Abstract
The mammalian basal forebrain (BF) has important roles in controlling sleep and wakefulness, but the underlying neural circuit remains poorly understood. We examined the BF circuit by recording and optogenetically perturbing the activity of four genetically defined cell types across sleep-wake cycles and by comprehensively mapping their synaptic connections. Recordings from channelrhodopsin-2 (ChR2)-tagged neurons revealed that three BF cell types, cholinergic, glutamatergic and parvalbumin-positive (PV+) GABAergic neurons, were more active during wakefulness and rapid eye movement (REM) sleep (wake/REM active) than during non-REM (NREM) sleep, and activation of each cell type rapidly induced wakefulness. By contrast, activation of somatostatin-positive (SOM+) GABAergic neurons promoted NREM sleep, although only some of them were NREM active. Synaptically, the wake-promoting neurons were organized hierarchically by glutamatergic→cholinergic→PV+ neuron excitatory connections, and they all received inhibition from SOM+ neurons. Together, these findings reveal the basic organization of the BF circuit for sleep-wake control.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Siegel, J.M. Sleep viewed as a state of adaptive inactivity. Nat. Rev. Neurosci. 10, 747–753 (2009).
Von Economo, C. Sleep as a problem of localization. J. Nerv. Ment. Dis. 71, 249–259 (1930).
Nauta, W.J. Hypothalamic regulation of sleep in rats; an experimental study. J. Neurophysiol. 9, 285–316 (1946).
Moruzzi, G. & Magoun, H.W. Brain stem reticular formation and activation of the EEG. Electroencephalogr. Clin. Neurophysiol. 1, 455–473 (1949).
Saper, C.B., Fuller, P.M., Pedersen, N.P., Lu, J. & Scammell, T.E. Sleep state switching. Neuron 68, 1023–1042 (2010).
Jones, B.E. Neurobiology of waking and sleeping. Handb. Clin. Neurol. 98, 131–149 (2011).
Brown, R.E., Basheer, R., McKenna, J.T., Strecker, R.E. & McCarley, R.W. Control of sleep and wakefulness. Physiol. Rev. 92, 1087–1187 (2012).
Szymusiak, R. & McGinty, D. Sleep suppression following kainic acid-induced lesions of the basal forebrain. Exp. Neurol. 94, 598–614 (1986).
Szymusiak, R. & McGinty, D. Sleep-waking discharge of basal forebrain projection neurons in cats. Brain Res. Bull. 22, 423–430 (1989).
Alam, M.N., McGinty, D. & Szymusiak, R. Thermosensitive neurons of the diagonal band in rats: relation to wakefulness and non-rapid eye movement sleep. Brain Res. 752, 81–89 (1997).
Duque, A., Balatoni, B., Detari, L. & Zaborszky, L. EEG correlation of the discharge properties of identified neurons in the basal forebrain. J. Neurophysiol. 84, 1627–1635 (2000).
Lee, M.G., Manns, I.D., Alonso, A. & Jones, B.E. Sleep-wake related discharge properties of basal forebrain neurons recorded with micropipettes in head-fixed rats. J. Neurophysiol. 92, 1182–1198 (2004).
Hassani, O.K., Lee, M.G., Henny, P. & Jones, B.E. Discharge profiles of identified GABAergic in comparison to cholinergic and putative glutamatergic basal forebrain neurons across the sleep-wake cycle. J. Neurosci. 29, 11828–11840 (2009).
Takahashi, K., Lin, J.S. & Sakai, K. Characterization and mapping of sleep-waking specific neurons in the basal forebrain and preoptic hypothalamus in mice. Neuroscience 161, 269–292 (2009).
McGinty, D.J. & Sterman, M.B. Sleep suppression after basal forebrain lesions in the cat. Science 160, 1253–1255 (1968).
Buzsaki, G. et al. Nucleus basalis and thalamic control of neocortical activity in the freely moving rat. J. Neurosci. 8, 4007–4026 (1988).
Sallanon, M. et al. Long-lasting insomnia induced by preoptic neuron lesions and its transient reversal by muscimol injection into the posterior hypothalamus in the cat. Neuroscience 32, 669–683 (1989).
Lin, J.S., Sakai, K., Vanni-Mercier, G. & Jouvet, M. A critical role of the posterior hypothalamus in the mechanisms of wakefulness determined by microinjection of muscimol in freely moving cats. Brain Res. 479, 225–240 (1989).
Nishino, S. et al. Muscle atonia is triggered by cholinergic stimulation of the basal forebrain: implication for the pathophysiology of canine narcolepsy. J. Neurosci. 15, 4806–4814 (1995).
Kaur, S., Junek, A., Black, M.A. & Semba, K. Effects of ibotenate and 192IgG-saporin lesions of the nucleus basalis magnocellularis/substantia innominata on spontaneous sleep and wake states and on recovery sleep after sleep deprivation in rats. J. Neurosci. 28, 491–504 (2008).
Fuller, P.M., Sherman, D., Pedersen, N.P., Saper, C.B. & Lu, J. Reassessment of the structural basis of the ascending arousal system. J. Comp. Neurol. 519, 933–956 (2011).
Deurveilher, S. & Semba, K. Basal forebrain regulation of cortical activity and sleep-wake states: Roles of cholinergic and non-cholinergic neurons. Sleep Biol. Rhythms 9, 65–70 (2011).
Jones, B.E. The organization of central cholinergic systems and their functional importance in sleep-waking states. Prog. Brain Res. 98, 61–71 (1993).
Lee, M.G., Hassani, O.K., Alonso, A. & Jones, B.E. Cholinergic basal forebrain neurons burst with theta during waking and paradoxical sleep. J. Neurosci. 25, 4365–4369 (2005).
Everitt, B.J. & Robbins, T.W. Central cholinergic systems and cognition. Annu. Rev. Psychol. 48, 649–684 (1997).
Sarter, M., Hasselmo, M.E., Bruno, J.P. & Givens, B. Unraveling the attentional functions of cortical cholinergic inputs: interactions between signal-driven and cognitive modulation of signal detection. Brain Res. Brain Res. Rev. 48, 98–111 (2005).
Pinto, L. et al. Fast modulation of visual perception by basal forebrain cholinergic neurons. Nat. Neurosci. 16, 1857–1863 (2013).
Fu, Y. et al. A cortical circuit for gain control by behavioral state. Cell 156, 1139–1152 (2014).
Eggermann, E., Kremer, Y., Crochet, S. & Petersen, C.C. Cholinergic signals in mouse barrel cortex during active whisker sensing. Cell Reports 9, 1654–1660 (2014).
Zaborszky, L. & Duque, A. Local synaptic connections of basal forebrain neurons. Behav. Brain Res. 115, 143–158 (2000).
Lein, E.S. et al. Genome-wide atlas of gene expression in the adult mouse brain. Nature 445, 168–176 (2007).
Xu, X., Roby, K.D. & Callaway, E.M. Immunochemical characterization of inhibitory mouse cortical neurons: three chemically distinct classes of inhibitory cells. J. Comp. Neurol. 518, 389–404 (2010).
Madisen, L. et al. A robust and high-throughput Cre reporting and characterization system for the whole mouse brain. Nat. Neurosci. 13, 133–140 (2010).
Madisen, L. et al. A toolbox of Cre-dependent optogenetic transgenic mice for light-induced activation and silencing. Nat. Neurosci. 15, 793–802 (2012).
Anikeeva, P. et al. Optetrode: a multichannel readout for optogenetic control in freely moving mice. Nat. Neurosci. 15, 163–170 (2012).
Modirrousta, M., Mainville, L. & Jones, B.E. Gabaergic neurons with alpha2-adrenergic receptors in basal forebrain and preoptic area express c-Fos during sleep. Neuroscience 129, 803–810 (2004).
Irmak, S.O. & de Lecea, L. Basal forebrain cholinergic modulation of sleep transitions. Sleep 37, 1941–1951 (2014).
Han, Y. et al. Selective activation of cholinergic basal forebrain neurons induces immediate sleep-wake transitions. Curr. Biol. 24, 693–698 (2014).
Freund, T.F. & Meskenaite, V. Gamma-aminobutyric acid-containing basal forebrain neurons innervate inhibitory interneurons in the neocortex. Proc. Natl. Acad. Sci. USA 89, 738–742 (1992).
Kim, T. et al. Cortically projecting basal forebrain parvalbumin neurons regulate cortical gamma band oscillations. Proc. Natl. Acad. Sci. USA 112, 3535–3540 (2015).
Steriade, M. & McCarley, R.W. Brain Control of Wakefulness and Sleep (Plenum Press, New York, 2005).
Goard, M. & Dan, Y. Basal forebrain activation enhances cortical coding of natural scenes. Nat. Neurosci. 12, 1444–1449 (2009).
Szymusiak, R., Gvilia, I. & McGinty, D. Hypothalamic control of sleep. Sleep Med. 8, 291–301 (2007).
Sherin, J.E., Shiromani, P.J., McCarley, R.W. & Saper, C.B. Activation of ventrolateral preoptic neurons during sleep. Science 271, 216–219 (1996).
Szymusiak, R., Alam, N., Steininger, T.L. & McGinty, D. Sleep-waking discharge patterns of ventrolateral preoptic/anterior hypothalamic neurons in rats. Brain Res. 803, 178–188 (1998).
Lu, J., Greco, M.A., Shiromani, P. & Saper, C.B. Effect of lesions of the ventrolateral preoptic nucleus on NREM and REM sleep. J. Neurosci. 20, 3830–3842 (2000).
Sherin, J.E., Elmquist, J.K., Torrealba, F. & Saper, C.B. Innervation of histaminergic tuberomammillary neurons by GABAergic and galaninergic neurons in the ventrolateral preoptic nucleus of the rat. J. Neurosci. 18, 4705–4721 (1998).
Steininger, T.L., Gong, H., McGinty, D. & Szymusiak, R. Subregional organization of preoptic area/anterior hypothalamic projections to arousal-related monoaminergic cell groups. J. Comp. Neurol. 429, 638–653 (2001).
Zhang, S. et al. Selective attention. Long-range and local circuits for top-down modulation of visual cortex processing. Science 345, 660–665 (2014).
Lambolez, B., Audinat, E., Bochet, P., Crepel, F. & Rossier, J. AMPA receptor subunits expressed by single Purkinje cells. Neuron 9, 247–258 (1992).
Pfeffer, C.K., Xue, M., He, M., Huang, Z.J. & Scanziani, M. Inhibition of inhibition in visual cortex: the logic of connections between molecularly distinct interneurons. Nat. Neurosci. 16, 1068–1076 (2013).
Danik, M., Puma, C., Quirion, R. & Williams, S. Widely expressed transcripts for chemokine receptor CXCR1 in identified glutamatergic, gamma-aminobutyric acidergic, and cholinergic neurons and astrocytes of the rat brain: a single-cell reverse transcription-multiplex polymerase chain reaction study. J. Neurosci. Res. 74, 286–295 (2003).
Nordenankar, K. et al. Increased hippocampal excitability and impaired spatial memory function in mice lacking VGLUT2 selectively in neurons defined by tyrosine hydroxylase promoter activity. Brain Struct. Funct. 220, 2171–2190 (2015).
Weissbourd, B. et al. Presynaptic partners of dorsal raphe serotonergic and GABAergic neurons. Neuron 83, 645–662 (2014).
Watakabe, A., Komatsu, Y., Ohsawa, S. & Yamamori, T. Fluorescent in situ hybridization technique for cell type identification and characterization in the central nervous system. Methods 52, 367–374 (2010).
Acknowledgements
We thank J. Cox, F. Weber and N. Hirai for the help with data analysis, K. Kao for technical assistance, T. Hnasko (University of California, San Diego) for sharing VGLUT2-eGFP mice, and the Stanford Neuroscience Gene Vector and Virus Core for AAV-DJ.
Author information
Authors and Affiliations
Contributions
M.X., S.C. and Y.D. conceived and designed the experiments. M.X. performed all of the optrode recording experiments, some of the in situ hybridization experiments and some of the slice recording experiments. S.C. performed histological characterization of BF cell types and all of the optogenetic activation experiments. S.Z. performed some of the slice experiments. P.Z. performed some of the slice recording experiments. C.M. and W.-C.C. performed some of the in situ hybridization experiments. N.S. and S.N. helped to establish sleep recording and data analysis. B.W. and L.L. provided reagents and helped to establish in situ hybridization. M.X., S.C. and Y.D. wrote the manuscript, and all of the authors helped with the revision of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
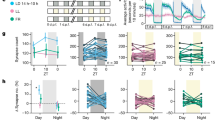
Supplementary Figure 1 Firing rates of BF neurons in natural wake, NREM, and REM states.
a. Upper panel, firing rates of each identified cholinergic neuron in the three brain states. Each line represents data from one neuron. Lower panel, firing rates averaged across all 12 cholinergic neurons. Error bar, ± s.e.m.
b. Similar to a, for glutamatergic neurons.
c. Similar to a, for PV+ neurons.
d. Similar to a, for SOM+ neurons. Right plot, expanded view of low firing-rate range. Red lines, strongly modulated NREM-active SOM+ neurons.
e. Firing rate of an example wake-active SOM+ neuron (blue line in d). Top panel, EEG power spectrogram (0-25 Hz). Middle panel, EMG trace. Bottom panel, firing rate of the SOM+ neuron. Brain states are color coded (wake, gray; REM, orange; NREM, white).
Supplementary Figure 2 ChR2-eYFP expression in the BF induced by AAV injection and optogenetically activated neurons identified by cFos staining.
a. Fluorescence images of coronal sections at virus injection site for optogenetic activation of each BF cell type, showing location of ChR2-eYFP expression. White outlines indicate BF, including the diagonal band of Broca (NDB), magnocellular preoptic nucleus (MA), and substantia innominata (SI), based on Allen Mouse Brain Reference Atlas (http://mouse.brain-map.org/static/atlas). Inset, superposition of ChR2-eYFP (green) and cFos staining (red), shown at a higher magnification for a small region of the BF; cFos staining was performed after optogenetic activation of ChAT+, VGLUT2+, PV+, or SOM+ neurons using the same protocol as that for the experiment shown in Fig. 3. Blue dotted line indicates position of the optic fiber.
b. Number of cFos+, ChR2-eYFP+ neurons near the optic fiber tip. Error bar, ± s.e.m.
Supplementary Figure 3 Effect of laser stimulation of BF neurons on transition probability between brain states.
a. Left, probability of NREM (NR) → wake (W) state transition within each 10 s period in ChAT-Cre mice expressing ChR2 in BF. The transition probability is defined as the number of trials in which the NR → W transition occurred within the given time bin divided by the number of trials in which the animal was in NR in the previous time bin. Blue shading, period of laser stimulation (60 s, 10 Hz). Error bar, ± s.d. (bootstrap). Baseline transition probability (red line) was computed after excluding the laser stimulation period. Right, probability of W → W transition.
b. Similar to a, for VGLUT2+ neuron activation.
c. Similar to a, for PV+ neuron activation.
d. W→ NR and NR → NR transitions with SOM+ neuron activation. In all cases the probability during laser stimulation was significantly higher than the baseline (P < 10-4, bootstrap). These analyses were based on the same data shown in Fig. 4.
Supplementary Figure 4 Synaptic interactions between BF cell types measured in brain slices.
a. ChAT → VGLUT2 connections. Left, responses of an example VGLUT2+ neuron to activation of ChAT+ neurons by a single 5-ms pulse of blue light (cyan dot) and a train of 5 pulses (10 Hz). Right, responses of another VGLUT2+ neuron (same as in Fig. 5c) before (blue) and after successive blockade of GABAA receptors (brown), mAChRs (red), and nAChRs (black). These pharmacological experiments indicated that the large, prolonged inhibitory response was mediated not by GABAA receptors but by mAChRs, and the small transient excitatory response was mediated by nAChRs.
b. Diversity of ChAT → SOM connections. Left, an example SOM+ neuron exhibiting fasting spiking in response to depolarizing current injection and fast excitatory response evoked by light activation of ChAT+ neurons. Right, another SOM+ neuron with low-frequency spiking evoked by current injection and slow inhibitory response to ChAT+ neuron activation.
c. Lack of PV → ChAT input. Left, confocal image of BF slice showing presynaptic PV+ neurons expressing ChR2-mCherry (red) and postsynaptic ChAT+ neurons expressing eGFP (green). Upper right, reliable spiking of an example PV+ neuron evoked by a light pulse (5 ms) or light step (200 ms), recorded with cell-attached recording. Lower right, whole-cell recording from an example ChAT+ neuron showing many spontaneous IPSCs but no light-evoked response. The recording was made under voltage clamp at 0 mV.
Supplementary Figure 5 Unclassified brain state induced by optogenetic activation of cholinergic BF neurons.
a. Example trials in which laser stimulation induced wakefulness of ChAT-ChR2 mice. Shown are EEG power spectrum, EEG traces during selected periods (indicated by boxes), and EMG trace in each trial. Blue bar, period of laser stimulation. Note the increase in EMG activity upon laser stimulation.
b. Similar to a, but for trials in which laser stimulation induced EEG desynchronization but no change in EMG.
c. Average EEG power spectra during natural wake, NREM and REM sleep, and during laser stimulation in trials without EMG change (see examples in b). Note that the EEG power spectrum in these trials is different from any of the natural brain states, thus the brain state was left unclassified in our study. The peak at ~6 Hz (lower than the frequency of theta band observed in natural REM sleep) has also been observed in the study of Han et al.38, and in their study the brain state was classified as REM sleep.
d. Probability of wake, NREM, REM, and unclassified states before, during, and after laser stimulation, averaged across 5 ChAT-ChR2 mice (same as Fig. 4c, but including probability of unclassified state).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5 and Supplementary Tables 1 and 2 (PDF 1651 kb)
Optrode recording from an identified BF cholinergic neuron across different brain states.
Shown are EEG power spectrogram, EMG trace, and spiking activity of the neuron, together with video recording of the ChAT-ChR2 mouse during wake, NREM and REM periods. Blue square indicates laser pulse train, which reliably evoked spiking from this neuron. The movie is shown at 4 × the original speed. (MOV 12586 kb)
Effect of optogenetic activation of glutamatergic BF neurons on brain states.
Shown are EEG power spectrogram, EMG trace, brain state, and video recording of the VGLUT2-Cre mouse (injected with Cre-inducible AAV expressing ChR2-eYFP in the BF). Laser stimulation (10 ms/pulse, 10 Hz, 30 s) during NREM (trials 1 and 2) or REM (trial 3) sleep caused immediate transitions into wakefulness. The movie is shown at 7 × the original speed. (MOV 7521 kb)
Rights and permissions
About this article
Cite this article
Xu, M., Chung, S., Zhang, S. et al. Basal forebrain circuit for sleep-wake control. Nat Neurosci 18, 1641–1647 (2015). https://doi.org/10.1038/nn.4143
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nn.4143
This article is cited by
-
Improved green and red GRAB sensors for monitoring spatiotemporal serotonin release in vivo
Nature Methods (2024)
-
Rethinking Neuroscientific Methodology: Lived Experience in Behavioral Studies
Biological Theory (2024)
-
Dexmedetomidine modulates neuronal activity of horizontal limbs of diagonal band via α2 adrenergic receptor in mice
BMC Anesthesiology (2023)
-
Selective optogenetic modulation of the PBN terminals in the lateral hypothalamic area and basal forebrain regulates emergence from isoflurane anesthesia in mice
BMC Anesthesiology (2023)
-
Activation of astrocytes in the basal forebrain in mice facilitates isoflurane-induced loss of consciousness and prolongs recovery
BMC Anesthesiology (2023)