B. and G.B. Mouse data were recorded by A.B.S. and M.C. Marmoset data were recorded by S.S. The procedure for non-chronic laminar recordings with NeuroNexus Vector probes in awake, behaving macaques was developed by G.H.D., A.S.E. and A.S.T., who also collected the data. K.D.H., S.N.K. and C.R. wrote the manuscript with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–17 and Supplementary Table 1 (PDF 4198 kb)
Rights and permissions
About this article
Cite this article
Rossant, C., Kadir, S., Goodman, D. et al. Spike sorting for large, dense electrode arrays. Nat Neurosci 19, 634–641 (2016). https://doi.org/10.1038/nn.4268
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nn.4268
This article is cited by
-
Neuronal dynamics direct cerebrospinal fluid perfusion and brain clearance
Nature (2024)
-
An actor-model framework for visual sensory encoding
Nature Communications (2024)
-
Temporal scaling of motor cortical dynamics reveals hierarchical control of vocal production
Nature Neuroscience (2024)
-
A developmental increase of inhibition promotes the emergence of hippocampal ripples
Nature Communications (2024)
-
Local origin of excitatory–inhibitory tuning equivalence in a cortical network
Nature Neuroscience (2024)