Abstract
Macroautophagy (hereafter autophagy) is a key pathway in neurodegeneration. Despite protective actions, autophagy may contribute to neuron demise when dysregulated. Here we consider X-linked spinal and bulbar muscular atrophy (SBMA), a repeat disorder caused by polyglutamine-expanded androgen receptor (polyQ-AR). We found that polyQ-AR reduced long-term protein turnover and impaired autophagic flux in motor neuron–like cells. Ultrastructural analysis of SBMA mice revealed a block in autophagy pathway progression. We examined the transcriptional regulation of autophagy and observed a functionally significant physical interaction between transcription factor EB (TFEB) and AR. Normal AR promoted, but polyQ-AR interfered with, TFEB transactivation. To evaluate physiological relevance, we reprogrammed patient fibroblasts to induced pluripotent stem cells and then to neuronal precursor cells (NPCs). We compared multiple SBMA NPC lines and documented the metabolic and autophagic flux defects that could be rescued by TFEB. Our results indicate that polyQ-AR diminishes TFEB function to impair autophagy and promote SBMA pathogenesis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Malgaroli, A., Vallar, L. & Zimarino, V. Protein homeostasis in neurons and its pathological alterations. Curr. Opin. Neurobiol. 16, 270–274 (2006).
Taylor, J.P., Hardy, J. & Fischbeck, K.H. Toxic proteins in neurodegenerative disease. Science 296, 1991–1995 (2002).
Levine, B. & Kroemer, G. Autophagy in the pathogenesis of disease. Cell 132, 27–42 (2008).
Hara, T. et al. Suppression of basal autophagy in neural cells causes neurodegenerative disease in mice. Nature 441, 885–889 (2006).
Komatsu, M. et al. Loss of autophagy in the central nervous system causes neurodegeneration in mice. Nature 441, 880–884 (2006).
Komatsu, M. et al. Essential role for autophagy protein Atg7 in the maintenance of axonal homeostasis and the prevention of axonal degeneration. Proc. Natl. Acad. Sci. USA 104, 14489–14494 (2007).
Ravikumar, B. et al. Inhibition of mTOR induces autophagy and reduces toxicity of polyglutamine expansions in fly and mouse models of Huntington disease. Nat. Genet. 36, 585–595 (2004).
Shintani, T. & Klionsky, D.J. Autophagy in health and disease: a double-edged sword. Science 306, 990–995 (2004).
Klionsky, D.J. Autophagy: from phenomenology to molecular understanding in less than a decade. Nat. Rev. Mol. Cell Biol. 8, 931–937 (2007).
Pandey, U.B. et al. HDAC6 rescues neurodegeneration and provides an essential link between autophagy and the UPS. Nature 447, 859–863 (2007).
Mizushima, N., Yamamoto, A., Matsui, M., Yoshimori, T. & Ohsumi, Y. In vivo analysis of autophagy in response to nutrient starvation using transgenic mice expressing a fluorescent autophagosome marker. Mol. Biol. Cell 15, 1101–1111 (2004).
Boland, B. et al. Autophagy induction and autophagosome clearance in neurons: relationship to autophagic pathology in Alzheimer's disease. J. Neurosci. 28, 6926–6937 (2008).
Nixon, R.A. et al. Extensive involvement of autophagy in Alzheimer disease: an immuno-electron microscopy study. J. Neuropathol. Exp. Neurol. 64, 113–122 (2005).
Wong, E. & Cuervo, A.M. Autophagy gone awry in neurodegenerative diseases. Nat. Neurosci. 13, 805–811 (2010).
Sardiello, M. et al. A gene network regulating lysosomal biogenesis and function. Science 325, 473–477 (2009).
Settembre, C. et al. TFEB links autophagy to lysosomal biogenesis. Science 332, 1429–1433 (2011).
Tsunemi, T. et al. PGC-1alpha rescues huntington's disease proteotoxicity by preventing oxidative stress and promoting TFEB function. Sci. Transl. Med. 4, 142ra197 (2012).
Decressac, M. et al. TFEB-mediated autophagy rescues midbrain dopamine neurons from alpha-synuclein toxicity. Proc. Natl. Acad. Sci. USA 110, E1817–E1826 (2013).
Dejager, S. et al. A comprehensive endocrine description of Kennedy's disease revealing androgen insensitivity linked to CAG repeat length. J. Clin. Endocrinol. Metab. 87, 3893–3901 (2002).
La Spada, A.R., Wilson, E.M., Lubahn, D.B., Harding, A.E. & Fischbeck, K.H. Androgen receptor gene mutations in X-linked spinal and bulbar muscular atrophy. Nature 352, 77–79 (1991).
La Spada, A.R. & Taylor, J.P. Repeat expansion disease: progress and puzzles in disease pathogenesis. Nat. Rev. Genet. 11, 247–258 (2010).
Cary, G.A. & La Spada, A.R. Androgen receptor function in motor neuron survival and degeneration. Phys. Med. Rehabil. Clin. N. Am. 19, 479–494 viii (2008).
Thomas, P.S. Jr. et al. Loss of endogenous androgen receptor protein accelerates motor neuron degeneration and accentuates androgen insensitivity in a mouse model of X-linked spinal and bulbar muscular atrophy. Hum. Mol. Genet. 15, 2225–2238 (2006).
Klionsky, D.J. et al. Guidelines for the use and interpretation of assays for monitoring autophagy. Autophagy 8, 445–544 (2012).
Sopher, B.L. et al. Androgen receptor YAC transgenic mice recapitulate SBMA motor neuronopathy and implicate VEGF164 in the motor neuron degeneration. Neuron 41, 687–699 (2004).
Chua, J.P. et al. Transcriptional activation of TFEB/ZKSCAN3 target genes underlies enhanced autophagy in spinobulbar muscular atrophy. Hum. Mol. Genet. 23, 1376–1386 (2014).
Martina, J.A., Chen, Y., Gucek, M. & Puertollano, R. MTORC1 functions as a transcriptional regulator of autophagy by preventing nuclear transport of TFEB. Autophagy 8, 903–914 (2012).
Takahashi, K. & Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126, 663–676 (2006).
Marchetto, M.C. et al. A model for neural development and treatment of Rett syndrome using human induced pluripotent stem cells. Cell 143, 527–539 (2010).
Ranganathan, S. et al. Mitochondrial abnormalities in spinal and bulbar muscular atrophy. Hum. Mol. Genet. 18, 27–42 (2009).
Boulting, G.L. et al. A functionally characterized test set of human induced pluripotent stem cells. Nat. Biotechnol. 29, 279–286 (2011).
Lieberman, A.P., Harmison, G., Strand, A.D., Olson, J.M. & Fischbeck, K.H. Altered transcriptional regulation in cells expressing the expanded polyglutamine androgen receptor. Hum. Mol. Genet. 11, 1967–1976 (2002).
McCampbell, A. et al. CREB-binding protein sequestration by expanded polyglutamine. Hum. Mol. Genet. 9, 2197–2202 (2000).
Miller, A.J., Levy, C., Davis, I.J., Razin, E. & Fisher, D.E. Sumoylation of MITF and its related family members TFE3 and TFEB. J. Biol. Chem. 280, 146–155 (2005).
He, B., Kemppainen, J.A., Voegel, J.J., Gronemeyer, H. & Wilson, E.M. Activation function 2 2 in the human androgen receptor ligand binding domain mediates interdomain communication with the NH2-terminal domain. J. Biol. Chem. 274, 37219–37225 (1999).
Lam, Y.C. et al. ATAXIN-1 interacts with the repressor Capicua in its native complex to cause SCA1 neuropathology. Cell 127, 1335–1347 (2006).
Nedelsky, N.B. et al. Native functions of the androgen receptor are essential to pathogenesis in a Drosophila model of spinobulbar muscular atrophy. Neuron 67, 936–952 (2010).
Young, J.E. et al. Polyglutamine-expanded androgen receptor truncation fragments activate a Bax-dependent apoptotic cascade mediated by DP5/Hrk. J. Neurosci. 29, 1987–1997 (2009).
Doi, H. et al. p62/SQSTM1 differentially removes the toxic mutant androgen receptor via autophagy and inclusion formation in a spinal and bulbar muscular atrophy mouse model. J. Neurosci. 33, 7710–7727 (2013).
Bonaldo, P. & Sandri, M. Cellular and molecular mechanisms of muscle atrophy. Dis. Model. Mech. 6, 25–39 (2013).
Cortes, C.J. et al. Muscle expression of mutant androgen receptor accounts for systemic and motor neuron disease phenotypes in spinal and bulbar muscular atrophy. Neuron 82, 295–307 (2014).
Marchetto, M.C., Brennand, K.J., Boyer, L.F. & Gage, F.H. Induced pluripotent stem cells (iPSCs) and neurological disease modeling: progress and promises. Hum. Mol. Genet. 20, R109–R115 (2011).
HD iPSC Consortium. Induced pluripotent stem cells from patients with Huntington's disease show CAG-repeat-expansion-associated phenotypes. Cell Stem Cell 11, 264–278 (2012).
Nihei, Y. et al. Enhanced aggregation of androgen receptor in induced pluripotent stem cell-derived neurons from spinal and bulbar muscular atrophy. J. Biol. Chem. 288, 8043–8052 (2013).
Sánchez-Danes, A. et al. Disease-specific phenotypes in dopamine neurons from human iPS-based models of genetic and sporadic Parkinson's disease. EMBO Mol. Med. 4, 380–395 (2012).
Burkhardt, M.F. et al. A cellular model for sporadic ALS using patient-derived induced pluripotent stem cells. Mol. Cell. Neurosci. 56C, 355–364 (2013).
Yang, Y.M. et al. A small molecule screen in stem-cell-derived motor neurons identifies a kinase inhibitor as a candidate therapeutic for ALS. Cell Stem Cell 12, 713–726 (2013).
Brooks, B.P. et al. Characterization of an expanded glutamine repeat androgen receptor in a neuronal cell culture system. Neurobiol. Dis. 3, 313–323 (1997).
Malik, B. et al. Absence of disturbed axonal transport in spinal and bulbar muscular atrophy. Hum. Mol. Genet. 20, 1776–1786 (2011).
Bailey, C.K., Andriola, I.F., Kampinga, H.H. & Merry, D.E. Molecular chaperones enhance the degradation of expanded polyglutamine repeat androgen receptor in a cellular model of spinal and bulbar muscular atrophy. Hum. Mol. Genet. 11, 515–523 (2002).
Acknowledgements
The authors wish to thank L.I. Macedo de Souza, A.C. Smith and H. Burke for technical assistance, N. Mizushima (University of Tokyo) and Z. Yue for (Icahn School of Medicine at Mount Sinai) providing GFP-LC3 transgenic mice, J.P. Taylor (St. Jude's Children's Research Hospital) for supplying the mCherry-GFP-LC3 MEFs and T. Johansen (Arctic University of Norway) for the gift of the mCherry-EGFP-LC3 construct. This work was supported by grants from the US National Institutes of Health (R01 NS041648 to A.R.L., R01 AG033082 to A.R.L. and DP2-OD006495-01 to A.R.M.), the Muscular Dystrophy Association (Basic Research Grant to A.R.L., Development Grant to C.J.C. and Basic Research Grant to A.R.M.), and the California Institute for Regenerative Medicine (CIRM TR2-01814 to A.R.M.).
Author information
Authors and Affiliations
Contributions
C.J.C., H.C.M., H.F., Y.B., J.E.Y., A.R.M., G.A.G. and A.R.L.S. designed the experiments. C.J.C., H.C.M., H.F., Y.B., J.E.Y., A.L., N.I., B.L.S., C.C. and G.A.G. performed the experiments. C.J.C., H.C.M., H.F., Y.B., J.E.Y., A.R.M., G.A.G. and A.R.L.S. analyzed the data. C.J.C., H.C.M., G.A.G. and A.R.L.S. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Reduced long-term protein turnover in MN-1 AR65Q cells.
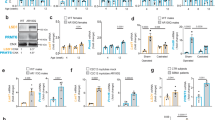
After labeling of newly synthesized proteins with L-azidohomoalanine (AHA), an analog of methionine, we measured rates of protein turnover over 100 hrs after the AHA pulse. We noted a significantly reduced rate of long-term protein turnover for MN-1 AR65Q cells in comparison to wild-type MN-1 cells (n = 3 independent experiments, ANOVA with post-hoc Tukey test, *P <.05). MN-1 AR24Q cells displayed a rate of protein turnover intermediate to wild-type MN-1 cells and MN-1 AR65Q cells, but the MN-1 AR24Q protein turnover rate was not significantly different from either cell line. MN-1 WT cells treated with leupeptin served as a positive control. Data are presented as mean + s.e.m.
Supplementary Figure 2 Detection of autophagosomes and autolysosomes.
Here we see representative images of autophagosomes (yellow arrowheads) and autolysosomes (red arrowhead). Autophagosomes were identified as double-membrane bound vacuoles containing heterogeneous cytoplasmic material within the double membrane, based upon well-established criteria (Eskelinen et. al., 2004, Mol Biol Cell 15: 3132–3145; Nixon et al., 2005, Neuropathol Exp Neurol 64: 113–122). Magnification of autophagosomes is at 12,460x. Autolysosomes were identified as vacuoles with homogeneous contents that were associated with electron-dense lysosomes, as delineated in previous studies (Eskelinen et. al., 2004, Mol Biol Cell 15: 3132–3145; Nixon et al., 2005, Neuropathol Exp Neurol 64: 113–122). The autolysosome panel is at 10,150x magnification.
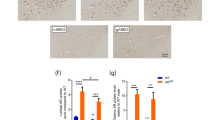
Supplementary Figure 3 Increased expression of TFEB target genes in skeletal muscle of SBMA YAC AR100 mice.
We measured expression levels of TFEB target genes in quadriceps muscle from non-transgenic (Nt), YAC AR20, and YAC AR100 transgenic mice. RT-PCR analysis of isolated RNAs revealed significant increases in TFEB target gene expression in YAC AR20 and YAC AR100 mice: Lamp1 (lysosomal-associated membrane protein 1) [F=2,12]=8.15, Atp6v1h (vesicular ATPase V1 subunit H) [F(2,12)=49.95]; Gla (galactosidase-α) F(2,12)=8.63]; Ctsd (cathepsin D) [F(2,12)=25.57]; Mcoln1 (mucolipin1) [F(2,12)=31.164]; Ctsf (cathepsin F) [F(2,12)=264.43]; Sqstm1 (sequesterome 1, p62) [F(2,12)=280.48], and Tfeb [F(2,12)=8.32]; n = 3 independent experiments, *P <.05, **P <.01, ***P <.001, ANOVA with post-hoc Tukey test. Data are presented as mean + s.e.m.
Supplementary Figure 4 TFEB promotes autophagic flux in MN-1 WT cells
Quantification of autophagic vesicle type in MN-1 WT cells that were transfected with BFP-empty, or transfected with BFP-TFEB. Note marked increase in autolysosomes in MN-1 WT cells expressing BFP-TFEB; n = 3 independent experiments, F(2,87)=4.97, ANOVA with post-hoc Tukey test. **P <.01. Data are presented as mean + s.e.m
Supplementary Figure 5 Summary of human sample production and derivation for stem cell model work.
Here we see a diagram delineating the number of different human patients lines and corresponding number of induced pluripotent stem cell (iPSC) lines and their clonal derivatives utilized in this study. As shown here, we derived three unique clonal lines per iPSC line, and we created neuron precursor cell (NPC) lines for EACH unique clone shown here. Hence, all NPC experimentation involved the use of three unique clones per human patient sample. This was done to dramatically reduce the likelihood of random variation creating artifact results in affected samples when compared to control samples. The CAG repeat allele size for each patient sample is indicated.
Supplementary Figure 6 Characterization of iPSC and NPC cell lines.
(a) We confirmed expression of the pluripotency markers Oct4, Nanog, Lin28, and Sox2 in all iPSC lines. Representative images, with the nuclei stained with DAPI, are shown here. Scale bar = 40 μM. (b) We confirmed that embryoid bodies generated from iPSC lines display expression of markers from all three germ layers based upon RT-PCR analysis of isolated RNAs. Representative results for endoderm markers (ASP, ESG), mesoderm markers (BRACH, GATA4), and ectoderm markers (NESTIN, PAX6) are shown. (c) We confirmed normal chromosome numbers for all six iPSC lines by karyotyping analysis. Representative normal karyotyping results for one iPSC line are shown here. (d) We confirmed expression of the neural progenitor markers Sox2 and Nestin in NPC lines derived from iPSC lines. Representative images, with the nuclei stained with DAPI, are shown here. Scale bar = 40 μm.
Supplementary Figure 7 Measurement of mitochondrial membrane polarization state in NPCs.
We cultured control and SBMA NPCs in the presence of JC-1 dye, a ratiometric probe that fluoresces differentially based upon whether it is taken up and retained by normally polarized mitochondria (red) or remains in the cytosol (green), and sorted cells based upon the red: green ratio by FACS. Here we see a representative run demonstrating an increased number of SBMA NPCs with a decreased red: green ratio.
Supplementary Figure 8 Effect of TFEB on control NPCs.
Quantification of autophagic vesicle type in control NPCs that were transfected with BFP-empty, or transfected with BFP-TFEB. Note the trend toward an increase in autolysosomes in control NPCs expressing BFP-TFEB, but lack of significant increase; n = 3 independent experiments. Data are presented as mean + s.e.m.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–9 and Supplementary Table 1 (PDF 3397 kb)
Supplementary Methods Checklist
(PDF 516 kb)
Rights and permissions
About this article
Cite this article
Cortes, C., Miranda, H., Frankowski, H. et al. Polyglutamine-expanded androgen receptor interferes with TFEB to elicit autophagy defects in SBMA. Nat Neurosci 17, 1180–1189 (2014). https://doi.org/10.1038/nn.3787
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nn.3787
This article is cited by
-
Transcriptional regulation of autophagy and its implications in human disease
Cell Death & Differentiation (2023)
-
Bicalutamide and Trehalose Ameliorate Spinal and Bulbar Muscular Atrophy Pathology in Mice
Neurotherapeutics (2023)
-
An mTORC1-mediated negative feedback loop constrains amino acid-induced FLCN-Rag activation in renal cells with TSC2 loss
Nature Communications (2022)
-
AR cooperates with SMAD4 to maintain skeletal muscle homeostasis
Acta Neuropathologica (2022)
-
Machinery, regulation and pathophysiological implications of autophagosome maturation
Nature Reviews Molecular Cell Biology (2021)