Abstract
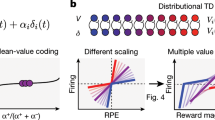
Dopamine neurons are thought to signal reward prediction error, or the difference between actual and predicted reward. How dopamine neurons jointly encode this information, however, remains unclear. One possibility is that different neurons specialize in different aspects of prediction error; another is that each neuron calculates prediction error in the same way. We recorded from optogenetically identified dopamine neurons in the lateral ventral tegmental area (VTA) while mice performed classical conditioning tasks. Our tasks allowed us to determine the full prediction error functions of dopamine neurons and compare them to each other. We found marked homogeneity among individual dopamine neurons: their responses to both unexpected and expected rewards followed the same function, just scaled up or down. As a result, we were able to describe both individual and population responses using just two parameters. Such uniformity ensures robust information coding, allowing each dopamine neuron to contribute fully to the prediction error signal.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Wise, R.A. Dopamine, learning and motivation. Nat. Rev. Neurosci. 5, 483–494 (2004).
Salamone, J.D. & Correa, M. The mysterious motivational functions of mesolimbic dopamine. Neuron 76, 470–485 (2012).
Bromberg-Martin, E.S., Matsumoto, M. & Hikosaka, O. Dopamine in motivational control: rewarding, aversive, and alerting. Neuron 68, 815–834 (2010).
Matsuda, W. et al. Single nigrostriatal dopaminergic neurons form widely spread and highly dense axonal arborizations in the neostriatum. J. Neurosci. 29, 444–453 (2009).
Fuxe, K. et al. The discovery of central monoamine neurons gave volume transmission to the wired brain. Prog. Neurobiol. 90, 82–100 (2010).
Grace, A.A. & Bunney, B.S. Intracellular and extracellular electrophysiology of nigral dopaminergic neurons. 1. Identification and characterization. Neuroscience 10, 301–315 (1983).
Vandecasteele, M., Glowinski, J. & Venance, L. Electrical synapses between dopaminergic neurons of the substantia nigra pars compacta. J. Neurosci. 25, 291–298 (2005).
Joshua, M. et al. Synchronization of midbrain dopaminergic neurons is enhanced by rewarding events. Neuron 62, 695–704 (2009).
Morris, G., Arkadir, D., Nevet, A., Vaadia, E. & Bergman, H. Coincident but distinct messages of midbrain dopamine and striatal tonically active neurons. Neuron 43, 133–143 (2004).
Kim, Y., Wood, J. & Moghaddam, B. Coordinated activity of ventral tegmental neurons adapts to appetitive and aversive learning. PLoS One 7, e29766 (2012).
Schultz, W. Predictive reward signal of dopamine neurons. J. Neurophysiol. 80, 1–27 (1998).
Schultz, W. Updating dopamine reward signals. Curr. Opin. Neurobiol. 23, 229–238 (2013).
Roeper, J. Dissecting the diversity of midbrain dopamine neurons. Trends Neurosci. 36, 336–342 (2013).
Blaess, S. et al. Temporal-spatial changes in Sonic Hedgehog expression and signaling reveal different potentials of ventral mesencephalic progenitors to populate distinct ventral midbrain nuclei. Neural Dev. 6, 29 (2011).
Margolis, E.B., Lock, H., Hjelmstad, G.O. & Fields, H.L. The ventral tegmental area revisited: is there an electrophysiological marker for dopaminergic neurons? J. Physiol. (Lond.) 577, 907–924 (2006).
Neuhoff, H., Neu, A., Liss, B. & Roeper, J. I(h) channels contribute to the different functional properties of identified dopaminergic subpopulations in the midbrain. J. Neurosci. 22, 1290–1302 (2002).
Lammel, S. et al. Input-specific control of reward and aversion in the ventral tegmental area. Nature 491, 212–217 (2012).
Watabe-Uchida, M., Zhu, L., Ogawa, S.K., Vamanrao, A. & Uchida, N. Whole-brain mapping of direct inputs to midbrain dopamine neurons. Neuron 74, 858–873 (2012).
Lammel, S. et al. Unique properties of mesoprefrontal neurons within a dual mesocorticolimbic dopamine system. Neuron 57, 760–773 (2008).
Fiorillo, C.D., Yun, S.R. & Song, M.R. Diversity and homogeneity in responses of midbrain dopamine neurons. J. Neurosci. 33, 4693–4709 (2013).
Matsumoto, M. & Hikosaka, O. Two types of dopamine neuron distinctly convey positive and negative motivational signals. Nature 459, 837–841 (2009).
Bayer, H.M. & Glimcher, P.W. Midbrain dopamine neurons encode a quantitative reward prediction error signal. Neuron 47, 129–141 (2005).
Cohen, J.Y., Haesler, S., Vong, L., Lowell, B.B. & Uchida, N. Neuron-type-specific signals for reward and punishment in the ventral tegmental area. Nature 482, 85–88 (2012).
Pan, W.-X., Schmidt, R., Wickens, J.R. & Hyland, B.I. Dopamine cells respond to predicted events during classical conditioning: evidence for eligibility traces in the reward-learning network. J. Neurosci. 25, 6235–6242 (2005).
D'Ardenne, K., McClure, S.M., Nystrom, L.E. & Cohen, J.D. BOLD responses reflecting dopaminergic signals in the human ventral tegmental area. Science 319, 1264–1267 (2008).
Waelti, P., Dickinson, A. & Schultz, W. Dopamine responses comply with basic assumptions of formal learning theory. Nature 412, 43–48 (2001).
Schultz, W., Dayan, P. & Montague, P.R. A neural substrate of prediction and reward. Science 275, 1593–1599 (1997).
Eshel, N. et al. Arithmetic and local circuitry underlying dopamine prediction errors. Nature 525, 243–246 (2015).
Tobler, P.N., Fiorillo, C.D. & Schultz, W. Adaptive coding of reward value by dopamine neurons. Science 307, 1642–1645 (2005).
Averbeck, B.B., Latham, P.E. & Pouget, A. Neural correlations, population coding and computation. Nat. Rev. Neurosci. 7, 358–366 (2006).
Cohen, M.R. & Kohn, A. Measuring and interpreting neuronal correlations. Nat. Neurosci. 14, 811–819 (2011).
Dayan, P. Twenty-five lessons from computational neuromodulation. Neuron 76, 240–256 (2012).
Abbott, L.F. & Dayan, P. The effect of correlated variability on the accuracy of a population code. Neural Comput. 11, 91–101 (1999).
Freund, T.F., Powell, J.F. & Smith, A.D. Tyrosine hydroxylase-immunoreactive boutons in synaptic contact with identified striatonigral neurons, with particular reference to dendritic spines. Neuroscience 13, 1189–1215 (1984).
Zahm, D.S. An electron microscopic morphometric comparison of tyrosine hydroxylase immunoreactive innervation in the neostriatum and the nucleus accumbens core and shell. Brain Res. 575, 341–346 (1992).
Turrigiano, G. Homeostatic signaling: the positive side of negative feedback. Curr. Opin. Neurobiol. 17, 318–324 (2007).
Davis, G.W. Homeostatic control of neural activity: from phenomenology to molecular design. Annu. Rev. Neurosci. 29, 307–323 (2006).
Fiorillo, C.D., Song, M.R. & Yun, S.R. Multiphasic temporal dynamics in responses of midbrain dopamine neurons to appetitive and aversive stimuli. J. Neurosci. 33, 4710–4725 (2013).
Fiorillo, C.D. Towards a general theory of neural computation based on prediction by single neurons. PLoS One 3, e3298 (2008).
Ganguli, S. et al. One-dimensional dynamics of attention and decision making in LIP. Neuron 58, 15–25 (2008).
Fitzgerald, J.K. et al. Biased associative representations in parietal cortex. Neuron 77, 180–191 (2013).
Matsumoto, M. & Takada, M. Distinct representations of cognitive and motivational signals in midbrain dopamine neurons. Neuron 79, 1011–1024 (2013).
Zweifel, L.S. et al. Activation of dopamine neurons is critical for aversive conditioning and prevention of generalized anxiety. Nat. Neurosci. 14, 620–626 (2011).
Grace, A.A., Floresco, S.B., Goto, Y. & Lodge, D.J. Regulation of firing of dopaminergic neurons and control of goal-directed behaviors. Trends Neurosci. 30, 220–227 (2007).
Menegas, W. et al. Dopamine neurons projecting to the posterior striatum form an anatomically distinct subclass. eLife 4, e10032 (2015).
Lerner, T.N. et al. Intact-brain analyses reveal distinct information carried by SNc dopamine subcircuits. Cell 162, 635–647 (2015).
Beier, K.T. et al. Circuit architecture of VTA dopamine neurons revealed by systematic input-output mapping. Cell 162, 622–634 (2015).
Zhang, L., Doyon, W.M., Clark, J.J., Phillips, P.E.M. & Dani, J.A. Controls of tonic and phasic dopamine transmission in the dorsal and ventral striatum. Mol. Pharmacol. 76, 396–404 (2009).
Tritsch, N.X., Ding, J.B. & Sabatini, B.L. Dopaminergic neurons inhibit striatal output through non-canonical release of GABA. Nature 490, 262–266 (2012).
Stuber, G.D., Hnasko, T.S., Britt, J.P., Edwards, R.H. & Bonci, A. Dopaminergic terminals in the nucleus accumbens but not the dorsal striatum corelease glutamate. J. Neurosci. 30, 8229–8233 (2010).
Bäckman, C.M. et al. Characterization of a mouse strain expressing Cre recombinase from the 3′ untranslated region of the dopamine transporter locus. Genesis 44, 383–390 (2006).
Boyden, E.S., Zhang, F., Bamberg, E., Nagel, G. & Deisseroth, K. Millisecond-timescale, genetically targeted optical control of neural activity. Nat. Neurosci. 8, 1263–1268 (2005).
Atasoy, D., Aponte, Y., Su, H.H. & Sternson, S.M. A FLEX switch targets Channelrhodopsin-2 to multiple cell types for imaging and long-range circuit mapping. J. Neurosci. 28, 7025–7030 (2008).
Uchida, N. & Mainen, Z.F. Speed and accuracy of olfactory discrimination in the rat. Nat. Neurosci. 6, 1224–1229 (2003).
Tian, J. & Uchida, N. Habenula lesions reveal that multiple mechanisms underlie dopamine prediction errors. Neuron 87, 1304–1316 (2015).
Schmitzer-Torbert, N. & Redish, A.D. Neuronal activity in the rodent dorsal striatum in sequential navigation: separation of spatial and reward responses on the multiple T task. J. Neurophysiol. 91, 2259–2272 (2004).
Lima, S.Q., Hromádka, T., Znamenskiy, P. & Zador, A.M. PINP: a new method of tagging neuronal populations for identification during in vivo electrophysiological recording. PLoS One 4, e6099 (2009).
Kvitsiani, D. et al. Distinct behavioural and network correlates of two interneuron types in prefrontal cortex. Nature 498, 363–366 (2013).
Olsen, S.R., Bhandawat, V. & Wilson, R.I. Divisive normalization in olfactory population codes. Neuron 66, 287–299 (2010).
Acknowledgements
We thank J. Fitzgerald for assistance with analysis, J. Assad, R. Born, J. Maunsell, R. Wilson and members of the Uchida laboratory for discussions, C. Dulac for sharing resources, and K. Deisseroth (Stanford University) for the AAV-FLEX-ChR2 construct. This work was supported by a Sackler Fellowship in Psychobiology (N.E.) and US National Institutes of Health grants T32GM007753 (N.E.), F30MH100729 (N.E.), 2T32MH020017-16 (M.B.), 5T32MH020017-17 (M.B.), R01MH095953 (N.U.) and R01MH101207 (N.U.).
Author information
Authors and Affiliations
Contributions
N.E., J.T. and N.U. designed the experiments. N.E. and M.B. collected data for the variable-reward task. J.T. collected data for the variable-expectation task. N.E. analyzed data and wrote the manuscript, with comments from J.T. and N.U.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Recording sites.
(a, b) Schematic of recording locations for mice used in the variable-reward task (a, n = 5) and the variable-expectation task (b, n = 5). RN, red nucleus. SNc, substantia nigra pars compacta. SNr, substantia nigra pars reticulata.
Supplementary Figure 2 VTA neurons cluster into three response types.
(a) Responses of all neurons recorded in the variable-reward task (n = 170). Each row reflects the auROC values for a single neuron in the second before and after delivery of expected reward. Baseline is taken as one second before odor onset. Yellow, increase from baseline; cyan, decrease from baseline. Light-identified neurons are denoted by an * to the left of each row. (b) The first three principal components of the auROC curves. These values were used for unsupervised hierarchical clustering, as shown in the dendrogram on the right. (c) Average firing rates for the three clusters of neurons. Orange, unexpected reward trials. Black, expected reward trials. (d-f) Same conventions as a-c, except for neurons recorded in the variable-expectation task. All 31 light-identified dopamine neurons were classified as Type 1.
Supplementary Figure 3 Light identification of dopamine neurons.
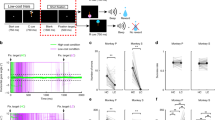
(a) Raw signal from one example light-identified dopamine neuron in the variable-reward task. Blue bars, light pulses. (b) For the same neuron, mean waveforms for spontaneous (black) and light-evoked (blue) action potentials. (c) For the same neuron, raster plots for 20 Hz (left) and 50 Hz (right) laser stimulation. Each row is one trial of laser stimulation (10 pulses of laser). (d) Histogram of log P values for each neuron recorded in the variable-reward task (n = 170). The P values were derived from SALT (see Methods). Neurons with P < 0.001 and waveform correlations > 0.9 were considered identified (filled bars). (e, f) For light-identified neurons, probability of spiking (e) and latency to first spike (f) after laser pulses at different frequencies. Orange circles, mean across neurons. (g) Histogram of mean latencies (left) and latency standard deviations (right) in response to laser stimulation for all light-identified dopamine neurons in the variable-reward task. (h-n) Same conventions as a-g, but for neurons recorded in the variable-expectation task (n = 106).
Supplementary Figure 4 Putative and identified dopamine neurons respond similarly on the variable-reward task.
(a) Average putative dopamine neuron responses (mean ± s.e.m.) for different sizes of unexpected (orange circle) and expected (black circle) reward. Orange line, best-fit Hill function for unexpected reward. Black line, subtractive shift of the orange line. n = 84 neurons. (b) Response to unexpected 2.5 μL reward versus effect of expectation for this reward size. Line, best-fit linear regression. Grey dots, putative dopamine neurons. Blue dots, light-identified dopamine neurons. Pearson’s correlation across all neurons, P = 1 x 10−10. R, correlation coefficient. (c) Baseline firing rates versus effect of expectation (averaged across reward sizes). P = 0.01. (d) Difference between reward-predicting odor and nothing-predicting odor versus difference between unexpected reward and expected reward. P = 3 x 10−6.
Supplementary Figure 5 Subtraction is scaled for each reward size.
(a-f) For identified dopamine neurons in the variable-reward experiment (n = 40), response to unexpected reward versus effect of expectation for 0.1 μL (a, P = 1.1 x 10−10), 0.3 μL (b, P = 4.4 x 10−9), 1.2 μL (c, P = 3.4 x 10−9), 5 μL (d, P = 1.9 x 10−8), 10 μL (e, P = 5 x 10−5), and 20 μL (f, P = 8.1 x 10−5) reward. R, correlation coefficient.
Supplementary Figure 6 Noise correlations for pairs of putative dopamine and GABA neurons.
(a, b) Noise correlations (mean ± s.e.m.) between pairs of simultaneously-recorded putative dopamine (Type 1) and GABA (Type 2) neurons in the variable-reward experiment (a, n = 59 pairs) and the variable-expectation experiment (b, n = 44 pairs). Correlations were calculated by examining trial-by-trial variations in spiking during different task epochs (see Methods). Grey bars, correlations on simultaneous trials. Black bars, correlations in which one neuron’s data was shifted by one trial. (c, d) Histograms of noise correlations between pairs of simultaneously-recorded putative dopamine and GABA neurons. Data are combined from both the variable-reward and variable-expectation experiments, and reflect correlations during the reward-predicting cue (c) and during delivery of expected reward (d). Filled bars, significant noise correlation (P < 0.05, Pearson’s correlation). Empty bars, n.s. Dotted lines, mean noise correlation.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 and Supplementary Note (PDF 1345 kb)
Rights and permissions
About this article
Cite this article
Eshel, N., Tian, J., Bukwich, M. et al. Dopamine neurons share common response function for reward prediction error. Nat Neurosci 19, 479–486 (2016). https://doi.org/10.1038/nn.4239
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nn.4239