Abstract
The role of developmental transcription factors in maintenance of neuronal properties and in disease remains poorly understood. Lmx1a and Lmx1b are key transcription factors required for the early specification of ventral midbrain dopamine (mDA) neurons. Here we show that conditional ablation of Lmx1a and Lmx1b after mDA neuron specification resulted in abnormalities that show striking resemblance to early cellular abnormalities seen in Parkinson's disease. We found that Lmx1b was required for the normal execution of the autophagic-lysosomal pathway and for the integrity of dopaminergic nerve terminals and long-term mDA neuronal survival. Notably, human LMX1B expression was decreased in mDA neurons in brain tissue affected by Parkinson's disease. Thus, these results reveal a sustained and essential requirement of Lmx1b for the function of midbrain mDA neurons and suggest that its dysfunction is associated with Parkinson's disease pathogenesis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Björklund, A. & Dunnett, S.B. Dopamine neuron systems in the brain: an update. Trends Neurosci. 30, 194–202 (2007).
Cheng, H.C., Ulane, C.M. & Burke, R.E. Clinical progression in Parkinson disease and the neurobiology of axons. Ann. Neurol. 67, 715–725 (2010).
Deneris, E.S. & Hobert, O. Maintenance of postmitotic neuronal cell identity. Nat. Neurosci. 17, 899–907 (2014).
Holmberg, J. & Perlmann, T. Maintaining differentiated cellular identity. Nat. Rev. Genet. 13, 429–439 (2012).
Kadkhodaei, B. et al. Transcription factor Nurr1 maintains fiber integrity and nuclear-encoded mitochondrial gene expression in dopamine neurons. Proc. Natl. Acad. Sci. USA 110, 2360–2365 (2013).
Stott, S.R. et al. Foxa1 and foxa2 are required for the maintenance of dopaminergic properties in ventral midbrain neurons at late embryonic stages. J. Neurosci. 33, 8022–8034 (2013).
Di Salvio, M. et al. Otx2 controls neuron subtype identity in ventral tegmental area and antagonizes vulnerability to MPTP. Nat. Neurosci. 13, 1481–1488 (2010).
Bergman, O. et al. Do polymorphisms in transcription factors LMX1A and LMX1B influence the risk for Parkinson's disease? J. Neural Transm. 116, 333–338 (2009).
Bergman, O. et al. PITX3 polymorphism is associated with early onset Parkinson's disease. Neurobiol. Aging 31, 114–117 (2010).
Fuchs, J. et al. The transcription factor PITX3 is associated with sporadic Parkinson's disease. Neurobiol. Aging 30, 731–738 (2009).
Haubenberger, D. et al. Association of transcription factor polymorphisms PITX3 and EN1 with Parkinson's disease. Neurobiol. Aging 32, 302–307 (2011).
Le, W.D. et al. Mutations in NR4A2 associated with familial Parkinson disease. Nat. Genet. 33, 85–89 (2003).
Sleiman, P.M. et al. Characterisation of a novel NR4A2 mutation in Parkinson's disease brain. Neurosci. Lett. 457, 75–79 (2009).
Deng, Q. et al. Specific and integrated roles of Lmx1a, Lmx1b and Phox2a in ventral midbrain development. Development 138, 3399–3408 (2011).
Yan, C.H., Levesque, M., Claxton, S., Johnson, R.L. & Ang, S.L. Lmx1a and lmx1b function cooperatively to regulate proliferation, specification, and differentiation of midbrain dopaminergic progenitors. J. Neurosci. 31, 12413–12425 (2011).
Andersson, E. et al. Identification of intrinsic determinants of midbrain dopamine neurons. Cell 124, 393–405 (2006).
Chung, S. et al. Wnt1-lmx1a forms a novel autoregulatory loop and controls midbrain dopaminergic differentiation synergistically with the SHH-FoxA2 pathway. Cell Stem Cell 5, 646–658 (2009).
Friling, S. et al. Efficient production of mesencephalic dopamine neurons by Lmx1a expression in embryonic stem cells. Proc. Natl. Acad. Sci. USA 106, 7613–7618 (2009).
Sánchez-Danés, A. et al. Efficient generation of A9 midbrain dopaminergic neurons by lentiviral delivery of LMX1A in human embryonic stem cells and induced pluripotent stem cells. Hum. Gene Ther. 23, 56–69 (2012).
Asbreuk, C.H., Vogelaar, C.F., Hellemons, A., Smidt, M.P. & Burbach, J.P. CNS expression pattern of Lmx1b and coexpression with ptx genes suggest functional cooperativity in the development of forebrain motor control systems. Mol. Cell. Neurosci. 21, 410–420 (2002).
Dai, J.X., Hu, Z.L., Shi, M., Guo, C. & Ding, Y.Q. Postnatal ontogeny of the transcription factor Lmx1b in the mouse central nervous system. J. Comp. Neurol. 509, 341–355 (2008).
Zou, H.L. et al. Expression of the LIM-homeodomain gene Lmx1a in the postnatal mouse central nervous system. Brain Res. Bull. 78, 306–312 (2009).
Ekstrand, M.I. et al. Progressive parkinsonism in mice with respiratory-chain-deficient dopamine neurons. Proc. Natl. Acad. Sci. USA 104, 1325–1330 (2007).
Kadkhodaei, B. et al. Nurr1 is required for maintenance of maturing and adult midbrain dopamine neurons. J. Neurosci. 29, 15923–15932 (2009).
Hwang, D.Y. et al. 3,4-dihydroxyphenylalanine reverses the motor deficits in Pitx3-deficient aphakia mice: behavioral characterization of a novel genetic model of Parkinson's disease. J. Neurosci. 25, 2132–2137 (2005).
Lisman, J.E. & Grace, A.A. The hippocampal-VTA loop: controlling the entry of information into long-term memory. Neuron 46, 703–713 (2005).
Bliss, T.V. & Collingridge, G.L. A synaptic model of memory: long-term potentiation in the hippocampus. Nature 361, 31–39 (1993).
Friedman, L.G. et al. Disrupted autophagy leads to dopaminergic axon and dendrite degeneration and promotes presynaptic accumulation of alpha-synuclein and LRRK2 in the brain. J. Neurosci. 32, 7585–7593 (2012).
Hernandez, D. et al. Regulation of presynaptic neurotransmission by macroautophagy. Neuron 74, 277–284 (2012).
Bové, J., Martinez-Vicente, M. & Vila, M. Fighting neurodegeneration with rapamycin: mechanistic insights. Nat. Rev. Neurosci. 12, 437–452 (2011).
Dehay, B. et al. Pathogenic lysosomal depletion in Parkinson's disease. J. Neurosci. 30, 12535–12544 (2010).
Burke, R.E. & O'Malley, K. Axon degeneration in Parkinson's disease. Exp. Neurol. 246, 72–83 (2013).
Le, W. et al. Decreased NURR1 gene expression in patients with Parkinson's disease. J. Neurol. Sci. 273, 29–33 (2008).
Liu, H. et al. Decreased NURR1 and PITX3 gene expression in Chinese patients with Parkinson's disease. Eur. J. Neurol. 19, 870–875 (2012).
Chu, Y., Dodiya, H., Aebischer, P., Olanow, C.W. & Kordower, J.H. Alterations in lysosomal and proteasomal markers in Parkinson's disease: relationship to alpha-synuclein inclusions. Neurobiol. Dis. 35, 385–398 (2009).
Dehay, B. et al. Loss of P-type ATPase ATP13A2/PARK9 function induces general lysosomal deficiency and leads to Parkinson disease neurodegeneration. Proc. Natl. Acad. Sci. USA 109, 9611–9616 (2012).
Sánchez-Danés, A. et al. Disease-specific phenotypes in dopamine neurons from human iPS-based models of genetic and sporadic Parkinson's disease. EMBO Mol. Med. 4, 380–395 (2012).
Lynch-Day, M.A., Mao, K., Wang, K., Zhao, M. & Klionsky, D.J. The role of autophagy in Parkinson's disease. Cold Spring Harb. Perspect. Med. 2, a009357 (2012).
Decressac, M. et al. TFEB-mediated autophagy rescues midbrain dopamine neurons from alpha-synuclein toxicity. Proc. Natl. Acsad. Sci. USA 110, E1817–E1826 (2013).
Kanaan, N.M., Kordower, J.H. & Collier, T.J. Age-related accumulation of Marinesco bodies and lipofuscin in rhesus monkey midbrain dopamine neurons: relevance to selective neuronal vulnerability. J. Comp. Neurol. 502, 683–700 (2007).
Kaksonen, M. et al. Syndecan-3-deficient mice exhibit enhanced LTP and impaired hippocampus-dependent memory. Mol. Cell. Neurosci. 21, 158–172 (2002).
Migaud, M. et al. Enhanced long-term potentiation and impaired learning in mice with mutant postsynaptic density-95 protein. Nature 396, 433–439 (1998).
Vaillend, C., Billard, J.M. & Laroche, S. Impaired long-term spatial and recognition memory and enhanced CA1 hippocampal LTP in the dystrophin-deficient Dmd(mdx) mouse. Neurobiol. Dis. 17, 10–20 (2004).
Kordower, J.H. et al. Disease duration and the integrity of the nigrostriatal system in Parkinson's disease. Brain 136, 2419–2431 (2013).
Yang, Y., Coleman, M., Zhang, L., Zheng, X. & Yue, Z. Autophagy in axonal and dendritic degeneration. Trends Neurosci. 36, 418–428 (2013).
Taylor, T.N. et al. Nonmotor symptoms of Parkinson's disease revealed in an animal model with reduced monoamine storage capacity. J. Neurosci. 29, 8103–8113 (2009).
Doty, R.L. Olfaction in Parkinson's disease and related disorders. Neurobiol. Dis. 46, 527–552 (2012).
Engblom, D. et al. Glutamate receptors on dopamine neurons control the persistence of cocaine seeking. Neuron 59, 497–508 (2008).
Zhao, Z.Q. et al. Lmx1b is required for maintenance of central serotonergic neurons and mice lacking central serotonergic system exhibit normal locomotor activity. J. Neurosci. 26, 12781–12788 (2006).
Laguna, A. et al. The protein kinase DYRK1A regulates caspase-9-mediated apoptosis during retina development. Dev. Cell 15, 841–853 (2008).
Eriksson, T.M. et al. Bidirectional regulation of emotional memory by 5–HT1B receptors involves hippocampal p11. Mol. Psychiatry 18, 1096–1105 (2013).
Schintu, N. et al. PPAR-gamma-mediated neuroprotection in a chronic mouse model of Parkinson's disease. Eur. J. Neurosci. 29, 954–963 (2009).
Tillerson, J.L. et al. Olfactory discrimination deficits in mice lacking the dopamine transporter or the D2 dopamine receptor. Behav. Brain Res. 172, 97–105 (2006).
Deacon, R.M. & Rawlins, J.N. T-maze alternation in the rodent. Nat. Protoc. 1, 7–12 (2006).
Krishnan, V. et al. Molecular adaptations underlying susceptibility and resistance to social defeat in brain reward regions. Cell 131, 391–404 (2007).
Cryan, J.F. et al. Use of dopamine-beta-hydroxylase-deficient mice to determine the role of norepinephrine in the mechanism of action of antidepressant drugs. J. Pharmacol. Exp. Ther. 298, 651–657 (2001).
Yoshitake, T., Kehr, J., Todoroki, K., Nohta, H. & Yamaguchi, M. Derivatization chemistries for determination of serotonin, norepinephrine and dopamine in brain microdialysis samples by liquid chromatography with fluorescence detection. Biomed. Chromatogr. 20, 267–281 (2006).
Pruszak, J., Just, L., Isacson, O. & Nikkhah, G. Isolation and culture of ventral mesencephalic precursor cells and dopaminergic neurons from rodent brains. Curr. Protoc. Stem Cell Biol. Ch. 2, unit 2D 5 (2009).
Malewicz, M. et al. Essential role for DNA-PK-mediated phosphorylation of NR4A nuclear orphan receptors in DNA double-strand break repair. Genes Dev. 25, 2031–2040 (2011).
Beach, T.G. et al. The Sun Health Research Institute Brain Donation Program: description and experience, 1987–2007. Cell Tissue Bank 9, 229–245 (2008).
Acknowledgements
We thank T. Samuelsson, H. Lunden-Miguel and C.-Y. Leung for technical assistance, and members of the Ericson and Perlmann laboratories for discussions. We thank S.-L. Ang (National Institute for Medical Research, London), N.-G. Larsson (Max Planck Institute, Köln) and G. Schütz (DKFZ, Heidelberg) for providing mouse lines. We thank T. Beach at the Banner Sun Health Research Institute, Arizona, USA for providing postmortem human brain samples. This work was supported by funding from the European Union, Seventh Framework Programme under grant agreement mdDANeurodev, NeuroStemCell and Synsys (T.P., J.E. and O.S.), from the Swedish Strategic Research Foundation (SSF; T.P. and P.S.), from the Swedish Research Council, and from Hjärnfonden and Parkinsonfonden (O.S.). The human tissue donation program was supported by the US National Institutes of Health (U24 NS072026 and P30 AG19610), the Arizona Department of Health Services, the Arizona Biomedical Research Commission and the Michael J. Fox Foundation for Parkinson's Research. A.L. was supported by a Marie Curie Intra-European Fellowship for Career Development.
Author information
Authors and Affiliations
Contributions
A.L. planned all experiments, performed histological, gene expression and western blot analyses and wrote the manuscript; N.S. performed behavioral analysis and dissection of mouse brain tissue; A.N. performed primary ventral midbrain cultures and gene expression analysis; A.A. performed behavioral analysis; N.V. cloned and produced lentiviruses; J.K.J. performed stereological and western blot analyses; M.G.G. performed electrophysiological analysis; E.S. performed electron microscopy analysis; E.J. performed histological analysis; T.Y. and J.K. performed high-performance liquid chromatography; Q.D. and J.E. generated conditional Lmx1a gene targeted mice and helped with planning; P.S. helped with analysis and planning; O.S. performed electron microscopy analysis, helped with analysis and with writing the manuscript; T.P. together with A.L. planned all experiments and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
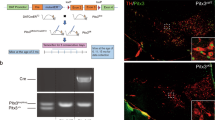
Supplementary Figure 1 Human LMX1A expression and recombination of Lmx1a and Lmx1b alleles in postmitotic DA neurons.
(a) Representative LMX1A immunostaining (blue; brown pigment corresponds to neuromelanin) in postmortem substantia nigra sections from human control subject. Scale bar, 25μm. (b) qRT-PCR analysis of Lmx1a and Lmx1b mRNA expression in dissected ventral midbrain region of E14.5 cCtrl and cDatCre embryos. n=5 embryos per genotype. Data is represented as mean±sem of the fold change normalized against Rpl19 expression and relative to cCtrl expression; Mann Whitney test *p<0.05. (c) Immunostaining in ventral midbrain sections of E16.5 cLmx1a/bCtrl (cCtrl) and cLmx1a/bDatCre (cDatCre) embryos showing Lmx1a, Lmx1b and tyrosine hydroxylase (TH) protein expression. Scale bar, 20μm. n=2 embryos per genotype.
Supplementary Figure 2 Behavioral characterization of adult and old cLmx1a/bCtrl and cLmx1a/bDatCre mice.
(a-e) Battery of behavioral tests performed with cLmx1a/bCtrl (cCtrl) and cLmx1a/bDatCre (cDatCre) mice. No differences in performance were revealed in anxiety or depression-like tests. (a) Passive avoidance test to evaluate emotional contextual memory. (b) T-maze test to measure spontaneous alternation. (c) Elevated plus-maze test to analyze anxiety like- behaviors. (d) Sucrose consumption test to evaluate anedonic-like behaviors. (e) Forced swin test to analyze depressive like-behaviors. Adult mice were aged 6 to 9 months and old mice 18 to 21 months. n=6–9 cCtrl and 8–14 cDatCre young animals, n=11–17 cCtrl and 11–18 cDatCre old animals; All data is represented as mean±sem; Two-way ANOVA was applied in (a), Unpaired t test was applied in (b-e); *p<0.05.
Supplementary Figure 3 Expression analysis of mDA neuron markers and catecholamines in Lmx1a-, Lmx1b- and Lmx1a/b-ablated mice.
(a) Tyrosine hydroxylase (TH) immunostaining in ventral midbrain sections of 2 months (2m) old cLmx1a/bCtrl (cCtrl) and cLmx1a/bDatCre (cDatCre) mice. Scale bar, 100 μm. (b) Unbiased stereological counting of TH-immunopositive cells (TH-positive) in the substantia nigra (SN) and the ventral tegmental area (VTA) of 2m old cCtrl and cDatCre mice. The sections were counterstained with Nissl staining and the Nissl-positive neuronal cells that were negative for TH immunostaining (Nissl) were counted. The total number of TH-positive and TH-negative neurons in the SN and VTA was also estimated (Total). Numbers correspond to the estimated cell numbers in one hemisphere. n=6 animals per genotype. (c) Tyrosine hydroxylase (TH) immunostaining in striatal sections of 2m old cLmx1aDatCre, cLmx1bDatCre mice and their corresponding control littermates (cLmx1aCtrl and cLmx1bCtrl). Scale bar, 100 μm. (d) TH immunostaining in ventral midbrain sections of 2m old cLmx1aDatCre, cLmx1bDatCre mice and their corresponding control littermates (cLmx1aCtrl and cLmx1bCtrl). Scale bar, 100 μm. (e) Optical densitometry from TH and Dopamine transporter (DAT) immunostained terminals from the dorsal striatum (Dorsal) and the ventral striatum (Ventral) areas of young (2m) cLmx1aDatCre, cLmx1bDatCre mice and their corresponding control littermates (cLmx1aCtrl and cLmx1bCtrl), n=3 animals per genotype. Data is shown in Arbitrary Units (AU). (f) TH immunostaining in striatal sections of 2m old cLmx1aDatCre, cLmx1bDatCre mice and their corresponding control littermates (cLmx1aCtrl and cLmx1bCtrl), showing nerve terminals and abnormally large profiles in cLmx1bDatCre mice. Scale bar, 50 μm. (g) HPLC measurements of Dopamine and the dopamine metabolites 3,4-dihydroxyphenylacetic acid (DOPAC) and Homovanillic acid (HVA) of 16 to 20 months old cLmx1aCtrl and cLmx1aDatCre mice. Separate analyses were performed on tissue extracts from the Striatum and the Nucleus accumbens (NAcc). n=10 cCtrl and 10 cDatCre animals per genotype. (h) HPLC measurements of Dopamine, DOPAC and HVA of 20 months old cLmx1bCtrl and cLmx1bDatCre mice. Separate analyses were performed on tissue extracts from the Striatum and the NAcc. n=10 cCtrl and 10 cDatCre animals per genotype. (i) HPLC measurements of Dopamine, DOPAC and HVA of 9 months old cLmx1a/bCtrl (cCtrl) and cLmx1a/bDatCre (cDatCre) mice. Separate analyses were performed on tissue extracts from the different brain areas indicated (Substantia nigra, SN; Ventral tegmental area, VTA; Cerebellum, CB). n=10 cCtrl and 13 cDatCre animals per genotype. All data is represented as mean±sem; Mann Whitney test; *p<0.05, **p<0.05, #p=0.06.
Supplementary Figure 4 Abnormal presynaptic nerve terminals in Lmx1a/b-ablated DA neurons.
(a) Vesicular monoamine transporter 2 (VMAT2) and Bassoon immunostaining in cLmx1a/bCtrl (cCtrl) and cLmx1a/bDatCre (cDatCre) mice. (b) Dopamine transporter (DAT) and Bassoon immunostaining in cCtrl and cDatCre mice. Representative pictures from n= 3 animals per genotype. Scale bar, 20μm. Merge images result from the combination of the green and red channel, and colocalization maps display colocalizing pixels as a white overlay on the green and red channel merge. (c) Optical densitometry from Bassoon immunostained active zones in the striatum of adult (12 months) cLmx1a/bCtrl (cCtrl) and cLmx1a/bDatCre (cDatCre) mice showing reduced occurrence of synaptic active zones in cDatCre mice. n=3 animals per genotype. Data is shown in Arbitrary Units (AU). (d) Pearson correlation coefficient from VMAT2 and Bassoon immunostained striatal sections of adult (12 months) cLmx1a/bCtrl (cCtrl) and cLmx1a/bDatCre (cDatCre) mice. n=3 animals per genotype. (e) Pearson correlation coefficient from DAT and Bassoon immunostained striatal sections of adult (12 months) cLmx1a/bCtrl (cCtrl) and cLmx1a/bDatCre (cDatCre) mice. n=3 animals per genotype. Both Pearson correlation coefficients reveal poor overlap of Bassoon staining with both dopaminergic markers. All data is represented as mean±sem; Mann Whitney test; *p<0.05.
Supplementary Figure 5 Phenotypic characterization of mice after Lmx1a, Lmx1b and Lmx1a/b ablation in mature mDA neurons.
(a) TH immunostaining in striatal sections of 2 months (2m) old cLmx1a/bDatCreERT2, cLmx1aDatCreERT2, cLmx1bDatCreERT2 mice and their corresponding control littermates (cLmx1a/bCtrl, cLmx1aCtrl and cLmx1bCtrl). All mice were treated with tamoxifen (TAM) at 4 weeks of age and analyzed 4 weeks later. cLmx1a/bDatCreERT2 and cLmx1bDatCreERT2 show abnormally large TH-immunopositive profiles throughout the striatum (arrows). Scale bar, 50 μm. n= 2 cCtrl and 4 cLmx1aDatCreERT2; n= 4 cCtrl and 7 cLmx1bDatCreERT2; n= 6 cCtrl and 6 cLmx1a/bDatCreERT2. (b-c) Quantification of the number of synaptic vesicles (sv) per µm of active zone (az) and of the number of multilamellar autophagic-lysosomal vesicles (ALV) per µm2 of the terminal profile in TH-immunostained striatal terminals larger than 1 µm from serial ultrathin sections of cLmx1a/bCtrl (cCtrl) and cLmx1a/bDatCreERT2 (cERT2) mice. n=5 synapses per genotype; n= 2 animals per genotype. (d-g) Battery of behavioral tests performed with cCtrl and cERT2 mice 6 months after TAM treatment. Beam traversal test (d) and Pole test (e) indicate no significant impairments of motor coordination and postural control. Open field test (f) shows no major alteration in locomotor activity. Novel object recognition test (g) shows no impairment in short-term memory formation since both groups show a significant preference for the novel object. Mice were aged 6 to 9 months. n=14 cCtrl and 16 cERT2 animals; All data is represented as mean±sem; Mann Whitney test was applied in (b-c), Unpaired t test was applied in (d-f), one sample t test was applied in (g); *p<0.05, **p<0.005, ***p<0.0005.
Supplementary Figure 6 Analysis of lysosome-mediated autophagy homeostasis after Lmx1a/b ablation and in response to rapamycin treatment.
(a) Quantification of protein levels by western blot analysis of autophagic and lysosomal proteins in striatal total cell extracts from cLmx1a/bCtrl (cCtrl) and cLmx1a/bDatCreERT2 (cDatCreERT2) mice 9 months after tamoxifen treatment. n=5 animals per genotype. All data is represented as mean±sem; Mann Whitney test *p<0.05. (b) Western blot of autophagic and lysosomal proteins in striatal total cell extracts from 9 months old cLmx1a/bCtrl and cLmx1a/bDatcre mice. n=4–5 animals per genotype. (c) Western blot of autophagic and lysosomal proteins in striatal total cell extracts from cLmx1a/bCtrl and cLmx1a/bDatcreERT2 mice 9 months after tamoxifen treatment. n=5 animals per genotype. (d) Western blot of autophagic and lysosomal proteins after pharmacological induction of autophagy and lysosomal function by rapamycin injection compared to vehicle injection (IP, 5mg/Kg/day for 10 consecutive days prior to sacrifice) in 2 months old cLmx1a/bCtrl (cCtrl) and cLmx1a/bDatCre (cDatCre) mice. n=5 animals per genotype. (e) Full-length western blots of phospho-mTOR (P-mTOR) and mTOR expression after pharmacological induction of autophagy and lysosomal function by rapamycin injection compared to vehicle injection in cCtrl and cDatCre mice corresponding to the cropped images shown in the main Figure 6e. * Note the presence of other bands corresponding to the signal for other proteins because the same membrane was blotted with other antibodies. Molecular weights (kDa) for the protein marker (Mk) are indicated.
Supplementary Figure 7 Analysis of Lmx1a/b-mediated transcriptional effects and gain- and loss-of-function experiments.
(a) qRT-PCR expression analysis of genes encoding for neuronal and dopaminergic markers, and for dopaminergic transcription factors in dissected ventral midbrain region of E14.5 cLmx1a/bCtrl (cCtrl) and cLmx1a/bDatCre (cDatCre) embryos. n=5 embryos per genotype. (b) Immunostaining showing Lmx1b and tyrosine hydroxylase (TH) protein expression in primary cultures from dissected ventral midbrains 6 days after lentiviral infection with lentiviruses expressing Lmx1b (LV-Lmx1b), a short-hairpin RNA with a Scrambled sequence (shScrmbl) or a sequence targeting Lmx1b (shLmx1b). Scale bar, 100μm. n= 2 independent experiments. (c) qRT-PCR expression analysis of Lmx1a and Lmx1b in primary cultures from dissected ventral midbrains 6 days after lentiviral infection with lentiviruses expressing GFP (LV-GFP) or LV-Lmx1b, and shScrmbl or shLmx1b. n= 2 biological duplicates in at least 3 independent experiments. Data is normalized to their corresponding controls (either LV-GFP or shScrmbl). (d) qRT-PCR expression analysis of Lmx1a, Lmx1b, genes encoding for autophagic-lysosomal proteins, genes involved in intracellular transport, general neuronal and dopaminergic genes in primary cultures from dissected ventral midbrains 6 days after lentiviral infection with lentiviruses expressing RFP (LV-RFP) or Lmx1a (LV-Lmx1a). n= 3 biological duplicates. Data is normalized to the corresponding control LV-RFP. All data is represented as mean±sem of the fold change normalized against Rpl19 levels; Mann Whitney test *p<0.05, *** p<0.0005.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Tables 1 and 2 (PDF 11239 kb)
Rights and permissions
About this article
Cite this article
Laguna, A., Schintu, N., Nobre, A. et al. Dopaminergic control of autophagic-lysosomal function implicates Lmx1b in Parkinson's disease. Nat Neurosci 18, 826–835 (2015). https://doi.org/10.1038/nn.4004
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nn.4004
This article is cited by
-
Animal models of Parkinson’s disease: bridging the gap between disease hallmarks and research questions
Translational Neurodegeneration (2023)
-
Genes critical for development and differentiation of dopaminergic neurons are downregulated in Parkinson’s disease
Journal of Neural Transmission (2023)
-
Maintenance of mitochondrial integrity in midbrain dopaminergic neurons governed by a conserved developmental transcription factor
Nature Communications (2022)
-
CCAAT/enhancer binding protein δ is a transcriptional repressor of α-synuclein
Cell Death & Differentiation (2020)
-
Myelin bodies in LMX1B-associated nephropathy: potential for misdiagnosis
Pediatric Nephrology (2020)