Abstract
Noncoding variants in the human MIR137 gene locus increase schizophrenia risk with genome-wide significance. However, the functional consequence of these risk alleles is unknown. Here we examined induced human neurons harboring the minor alleles of four disease-associated single nucleotide polymorphisms in MIR137. We observed increased MIR137 levels compared to those in major allele–carrying cells. microRNA-137 gain of function caused downregulation of the presynaptic target genes complexin-1 (Cplx1), Nsf and synaptotagmin-1 (Syt1), leading to impaired vesicle release. In vivo, miR-137 gain of function resulted in changes in synaptic vesicle pool distribution, impaired induction of mossy fiber long-term potentiation and deficits in hippocampus-dependent learning and memory. By sequestering endogenous miR-137, we were able to ameliorate the synaptic phenotypes. Moreover, reinstatement of Syt1 expression partially restored synaptic plasticity, demonstrating the importance of Syt1 as a miR-137 target. Our data provide new insight into the mechanism by which miR-137 dysregulation can impair synaptic plasticity in the hippocampus.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
Change history
08 July 2016
It has come to our attention that our results, as presented, have caused some confusion among readers with regard to genetic variation at the MIR137 locus, gene expression of MIR137 and schizophrenia risk. It should be noted that it is currently unclear how gene variants at the MIR137 locus confers disease risk and which are the causative or functional alleles or the gene(s) that mediate risk. Also, it is not clear to what extent a single disease-associated SNP, in isolation, relates to disease. In our study we did not examine the effects of single alleles since they often segregate together. It remains to be determined how the multiple reported disease-associated SNPs in the MIR137 locus alone and in combination influence gene expression. To clarify this would require genome editing to alter SNPs, one by one, in cell lines. Finally, further studies are required to determine whether the synaptic and behavioral changes observed in this study can be linked to the specific genetic variants within the MIR137 locus associated with schizophrenia in genome-wide association studies.
References
Cross-Disorder Group of the Psychiatric Genomics Consortium. Identification of risk loci with shared effects on five major psychiatric disorders: a genome-wide analysis. Lancet 381, 1371–1379 (2013).
Ripke, S. et al. Genome-wide association analysis identifies 13 new risk loci for schizophrenia. Nat. Genet. 45, 1150–1159 (2013).
Bartel, D.P. MicroRNAs: target recognition and regulatory functions. Cell 136, 215–233 (2009).
Kosik, K.S. The neuronal microRNA system. Nat. Rev. Neurosci. 7, 911–920 (2006).
Esquela-Kerscher, A. & Slack, F.J. Oncomirs — microRNAs with a role in cancer. Nat. Rev. Cancer 6, 259–269 (2006).
Herzer, S., Silahtaroglu, A. & Meister, B. Locked nucleic acid-based in situ hybridisation reveals miR-7a as a hypothalamus-enriched microRNA with a distinct expression pattern. J. Neuroendocrinol. 24, 1492–1504 (2012).
Smrt, R.D. et al. MicroRNA miR-137 regulates neuronal maturation by targeting ubiquitin ligase Mind Bomb-1. Stem Cells 28, 1060–1070 (2010).
Kesner, R.P., Lee, I. & Gilbert, P. A behavioral assessment of hippocampal function based on a subregional analysis. Rev. Neurosci. 15, 333–351 (2004).
Szulwach, K.E. et al. Cross talk between microRNA and epigenetic regulation in adult neurogenesis. J. Cell Biol. 189, 127–141 (2010).
Heckers, S. et al. Impaired recruitment of the hippocampus during conscious recollection in schizophrenia. Nat. Neurosci. 1, 318–323 (1998).
Won, H., Mah, W. & Kim, E. Autism spectrum disorder causes, mechanisms, and treatments: focus on neuronal synapses. Front. Mol. Neurosci. 6, 19 (2013).
Hunt, R., Sauna, Z.E., Ambudkar, S.V., Gottesman, M.M. & Kimchi-Sarfaty, C. Silent (synonymous) SNPs: should we care about them? Single Nucleotide Polymorphisms: Methods and Protocols 2nd edn. 578, 23–39 (2009).
Vierbuchen, T. et al. Direct conversion of fibroblasts to functional neurons by defined factors. Nature 463, 1035–1041 (2010).
Busskamp, V. et al. Rapid neurogenesis through transcriptional activation in human stem cells. Mol. Syst. Biol. 10, 760 (2014).
Kwon, E., Wang, W. & Tsai, L.H. Validation of schizophrenia-associated genes CSMD1, C10orf26, CACNA1C and TCF4 as miR-137 targets. Mol. Psychiatry 18, 11–12 (2013).
Weinberger, D.R. Cell biology of the hippocampal formation in schizophrenia. Biol. Psychiatry 45, 395–402 (1999).
Wenzel, H.J., Cole, T.B., Born, D.E., Schwartzkroin, P.A. & Palmiter, R.D. Ultrastructural localization of zinc transporter-3 (ZnT-3) to synaptic vesicle membranes within mossy fiber boutons in the hippocampus of mouse and monkey. Proc. Natl. Acad. Sci. USA 94, 12676–12681 (1997).
Kamiya, H., Shinozaki, H. & Yamamoto, C. Activation of metabotropic glutamate receptor type 2/3 suppresses transmission at rat hippocampal mossy fibre synapses. J. Physiol. (Lond.) 493, 447–455 (1996).
Phillips, R.G. & LeDoux, J.E. Differential contribution of amygdala and hippocampus to cued and contextual fear conditioning. Behav. Neurosci. 106, 274–285 (1992).
Devanna, P. & Vernes, S.C. A direct molecular link between the autism candidate gene RORa and the schizophrenia candidate MIR137. Sci. Rep. 4, 3994 (2014).
Ebert, M.S., Neilson, J.R. & Sharp, P.A. MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells. Nat. Methods 4, 721–726 (2007).
Chapman, E.R. How does synaptotagmin trigger neurotransmitter release? Annu. Rev. Biochem. 77, 615–641 (2008).
Tang, J. et al. A complexin/synaptotagmin 1 switch controls fast synaptic vesicle exocytosis. Cell 126, 1175–1187 (2006).
Xu, J., Pang, Z.P.P., Shin, O.H. & Südhof, T.C. Synaptotagmin-1 functions as a Ca2+ sensor for spontaneous release. Nat. Neurosci. 12, 759–766 (2009).
Südhof, T.C. Calcium control of neurotransmitter release. Cold Spring Harb. Perspect. Biol. 4, a011353 (2012).
Bacaj, T. et al. Synaptotagmin-1 and Synaptotagmin-7 trigger synchronous and asynchronous phases of neurotransmitter release. Neuron 80, 947–959 (2013).
Nicoll, R.A. & Schmitz, D. Synaptic plasticity at hippocampal mossy fibre synapses. Nat. Rev. Neurosci. 6, 863–876 (2005).
Chen, Q. et al. Association and expression study of synapsin III and schizophrenia. Neurosci. Lett. 465, 248–251 (2009).
Sawada, K. et al. Hippocampal complexin proteins and cognitive dysfunction in schizophrenia. Arch. Gen. Psychiatry 62, 263–272 (2005).
Vawter, M.P. et al. Reduction of synapsin in the hippocampus of patients with bipolar disorder and schizophrenia. Mol. Psychiatry 7, 571–578 (2002).
Eastwood, S.L., Cotter, D. & Harrison, P.J. Cerebellar synaptic protein expression in schizophrenia. Neuroscience 105, 219–229 (2001).
Mirnics, K., Middleton, F.A., Marquez, A., Lewis, D.A. & Levitt, P. Molecular characterization of schizophrenia viewed by microarray analysis of gene expression in prefrontal cortex. Neuron 28, 53–67 (2000).
Castillo, P.E. et al. Rab3A is essential for mossy fibre long-term potentiation in the hippocampus. Nature 388, 590–593 (1997).
Lonart, G., Janz, R., Johnson, K.M. & Südhof, T.C. Mechanism of action of rab3A in mossy fiber LTP. Neuron 21, 1141–1150 (1998).
Cao, P., Yang, X.F. & Südhof, T.C. Complexin activates exocytosis of distinct secretory vesicles controlled by different synaptotagmins. J. Neurosci. 33, 1714–1727 (2013).
Martin, J.A., Hu, Z.T., Fenz, K.M., Fernandez, J. & Dittman, J.S. Complexin has opposite effects on two modes of synaptic vesicle fusion. Curr. Biol. 21, 97–105 (2011).
Xu, W. et al. Distinct neuronal coding schemes in memory revealed by selective erasure of fast synchronous synaptic transmission. Neuron 73, 990–1001 (2012).
Jorquera, R.A., Huntwork-Rodriguez, S., Akbergenova, Y., Cho, R.W. & Littleton, J.T. Complexin controls spontaneous and evoked neurotransmitter release by regulating the timing and properties of synaptotagmin activity. J. Neurosci. 32, 18234–18245 (2012).
Geppert, M. et al. Synaptotagmin-1: a major Ca2+ sensor for transmitter release at a central synapse. Cell 79, 717–727 (1994).
Jorgensen, E.M. et al. Defective recycling of synaptic vesicles in Synaptotagmin mutants of Caenorhabditis elegans. Nature 378, 196–199 (1995).
Littleton, J.T. et al. SNARE-complex disassembly by NSF follows synaptic-vesicle fusion. Proc. Natl. Acad. Sci. USA 98, 12233–12238 (2001).
Cesca, F., Baldelli, P., Valtorta, F. & Benfenati, F. The synapsins: key actors of synapse function and plasticity. Prog. Neurobiol. 91, 313–348 (2010).
Pieribone, V.A. et al. Expression of synapsin III in nerve terminals and neurogenic regions of the adult brain. J. Comp. Neurol. 454, 105–114 (2002).
Feng, J. et al. Regulation of neurotransmitter release by synapsin III. J. Neurosci. 22, 4372–4380 (2002).
Kao, H.T. & Porton, B. Synaptic vesicle associated proteins and schizophrenia. in Schizophrenia 3rd edn. (eds. Javitt, D. & Kantrowitz, J.) 267–284 (Springer, 2009).
Gordon Frankle, W., Lerma, J. & Laruelle, M. The synaptic hypothesis of schizophrenia. Neuron 39, 205–216 (2003).
Harrison, P.J. & Weinberger, D.R. Schizophrenia genes, gene expression, and neuropathology: on the matter of their convergence. Mol. Psychiatry 10, 40–68 (2005).
Föcking, M. et al. Proteomic and genomic evidence implicates the postsynaptic density in schizophrenia. Mol. Psychiatry 20, 424–432 (2015).
Wen, Z. et al. Synaptic dysregulation in a human iPS cell model of mental disorders. Nature 515, 414–418 (2014).
Yoshimizu, T. et al. Functional implications of a psychiatric risk variant within CACNA1C in induced human neurons. Mol. Psychiatry 20, 162–169 (2015).
Pfisterer, U. et al. Efficient induction of functional neurons from adult human fibroblasts. Cell Cycle 10, 3311–3316 (2011).
Dittgen, T. et al. Lentivirus-based genetic manipulations of cortical neurons and their optical and electrophysiological monitoring in vivo. Proc. Natl. Acad. Sci. USA 101, 18206–18211 (2004).
van Hooijdonk, L.W.A. et al. Lentivirus-mediated transgene delivery to the hippocampus reveals sub-field specific differences in expression. BMC Neurosci. 10, 2 (2009).
Vicario-Abejón, C. Long-term culture of hippocampal neurons. Curr. Protoc. Neurosci. 3.2.1–3.2.13 (2004).
Cetin, A., Komai, S., Eliava, M., Seeburg, P.H. & Osten, P. Stereotaxic gene delivery in the rodent brain. Nat. Protoc. 1, 3166–3173 (2006).
Morris, R. Developments of a water-maze procedure for studying spatial-learning in the rat. J. Neurosci. Methods 11, 47–60 (1984).
Moy, S.S. et al. Sociability and preference for social novelty in five inbred strains: an approach to assess autistic-like behavior in mice. Genes Brain Behav. 3, 287–302 (2004).
Peça, J. et al. Shank3 mutant mice display autistic-like behaviours and striatal dysfunction. Nature 472, 437–442 (2011).
Reid, C.A. & Clements, J.D. Postsynaptic expression of long-term potentiation in the rat dentate gyrus demonstrated by variance-mean analysis. J. Physiol. (Lond.) 518, 121–130 (1999).
Colino, A. & Malenka, R.C. Mechanisms underlying induction of long-term potentiation in rat medial and lateral perforate paths in vitro. J. Neurophysiol. 69, 1150–1159 (1993).
McNaughton, B.L. Evidence for 2 physiologically distinct perforant pathways to the fascia dentata. Brain Res. 199, 1–19 (1980).
Pasquinelli, A.E. Non-coding RNA microRNAs and their targets: recognition, regulation and an emerging reciprocal relationship. Nat. Rev. Genet. 13, 271–282 (2012).
Griffiths-Jones, S., Saini, H.K., van Dongen, S. & Enright, A.J. miRBase: tools for microRNA genomics. Nucleic Acids Res. 36, D154–D158 (2008).
Krek, A. et al. Combinatorial microRNA target predictions. Nat. Genet. 37, 495–500 (2005).
Lewis, B.P., Burge, C.B. & Bartel, D.P. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 120, 15–20 (2005).
Betel, D., Wilson, M., Gabow, A., Marks, D.S. & Sander, C. The microRNA.org resource: targets and expression. Nucleic Acids Res. 36, D149–D153 (2008).
Boyle, E.I. et al. GO:TermFinder – open source software for accessing Gene Ontology information and finding significantly enriched Gene Ontology terms associated with a list of genes. Bioinformatics 20, 3710–3715 (2004).
Livak, K.J. & Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(−ΔΔC(T)) method. Methods 25, 402–408 (2001).
Gaffield, M.A. & Betz, W.J. Imaging synaptic vesicle exocytosis and endocytosis with FM dyes. Nat. Protoc. 1, 2916–2921 (2006).
Cousin, M.A. Use of FM1-43 and other derivatives to investigate neuronal function. Curr. Protoc. Neurosci. Ch. 2, unit 2.6 (2008).
Acknowledgements
We thank B. Cohen (McLean Hospital) and J. Madison (Broad Institute/Stanley Center) for providing the human fibroblasts; N. Nadif Kasri (Radbound University Medical Center Nijmegen) and A. Aschrafi (Radbound University Medical Center Nijmegen) for sharing the SpongeControl and SpongemiR-137 plasmids; T. Südhof (Stanford School of Medicine) for providing us the Syt1-KD and Syt1-KDControl plasmid37; K. Jones for the synaptosomal preparation protocol; W. Xu, J. Penney and M. Benevento for comments on the manuscript; and the members of the Tsai laboratory for their overall comments on the project. S.S. was supported by a Human Frontier Science Program (HFSP) long-term postdoctoral fellowship and a Swiss National Science Foundation fellowship for prospective researchers. E.J.K. was supported by a Simons Foundation Postdoctoral Fellowship. A.R. was supported by a NARSAD Young Investigator Award. This work was supported by a Seed Grant from the Simons Center for the Social Brain and US National Institutes of Health grant RO1 MH 091115 to L.-H.T.
Author information
Authors and Affiliations
Contributions
The study was designed by S.S., E.J.K. and L.-H.T., and directed and coordinated by L.-H.T. S.S. and L.-H.T. designed the experiments and wrote the manuscript. E.J.K. initiated the study and contributed to the preliminary data. J.S. and S.C. performed electrophysiological experiments. A.R. and W.W. performed behavior experiments. W.W. and E.J.K. cloned the miR-137OE and ΔmiR-137OE constructs. Z.F. and A.J.M. provided technical support. M.E. performed electron microscopy. A.E.M. contributed to the induced neuron experiments and helped write the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Human induced neurons.
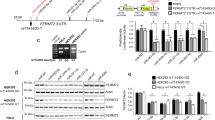
a. Transcriptional luciferase assay for the four SNPs in human neuroblastoma cell line SH-SY5Y, F(4) = 7.87, ***p < 0.001, posthoc analysis: tpGL3-rs2660304 (4.482), ***p < 0.001, r = 0.91; trs2660304-rs2802535 (–4.57), ***p < 0.001, r = 0.92; trs2660304-rs1625579 (–4.57), ***p < 0.001, r = 0.92; tpGL3-rs2660304 (1.65), nsp = 0.4781, r = 0.64, n: three experiments. b. Fold-change differences of MIR137 determined by qRT-PCR from Fig. 1c, separated by its individual lines, WiN= 65, *p = 0.0157, r = –0.40, box plot shows, in ascending order, the lowest maximum value, the first quartile, median, third quartile and the highest maximum value, dots are outlier. c. Summary table of the human fibroblasts and their genotype alleles for the four disease-associated SNPs. First four are homozygous for common major alleles; the last two are homozygous for the minor alleles. d. Examples of successfully induced neurons before FACS. Left: bright field, middle: transduced cells, right: expression of the neuron-specific HSV-CamKII-mCherry construct. Scale bar: 100 μm. Before any iN experiments, cells were visually controlled for successful reprogramming. e. Relative microRNA levels in induced neurons from common major and minor allele SNPs determined by RT-qPCR, tmiR-9(7.87) = –0.2, nsp = 0.8475, r = 0.07, tmiR-19b(8.28) = 0.12, nsp = 0.9047, r = 0.04, tmiR-124(13.89) = –0.16, nsp = 0.8773, r = 0.04, box plot shows, in ascending order, the lowest maximum value, the first quartile, median, third quartile and the highest maximum value, dots are outlier, n: number of samples from at least two reprogramming, s.e.m.
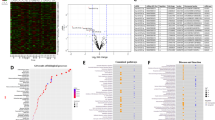
Supplementary Figure 2 Presynaptic target genes validation.
a. Summary table of putative in silico predicted miR-137 target genes involved in “synaptic transmission”. Red: targets tested in this study. b. Luciferase assay results for additional putative miR-137 targets, light grey and dark orange: miR-137 binding site deletion construct (Δ), WCalb1 = 170, *p = 0.0180, r = –0.43; tCamK2(14.58) = –0.14, nsp = 0.8888, r = 0.04; FNrxn1(3, 80) = 13.89, ***p < 0.001; FStx8(3, 56) = 0.60, nsp = 0.6159; WStxbp5 = 187, nsp = 0.4381, r = –0.13; FSv2a(3, 20) = 0.05, nsp = 0.9831; WSyn2a = 201, nsp = 0.6326, r = –0.07; FSyn2b(3, 74) = 0.72, nsp = 0.545; FSynj1(3, 56) = 1.30, nsp = 0.2845; WSynpr = 278, nsp = 0.1515, r = –0.22; WSyt9 = 121, nsp = 0.2983, r = –0.18; WVamp1 = 119, nsp = 0.1786 r = –0.22; FVamp2(3, 81) = 3.92, *p = 0.0115; WVamp7 = 172, nsp = 0.7637, r = –0.05. c. Example images of FM4–64 recording, white: induced neuron, green: FM4–64, arrow: terminal. ns: not significant, s.e.m.
Supplementary Figure 3 miR-137OE validation and in vivo expression.
a–b. Relative miR-137 expression level determined by qRT-PCR. a. in HEK-293T cells, Ty(2) = 44.89, ***p < 0.001, r = 1; b. in primary cortical neuronal culture at DIV14, Ty(3.97) = 4.48, *p = 0.0112, r = 0.91, n: number of experiments. c. Additional examples of hippocampi tile-scans injected with the lentivirus expressing the ΔmiR-137OE or miR-137OE constructs, immunostained for mCherry (magenta) and the nuclei dye DAPI (blue). CA3: Cornu Ammonis region 3. Staining was reliably throughout this study. d. Immunostaining of ΔmiR-137OE or miR-137OE-transduced dentate gyrus for DAPI (blue), mCherry (magenta), glial fibrillary acidic protein (GFAP), and ionized calcium-binding adapter molecule (Iba1). This experiment was repeated in at least three different animals. Scale bars: 100 μm. s.e.m.
Supplementary Figure 4 Ultrastructural analysis for miR-137OE and ΔmiR-137OE.
a. Ultrastructural images of the mossy fiber presynaptic terminals in ΔmiR-137OE and miR-137OE mice. Black arrow: mCherry gold-particle. Orange arrows: gap within the vesicle pool in miR-137OE synapses, * active zone, scale bar: 100 nm. b. Total number of vesicles for ΔmiR-137OE (black, n = 84) and miR-137OE (orange, n = 112), W = 4294, nsp = 0.2973, r = –0.07, box plot shows, in ascending order, the lowest maximum value, the first quartile, median, third quartile and the highest maximum value, dots are outlier, n: number of analyzed synapses of at least three animals. ns: not significant, s.e.m.
Supplementary Figure 5 Behavior analysis for miR-137OE and ΔmiR-137OE.
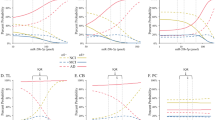
a–c. Equivalent levels of locomotion and a trend for anxiety in miR-137OE as measured by a. Open-field arena paradigm: margin distance: t(12.94) = –0.61, nsp = 0.5547, r = 0.17; margin time: t(12.20) = –1.89, nsp = 0.083, r = 0.48; center distance: t(13.83) = 2.55,*p = 0.023, r = 0.57; center time: t(12.20) = 1.89, nsp = 0.083, r = 0.48; total distance: t(12.89) = 1.31, nsp = 0.2117, r = 0.34; horizontal activity: t(13.52) = 1.98, nsp = 0.0685, r = 0.47; vertical activity: t(13.15) = 1.36, nsp = 0.1952, r = 0.35; stereotype: t(13.87) = 2.13, nsp = 0.05167, r = 0.50. b. Light-dark box paradigm: Frequency of exploration: t(9.90) = 0.94, nsp = 0.3709, r = 0.29, time spend in light: t(11.88) = 0.60, nsp = 0.5585, r = 0.17. c. Elevated-plus maze paradigm. Panel from left to right: time spent in closed or open arm, no significant main effect, F(1, 26) = 2.09, nsp = 0.161, tTime ΔmiR-137 (13.66) = 42.0, ***p < 0.001, r = 1, tTime miR-137 (9.86) = 39.68, ***p < 0.001, r = 1; Frequency of entering did not differ between open-closed arm and the condition, F(1, 26) = 0.40, nsp = 0.535, tFrequency ΔmiR-137 (11.49) = 2.49, *p < 0.05, r = 0.59, tFrequency miR-137 (11.49) = 2.49, *p < 0.05, r = 0.59). Distance and velocity, tdistance(12.98) = 0.36, nsp = 0.7259, r = 0.10, tvelocity(12.97) = 0.35, nsp = 0.7298, r = 0.10. d. Nociception response, W = 20.5, nsp = 0.7471, r = –0.09. e. Freezing response after context and before cue in %, t(41.97) = –1.20, nsp = 0.2381, r = 0.18. f. Thigmotaxis, percentage of the trail time and path that were spent in the defined thigmotaxis band (0.8) relative to the diameter of the pool, tTime(24.07) = –0.83, nsp = 0.4165, r = 0.17; tPath(25.57) = –0.15, nsp = 0.8838, r = 0.03. g. Three-chamber social interaction paradigm. Amount of time that the mouse spends to explore, left panel: new versus familiar mouse, F(2, 24) = 22.41, ***p < 0.001, right panel: mouse versus object, F(2, 24) = 45.5, ***p < 0.001. No significant difference for ΔmiR-137OE and miR-137OE in both experimental set-ups, Ffamiliar mouse (2, 48) = 2.41, nsp = 0.10 and Fempty cage (2, 48) = 2.86, nsp = 0.0673. Social paradigm for the familiar paradigm: for ΔmiR-137OE: tfamiliar-middle (–3.81), **p = 0.0023, tfamiliar-new (2.86), *p = 0.0225, tmiddle-new (6.67), ***p < 0.001; for miR137OE: tfamiliar-middle (–5.89), ***p < 0.001, tfamiliar-new (3.44), **p = 0.00574, tmiddle-new (9.33), ***p < 0.001; for the empty paradigm for ΔmiR-137OE: tempty-middle (–6.06), ***p < 0.001, tempty-mouse (3.35), **p = 0.0718, tmiddle-mouse (9.41), ***p < 0.001; for miR-137OE: tempty-middle (–4.03), **p = 0.0013, tempty-mouse (3.19), *p = 0.0106, tmiddle-mouse (7.22), ***p < 0.001, n: animals used for each experiment. ns: not significant, s.e.m.
Supplementary Figure 6 Sponge miR-137 and Sponge Control.
a. Additional ultrastructural images of the mossy fiber presynaptic terminals in SpongeControl and SpongemiR-137 mice. Black arrow: gold-particle labeled GFP, * active zone, scale bar: 100 nm. b. Total vesicle number for SpongeControl (black, n = 50) and SpongemiR-137 (blue, n = 43) mice in the presynaptic compartments at the mossy fiber synapse, t(88.82) = –1.80, nsp = 0.0751, r = –0.19, box plot shows, in ascending order, the lowest maximum value, the first quartile, median, third quartile and the highest maximum value, dots are outlier, n: number of analyzed synapses of at least three animals. c. Input-output curve estimated from fEPSP against the fiber volley amplitude at the mossy fiber-CA3 synapse in hippocampal slices from SpongeControl (black, n = 4) and SpongemiR-137 (blue, n = 4), F(1, 129) = 1.46, nsp = 0.229, shaded area: 95% confidence interval, n: number of analyzed hippocampal slices from at least three animals. d. Open-field arena paradigm. Total distance: t(16.5) = –0.72, nsp = 0.4809, r = 0.18; time moving: t(16.33) = –1.01, nsp = 0.3285, r = 0.24; moves/counts: t(17.51) = –0.88, nsp = 0.389, r = 0.21; distance periphery: t(16.37) = –0.18, nsp = 0.8617, r = 0.04; time periphery: t(16.19) = 2.62, *p = 0.0182, r = 0.55; distance center: t(12.89) = –2.80, *p = 0.0151, r = 0.62; time center: t(16.21) = –2.63, *p = 0.0182, r = 0.55. e. Nociception response, t(13.47) = 0.30, nsp = 0.7664, r = 0.08. f. Freezing response after context and before cue in %, t(17.12) = 0, nsp = 1, r = 0, n: number of animals. ns: not significant, s.e.m.
Supplementary Figure 7 Endogenous expression of synaptic target genes and Syt1 restoration (miR-137-Syt1-Venus).
a–c. Endogenous target gene expression in the dentate gyrus and the mossy fiber-CA3 pathway of naïve C57BL/6 animals. a. Absolute endogenous mRNA levels measured by RT-qPCR. b–c. Endogenous protein levels of Cplx1, Nsf, Syn3 and Syt1 by b. western blot (cropped. Full-length blots are presented in Supplementary Fig. 9), and c. immunohistochemistry. Vibratome sections, immunolabeled against the mossy fiber pathway marker zinc transporter 3 (ZnT3, green), the target gene (magenta), and stained with nuclei dye DAPI (blue). Scale bar: 10 μm. The observation was repeated in at least three different animals. d. Additional ultrastructural images of the mossy fiber presynaptic terminal of ΔmiR-137OE-Venus (grey), miR-137OE-Venus (red), miR-137OE-Syt1-Venus (green), and ΔmiR-137OE-Syt1-Venus (purple) mice, black arrow: gold particles staining of Venus, red arrows: gap within the vesicle pool in miR-137OE-Venus, * active zone, scale bar: 100 nm. e. Total vesicle number for ΔmiR-137OE-Venus (grey, n=56), miR-137OE-Venus (red, n=53), miR-137OE-Syt1-Venus (green, n=56), and ΔmiR-Syt1-Venus (purple, n=58), H(3) = 77.75, ***p < 0.001. Multiple comparison two-tailed test after Kruskal-Wallis, *p < 0.05, box plot shows, in ascending order, the lowest maximum value, the first quartile, median, third quartile and the highest maximum value, dots are outlier. f. Nociception response measured by mean speed of movement during shock in (cm s–1), H(3) = 6.10, nsp = 0.1068. ns: not significant, s.e.m.
Supplementary Figure 8 Syt1-KD and Syt1-KDControl in the DG-CA3 synapse and miR-137OE and ΔmiR-137OE at the entorhinal cortex-DG synapse.
a–c. Electrophysiological properties of Syt1 knockdown in acute hippocampal slices, black: Syt1-KDcontrol, brown: Syt1-shRNA. n: number of hippocampal slice preparation from at least 3 animal. a. LTP recording for Syt1-shRNA (n = 5) and Syt-1 control (n = 5), H(1) = 129,44 ***p < 0.001. Arrows: application of either high frequency stimulation or DCG-IV, embedded representative fEPSP traces, grey traces: baseline level before stimulation. Right bar chart: LTP magnitude calculated by averaging fEPSP amplitude during the last 10 min (50–60 min) recording, W = 2681, ***p < 0.001, r = –0.39. b. Input-output curve from fEPSP against the fiber volley amplitude at the mossy fiber-CA3 synapse from Syt1-shRNA (brown, n=5) and Syt-1 control (black, n=5), H(1) = 6.63, *p = 0.0100, shaded area: 95% confidence interval. c. fEPSP amplitude for 1 Hz sustained stimulation for Syt1-shRNA (n = 5) and Syt-1 control (black, n = 5), H(1) = 47.32, ***p < 0.001. Embedded: representative traces, grey traces: response to the 1st stimulation. d. BrdU incorporation in miR-137OE (orange) compared to ΔmiR-137OE (black) in the dentate gyrus, t(4.77) = –3.18, *p = 0.0264. Right panel: immunostaining for BrdU (green) and virus expression of mCherry (red), n: number of animals. b. LTP recording at the molecular layer of the dentate gyrus with stimulation of medial perfornant path in miR-137OE and control ΔmiR-137OE, H(1) = 3.19, nsp = 0.0741, arrows: application of stimulation paradigm, right bar chart: LTP magnitude during the last 10 min (50–60 min) recording, W = 1528, nsp = 0.9285, r = –0.01. c. Input-output curve estimated from fEPSP against the fiber volley amplitude measured at a range of stimulus intensities at the molecular layer of the dentate gyrus in hippocampal slices from ΔmiR-137OE (black, n=12) and miR-137OE (orange, n = 14), F(1,66) = 0.91, nsp = 0.4543, shaded area: 95% confidence interval. n: number of analyzed hippocampal slices from at least three animals, ns: not significant, s.e.m.
Supplementary Figure 9 Full-length western blot images.
Samples were run on a 4–15% TGX SDS-gradient gel (Bio-Rad Laboratories). Molecular weight (M, in kDa): BLUEstain 2 Protein ladder, 5–245 kDA (Goldbio). Images were taken with the infrared imaging system from Odyseey (Li-cor). This allows staining for two proteins at the same time on the same plot. Light-blue bands indicate oversaturation. a. for Fig. 2e, left and right panel show the same plot, but with adjustment of brightness and contrast; b. for Fig. 2d, the second and the fourth row are adjustment in brightness and contrast from the plots in the first and the third row, respectively; c. and d. for Supplementary Fig. 7. For Syt1, we always observed in varying intensity a band at ∼75 kDa. This band is likely to be unstripped Nsf. We tried to optimize the stripping procedure, however we could never remove the strong signal band for Nsf completely. For consistency, we decided to stain always Syt1 after Nsf.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–9 and Supplementary Table 1 (PDF 1566 kb)
Rights and permissions
About this article
Cite this article
Siegert, S., Seo, J., Kwon, E. et al. The schizophrenia risk gene product miR-137 alters presynaptic plasticity. Nat Neurosci 18, 1008–1016 (2015). https://doi.org/10.1038/nn.4023
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nn.4023
This article is cited by
-
Transition of allele-specific DNA hydroxymethylation at regulatory loci is associated with phenotypic variation in monozygotic twins discordant for psychiatric disorders
BMC Medicine (2023)
-
Mitochondrial, exosomal miR137-COX6A2 and gamma synchrony as biomarkers of parvalbumin interneurons, psychopathology, and neurocognition in schizophrenia
Molecular Psychiatry (2022)
-
Epigenome-wide DNA methylation analysis of whole blood cells derived from patients with GAD and OCD in the Chinese Han population
Translational Psychiatry (2022)
-
Circulating microRNA miR-137 as a stable biomarker for methamphetamine abstinence
Psychopharmacology (2022)
-
Regulation of the Glycine Transporter GLYT1 by microRNAs
Neurochemical Research (2022)