Abstract
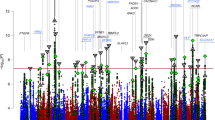
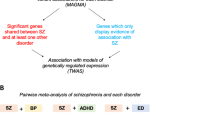
Genome-wide association studies (GWAS) of psychiatric disorders have identified multiple genetic associations with such disorders, but better methods are needed to derive the underlying biological mechanisms that these signals indicate. We sought to identify biological pathways in GWAS data from over 60,000 participants from the Psychiatric Genomics Consortium. We developed an analysis framework to rank pathways that requires only summary statistics. We combined this score across disorders to find common pathways across three adult psychiatric disorders: schizophrenia, major depression and bipolar disorder. Histone methylation processes showed the strongest association, and we also found statistically significant evidence for associations with multiple immune and neuronal signaling pathways and with the postsynaptic density. Our study indicates that risk variants for psychiatric disorders aggregate in particular biological pathways and that these pathways are frequently shared between disorders. Our results confirm known mechanisms and suggest several novel insights into the etiology of psychiatric disorders.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Change history
19 January 2015
In the version of this article initially published, the affiliation numbers given for several authors were incorrect. Astrid M. Vicente was listed as associated with affiliations 234–236 instead of 233–235, Veronica J. Vieland as 237 instead of 236, Kenneth S. Kendler as 110, 253 and 254 instead of 110, 252 and 253, and Zhaoming Zhao as 255 instead of 254 and 255. The errors have been corrected in the HTML and PDF versions of the article.
28 April 2015
In the version of this article initially published, the name of author Zhongming Zhao was misspelled Zhaoming Zhao. The error has been corrected in the HTML and PDF versions of the article.
References
Vos, T. et al. Years lived with disability (YLDs) for 1160 sequelae of 289 diseases and injuries 1990–2010: a systematic analysis for the Global Burden of Disease Study 2010. Lancet 380, 2163–2196 (2012).
Lee, S.H. et al. Genetic relationship between five psychiatric disorders estimated from genome-wide SNPs. Nat. Genet. 45, 984–994 (2013).
Cross-Disorder Group of the Psychiatric Genomics Consortium et al. Identification of risk loci with shared effects on five major psychiatric disorders: a genome-wide analysis. Lancet 381, 1371–1379 (2013).
Mirnics, K., Middleton, F.A., Marquez, A., Lewis, D.A. & Levitt, P. Molecular characterization of schizophrenia viewed by microarray analysis of gene expression in prefrontal cortex. Neuron 28, 53–67 (2000).
Nam, D. & Kim, S.Y. Gene-set approach for expression pattern analysis. Brief. Bioinform. 9, 189–197 (2008).
Ackermann, M. & Strimmer, K. A general modular framework for gene set enrichment analysis. BMC Bioinformatics 10, 47 (2009).
Baranzini, S.E. et al. Pathway and network-based analysis of genome-wide association studies in multiple sclerosis. Hum. Mol. Genet. 18, 2078–2090 (2009).
Wang, K. et al. Diverse genome-wide association studies associate the IL12/IL23 pathway with Crohn Disease. Am. J. Hum. Genet. 84, 399–405 (2009).
Holmans, P. et al. Gene ontology analysis of GWA study data sets provides insights into the biology of bipolar disorder. Am. J. Hum. Genet. 85, 13–24 (2009).
O'Dushlaine, C. et al. Molecular pathways involved in neuronal cell adhesion and membrane scaffolding contribute to schizophrenia and bipolar disorder susceptibility. Mol. Psychiatry 16, 286–292 (2011).
Psychiatric GWAS Consortium Bipolar Disorder Working Group. Large-scale genome-wide association analysis of bipolar disorder identifies a new susceptibility locus near ODZ4. Nat. Genet. 43, 977–983 (2011).
Sullivan, P.F. The psychiatric GWAS consortium: big science comes to psychiatry. Neuron 68, 182–186 (2010).
Neale, B.M. et al. Meta-analysis of genome-wide association studies of attention-deficit/hyperactivity disorder. J. Am. Acad. Child Adolesc. Psychiatry 49, 884–897 (2010).
Schizophrenia Psychiatric Genome-Wide Association Study Consortium. Genome-wide association study identifies five new schizophrenia loci. Nat. Genet. 43, 969–976 (2011).
Major Depressive Disorder Working Group of the PGC et al. A mega-analysis of genome-wide association studies for major depressive disorder. Mol. Psychiatry 18, 497–511 (2013).
Brown, M.B. A method for combining non-independent one-sided tests of significance. Biometrics 31, 987–992 (1975).
McLaren, P.J. et al. Association study of common genetic variants and HIV-1 acquisition in 6,300 infected cases and 7,200 controls. PLoS Pathog. 9, e1003515 (2013).
Kang, H.J. et al. Spatio-temporal transcriptome of the human brain. Nature 478, 483–489 (2011).
Gong, S. et al. A gene expression atlas of the central nervous system based on bacterial artificial chromosomes. Nature 425, 917–925 (2003).
Xu, X., Wells, A.B., O′Brien, D.R., Nehorai, A. & Dougherty, J.D. Cell type-specific expression analysis to identify putative cellular mechanisms for neurogenetic disorders. J. Neurosci. 34, 1420–1431 (2014).
Doyle, J.P. et al. Application of a translational profiling approach for the comparative analysis of CNS cell types. Cell 135, 749–762 (2008).
Schizophrenia Working Group of the Psychiatric Genomics. C. Biological insights from 108 schizophrenia-associated genetic loci. Nature 511, 421–427 (2014).
Kirov, G. et al. De novo CNV analysis implicates specific abnormalities of postsynaptic signalling complexes in the pathogenesis of schizophrenia. Mol. Psychiatry 17, 142–153 (2012).
Purcell, S.M. et al. A polygenic burden of rare disruptive mutations in schizophrenia. Nature 506, 185–190 (2014).
Voineagu, I. et al. Transcriptomic analysis of autistic brain reveals convergent molecular pathology. Nature 474, 380–384 (2011).
Ting, J.T., Peça, J. & Feng, G. Functional consequences of mutations in postsynaptic scaffolding proteins and relevance to psychiatric disorders. Annu. Rev. Neurosci. 35, 49–71 (2012).
Purcell, S.M. et al. A polygenic burden of rare disruptive mutations in schizophrenia. Nature 506, 185–190 (2014).
Fromer, M. et al. De novo mutations in schizophrenia implicate synaptic networks. Nature 506, 179–184 (2014).
Cruceanu, C. et al. H3K4 tri-methylation in synapsin genes leads to different expression patterns in bipolar disorder and major depression. Int. J. Neuropsychopharmacol. 16, 289–99 (2013).
Jarome, T.J. & Lubin, F.D. Histone lysine methylation: critical regulator of memory and behavior. Rev. Neurosci. 24, 375–387 (2013).
Ben-David, E. & Shifman, S. Combined analysis of exome sequencing points toward a major role for transcription regulation during brain development in autism. Mol. Psychiatry 18, 1054–1056 (2013).
Parikshak, N.N. et al. Integrative functional genomic analyses implicate specific molecular pathways and circuits in autism. Cell 155, 1008–1021 (2013).
Ronemus, M., Iossifov, I., Levy, D. & Wigler, M. The role of de novo mutations in the genetics of autism spectrum disorders. Nat. Rev. Genet. 15, 133–141 (2014).
Bauman, A.L. et al. Cocaine and antidepressant-sensitive biogenic amine transporters exist in regulated complexes with protein phosphatase 2A. J. Neurosci. 20, 7571–7578 (2000).
Dieperink, E., Willenbring, M. & Ho, S.B. Neuropsychiatric symptoms associated with hepatitis C and interferon alpha: a review. Am. J. Psychiatry 157, 867–876 (2000).
Susser, E., St Clair, D. & He, L. Latent effects of prenatal malnutrition on adult health: the example of schizophrenia. Ann. NY Acad. Sci. 1136, 185–192 (2008).
Heijmans, B.T., Tobi, E.W., Lumey, L.H. & Slagboom, P.E. The epigenome: archive of the prenatal environment. Epigenetics 4, 526–531 (2009).
Brykczynska, U. et al. Repressive and active histone methylation mark distinct promoters in human and mouse spermatozoa. Nat. Struct. Mol. Biol. 17, 679–687 (2010).
Huang, H.S. et al. Prefrontal dysfunction in schizophrenia involves mixed-lineage leukemia 1-regulated histone methylation at GABAergic gene promoters. J. Neurosci. 27, 11254–62 (2007).
Yauch, R.L. & Settleman, J. Recent advances in pathway-targeted cancer drug therapies emerging from cancer genome analysis. Curr. Opin. Genet. Dev. 22, 45–49 (2012).
Flicek, P. et al. Ensembl 2013. Nucleic Acids Res. 41, D48–D55 (2013).
Maston, G.A., Evans, S.K. & Green, M.R. Transcriptional regulatory elements in the human genome. Annu. Rev. Genomics Hum. Genet. 7, 29–59 (2006).
Pedroso, I. et al. Common genetic variants and gene-expression changes associated with bipolar disorder are over-represented in brain signaling pathway genes. Biol. Psychiatry 72, 311–317 (2012).
Moskvina, V. et al. Evaluation of an approximation method for assessment of overall significance of multiple-dependent tests in a genomewide association study. Genet. Epidemiol. 35, 861–866 (2011).
Lee, P.H., O'Dushlaine, C., Thomas, B. & Purcell, S.M. INRICH: interval-based enrichment analysis for genome-wide association studies. Bioinformatics 28, 1797–1799 (2012).
Segrè, A.V., Groop, L., Mootha, V.K., Daly, M.J. & Altshuler, D. Common inherited variation in mitochondrial genes is not enriched for associations with type 2 diabetes or related glycemic traits. PLoS Genet. 6, e1001058 (2010).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Storey, J.D. & Tibshirani, R. Statistical significance for genomewide studies. Proc. Natl. Acad. Sci. USA 100, 9440–9445 (2003).
Brown, M.B. A method for combining non-independent, one-sided tests of significance. Biometrics 31, 978–992 (1975).
Zhang, B. & Horvath, S. A general framework for weighted gene co-expression network analysis. Stat. Appl. Genet. Mol. Biol. 4, Article17 (2005).
Acknowledgements
G.B. and S.N. acknowledge funding support for this work from the National Institute for Health Research (NIHR) Mental Health Biomedical Research Centre at South London and Maudsley NHS Foundation Trust and King's College London. P.H.L. is supported by US National Institute of Mental Health (NIMH) grant K99MH101367. The PGC Cross-Disorder Group is supported by NIMH grant U01 MH085520. Statistical analyses were carried out on the Genetic Cluster Computer, which is financially supported by the Netherlands Scientific Organization (NOW; 480-05-003; principal investigator D.P.) along with a supplement from the Dutch Brain Foundation and VU University. Numerous (>100) grants from government agencies along with substantial private and foundation support worldwide enabled the collection of phenotype and genotype data, without which this research would not be possible.
Author information
Authors and Affiliations
Consortia
Contributions
Project conception: G.B., P.F.S., P.A.H. Analysis: C.O'D., P.H.L., P.A.H., G.B., S.R., L.R., L.D., S.N. Writing of the manuscript: G.B., C.O'D. P.A.H., P.H.L., L.R., L.D., N.P. Quality control for PGC data: S. Ripke and B.M.N. Revisions to the manuscript: G.B., D.H.G., C.O'D., L.R., P.H.L., N.P. PGC Network and Pathway Analysis Workgroup: S.R., B.M.N., S.M.P., D.H.G., P.A.H., P.H.L., M.M., C.O'D., D.P., L.R., L.D., P.F.S., J.W.S., N.R.W., Z.Z. PGC Workgroup Chairs: M.J.D. (analysis), S.V.F. (ADHD), M.J.D. and B.D. (co-chairs ASD), G.B. and P.A.H. (Network and Pathway Analysis subgroup), J.K. and P. Sklar (co-chairs bipolar disorder), P.F.S. (major depressive disorder), M.C.O'D. (schizophrenia) and J.W.S. and K.S.K. (co-chairs cross-disorder group). Collection, genotyping and analysis for PGC Working Groups. PGC ADHD Working Group: B.M.N., S.V.F., A.T., R.A., P.A., T. Banaschewski, M. Bayés, J.B., J.K.B., M.C., B.C., J.C., A.E.D., R.P.E., J.E., B.F., C.M.F., L. Kent, J.K., K.-P.L., S.K.L., J.M., J.J.M., S.E.M., J.M.S., A. Miranda, S.F.N., R.D.O., J.A.R.-Q., A. Reif, M. Ribasés, H.R., A. Rothenberger, J.A.S., R.S., S.L. Smalley, E.J.S.S.-B., H.-C.S., A.A.T. and N.W. PGC ASD Working Group: R.A., D.E.A., A.J.B., A.B., C.B., J.D. Buxbaum, A. Chakravarti, E.H.C., H.C., M.L.C., G.D., E.D., S.E., E.F., C.M.F., L. Gallagher, D.H.G., M. Gill, D.E.G., J.L.H., H.H., J.H., V.H., S.M.K., L. Klei, D.H. Ledbetter, C. Lord, J.K.L., E.M., S.M.M., C.L.M., W.M.M., A.P.M., D.M.-D.-L., E.M.M., M. Murtha, G.O., A.P., J.R.P., A.D.P., M.A.P.-V., J. Piven, F.P., K. Rehnström, K. Roeder, G.R., S.J.S., S.L. Santangelo, G.D.S., S.W.S., M. State, J.S. Sutcliffe, P. Szatmari, A.M.V., V.J.V., C.A.W., T.H.W., E.M.W., A.J.W., T.W.Y., B.D. and M.J.D. PGC BPD Working Group: S.M.P., D.A., H.A., O.A.A., A.A., L.B., J.A.B., J.D. Barchas, T.B.B., N.B., M. Bauer, F.B., S.E.B., W.B., D.H.R.B., C.S.B., M. Boehnke, G.B., R. Breuer, W.E.B., W.F.B., S. Caesar, K. Chambert, S. Cichon, D.A.C., A. Corvin, W.H.C., D.W.C., R.D., F. Degenhardt, S. Djurovic, F. Dudbridge, H.J.E., B.E., A.E.F., I.N.F., M. Flickinger, T.F., J.F., C.F., L.F., E.S.G., M. Gill, K.G.-S., E.K.G., T.A.G., D.G., W.G., H.G., M.L.H., M. Hautzinger, S. Herms, M. Hipolito, P.A.H., C.M.H., S.J., E.G.J., I.J., L.J., R. Kandaswamy, J.L.K., G.K.K., D.L.K., P.K., M. Landén, N.L., M. Lathrop, J. Lawrence, W.B.L., M. Leboyer, P.H.L., J. Li, P.L., D.-Y.L., C. Liu, F.W.L., S.L., P.B.M., W.M., N.G.M., M. Mattheisen, K.M., M. Mattingsdal, K.A.M., P.M., M.G.M., A. McIntosh, R.M., A.W.M., F.J.M., A. McQuillin, S.M., I.M., F.M., G.W.M., J.L.M., G.M., D.W.M., V. Moskvina, P.M., T.W.M., W.J.M., B.M.-M., R.M.M., C.M.N., I.N., V.N., M.M.N., J.I.N., E.A.N., C.O., U.O., M.J.O., B.S.P., J.B.P., P.P., E.M.Q., S. Raychaudhuri, A. Reif, J.P.R., M. Rietschel, D. Ruderfer, M. Schalling, A.F.S., W.A.S., N.J.S., T.G.S., J. Schumacher, M. Schwarz, E.S., L.J.S., P.D.S., E.N.S., D.S.C., M. Steffens, J.S. Strauss, J. Strohmaier, S.S., R.C.T., F.T., J.T., J.B.V., S.J.W., T.F.W., S.H.W., W.X., A.H.Y., P.P.Z., P.Z., S. Zöllner, J.R.K., P. Sklar, M.J.D., M.C.O. and N.C. PGC MDD Working Group: M.R.B., T. Bettecken, E.B.B., D.H.R.B., D.I.B., G.B., R. Breuer, S. Cichon, W.H.C., I.W.C., D. Czamara, E.J.D.G., F. Degenhardt, A.E.F., J.F., S.D.G., M. Gross, S.P.H., A.C.H., A.K.H., S. Herms, I.B.H., F.H., W.J.H., S. Hoefels, J.-J.H., M.I., I.J., L.J., J.-Y.T., J.A.K., M.A.K., A.K., W.B.L., D.F.L., C.M.L., D.-Y.L., S.L., D.J.M., P.A.F.M., W.M., N.G.M., M. Mattheisen, P.J.M., P.M., A. McIntosh, A.W.M., C.M.M., L.M., G.W.M., P.M., B.M.-M., W.A.N., M.M.N., D.R.N., B.W.P., M.L.P., J.B.P., M. Rietschel, W.A.S., T.G.S., J. Shi, S.I.S., S.L. Slager, J.H.S., M. Steffens, F.T., J.T., M.U., E.J.C.G.v.d.O., G.V.G., M.M.W., G.W., F.G.Z., P.F.S. and N.R.W. PGC SCZ Working Group: S. Ripke, B.M.N., S.M.P., B.J.M., I.A., F.A., O.A.A., M.H.A., N.B., D.W.B., D.H.R.B., R. Bruggeman, N.G.B., W.F.B., W.C., R.M.C., K. Choudhury, S. Cichon, C.R.C., P.C., A. Corvin, D. Curtis, S. Datta, S. Djurovic, G.J.D., J.D., F. Dudbridge, A.F., R.F., N.B.F., M. Friedl, P.V.G., L. Georgieva, I.G., M. Gill, H.G., L.D.H., M.L.H., T.F.H., A.M.H., P.A.H., C.M.H., A.I., A.K.K., R.S.K., M.C.K., E.K., Y.K., G.K.K., B.K., L. Krabbendam, R. Krasucki, J. Lawrence, P.H.L., T.L., D.F.L., J.A.L., D.-Y.L., D.H. Linszen, P.K.E.M., W.M., A.K.M., M. Mattheisen, M. Mattingsdal, S.M., S.A.M., A. McIntosh, A. McQuillin, H.M., I.M., V. Milanova, D.W.M., V. Moskvina, I.M.-G., M.M.N., C.O., A.O., L.O., R.A.O., M.J.O., C.N.P., M.T.P., B.S.P., J. Pimm, D.P., V.P., D.J.Q., H.B.R., M. Rietschel, L.R., D. Ruderfer, D. Rujescu, A.R.S., T.G.S., J. Shi, J.M.S., D.S.C., T.S.S., S.T., J.V.O., P.M.V., T.W., S. Zammit, P. Sklar, M.J.D., M.C.O., N.C., P.F.S. and K.S.K. PGC Cross-Disorder Group Working Group: S.H.L., S. Ripke, B.M.N., S.M.P., R.H.P., A.T., A.F., M.C.N., J.I.N., B.W.P., M. Rietschel, T.G.S., N.C., S.L. Santangelo, P.F.S., J.W.S., K.S.K. and N.R.W. PGC Analysis Working Group: S.H.L., S. Ripke, B.M.N., S.M.P., V.A., E.M.B., P.H.L., S.E.M., M.C.N., D.P., G.B., M.J.D. and N.R.W.
Corresponding authors
Ethics declarations
Competing interests
G.B. is a consultant in PreClinical Genetics at Eli Lilly.
Integrated supplementary information
Supplementary Figure 1 Observed and expected (shown as ranges representing 95% confidence intervals) number of significant pathways overlapping between different sets of results from different methods.
(a) Analysis conducted using the SCZ dataset, top 25% of each method. (b) Analysis conducted using the NULL dataset, top 25% of each method.
Supplementary Figure 2 Quantile-quantile plots summarizing combined p-values for each method on different phenotypes.
Order is (a) SCZ, (b) BIP, (c) MDD, (d) AUT, (e) ADD, (f) null data from permutations and, (g) HIV.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1 and 2, Supplementary Analysis, and Supplementary Tables 1–4, 6 and 12–16 (PDF 280 kb)
Supplementary Table 5
Pathway results for each disease/cross-disease dataset (XLS 21847 kb)
Supplementary Table 7
Ranked list of pathways when combined across 3 well-powered disease groups. (XLS 3804 kb)
Supplementary Table 8
Ranked list of pathways when combined across SCZ, HIV and NULL datasets. (XLS 3803 kb)
Supplementary Table 9
Details related to module-level summaries of spatial, temporal, and cell-type specific expression patterns. (XLSX 184 kb)
Supplementary Table 10
Ranked list of pathways when combined across all 5 disease groups. (XLS 3959 kb)
Supplementary Table 11
P values for the SNPs in each gene in each disorder and their pathways membership are presented for all pathways with q < 0.1. (XLSX 639 kb)
Supplementary Table 17
Information related to the supervised network analysis of 797 genes, specifying gene set membership from the 3 major pathways, module membership, and relative connectivity (kME) to each module. (XLSX 171 kb)
Supplementary Data
Source data for Figure 3 (ZIP 1200 kb)
Supplementary Software
PGC pathway network analysis code (TXT 22 kb)
Rights and permissions
About this article
Cite this article
The Network and Pathway Analysis Subgroup of the Psychiatric Genomics Consortium. Psychiatric genome-wide association study analyses implicate neuronal, immune and histone pathways. Nat Neurosci 18, 199–209 (2015). https://doi.org/10.1038/nn.3922
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nn.3922
This article is cited by
-
Systemic inflammatory biomarkers in Schizophrenia are changed by ECT administration and related to the treatment efficacy
BMC Psychiatry (2024)
-
CSF1R regulates schizophrenia-related stress response and vascular association of microglia/macrophages
BMC Medicine (2023)
-
Prioritizing genes associated with brain disorders by leveraging enhancer-promoter interactions in diverse neural cells and tissues
Genome Medicine (2023)
-
Impulsivity across severe mental disorders: a cross-sectional study of immune markers and psychopharmacotherapy
BMC Psychiatry (2023)
-
Inflammation and cognition in severe mental illness: patterns of covariation and subgroups
Molecular Psychiatry (2023)