Abstract
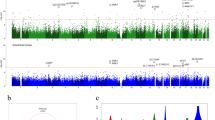
Alzheimer's disease (AD) is a chronic neurodegenerative disorder that is characterized by progressive neuropathology and cognitive decline. We performed a cross-tissue analysis of methylomic variation in AD using samples from four independent human post-mortem brain cohorts. We identified a differentially methylated region in the ankyrin 1 (ANK1) gene that was associated with neuropathology in the entorhinal cortex, a primary site of AD manifestation. This region was confirmed as being substantially hypermethylated in two other cortical regions (superior temporal gyrus and prefrontal cortex), but not in the cerebellum, a region largely protected from neurodegeneration in AD, or whole blood obtained pre-mortem from the same individuals. Neuropathology-associated ANK1 hypermethylation was subsequently confirmed in cortical samples from three independent brain cohorts. This study represents, to the best of our knowledge, the first epigenome-wide association study of AD employing a sequential replication design across multiple tissues and highlights the power of this approach for identifying methylomic variation associated with complex disease.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Accession codes
References
Brookmeyer, R., Johnson, E., Ziegler-Graham, K. & Arrighi, H.M. Forecasting the global burden of Alzheimer's disease. Alzheimers Dement. 3, 186–191 (2007).
Knapp, M. & Prince, M. Dementia UK: the Full report. (Alzheimer's Society, 2007).
Hardy, J.A. & Higgins, G.A. Alzheimer's disease: the amyloid cascade hypothesis. Science 256, 184–185 (1992).
Hardy, J. & Selkoe, D.J. The amyloid hypothesis of Alzheimer's disease: progress and problems on the road to therapeutics. Science 297, 353–356 (2002).
Wenk, G.L. Neuropathologic changes in Alzheimer's disease. J. Clin. Psychiatry 64 (suppl. 9), 7–10 (2003).
Lambert, J.C. et al. Meta-analysis of 74,046 individuals identifies 11 new susceptibility loci for Alzheimer's disease. Nat. Genet. 45, 1452–1458 (2013).
Encode Project Consortium et al. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Lunnon, K. & Mill, J. Epigenetic studies in Alzheimer's disease: current findings, caveats, and considerations for future studies. Am. J. Med. Genet. B. Neuropsychiatr. Genet. 162B, 789–799 (2013).
Mill, J. & Heijmans, B.T. From promises to practical strategies in epigenetic epidemiology. Nat. Rev. Genet. 14, 585–594 (2013).
Murphy, T.M. & Mill, J. Epigenetics in health and disease: heralding the EWAS era. Lancet 383, 1952–1954 (2014).
Pidsley, R. et al. A data-driven approach to preprocessing Illumina 450K methylation array data. BMC Genomics 14, 293 (2013).
Braak, H. & Braak, E. Neuropathological stageing of Alzheimer-related changes. Acta Neuropathol. 82, 239–259 (1991).
Kordeli, E. & Bennett, V. Distinct ankyrin isoforms at neuron cell bodies and nodes of Ranvier resolved using erythrocyte ankyrin-deficient mice. J. Cell Biol. 114, 1243–1259 (1991).
Boiko, T. et al. Ankyrin-dependent and -independent mechanisms orchestrate axonal compartmentalization of L1 family members neurofascin and L1/neuron-glia cell adhesion molecule. J. Neurosci. 27, 590–603 (2007).
Nelson, S.B., Janiesch, C. & Sander, M. Expression of Nkx6 genes in the hindbrain and gut of the developing mouse. J. Histochem. Cytochem. 53, 787–790 (2005).
Alanentalo, T. et al. Cloning and analysis of Nkx6.3 during CNS and gastrointestinal development. Gene Expr. Patterns 6, 162–170 (2006).
Guintivano, J., Aryee, M. & Kaminsky, Z. A cell epigenotype specific model for the correction of brain cellular heterogeneity bias and its application to age, brain region and major depression. Epigenetics 8, 290–302 (2013).
Pedersen, B.S., Schwartz, D.A., Yang, I.V. & Kechris, K.J. Comb-p: software for combining, analyzing, grouping and correcting spatially correlated P-values. Bioinformatics 28, 2986–2988 (2012).
Haroutunian, V. et al. Regional distribution of neuritic plaques in the nondemented elderly and subjects with very mild Alzheimer disease. Arch. Neurol. 55, 1185–1191 (1998).
Esiri, M.M. Brain banks: the Oxford experience. J. Neural Transm. Suppl. 39, 25–30 (1993).
De Jager, P.L. et al. Alzheimer's disease pathology is associated with early alterations in brain DNA methylation at ANK1, BIN1 and other loci. Nat. Neurosci. 10.1038/nn.3786 (17 August 2014).
Soranzo, N. et al. Common variants at 10 genomic loci influence hemoglobin A(1)(C) levels via glycemic and nonglycemic pathways. Diabetes 59, 3229–3239 (2010).
Imamura, M. et al. A single-nucleotide polymorphism in ANK1 is associated with susceptibility to type 2 diabetes in Japanese populations. Hum. Mol. Genet. 21, 3042–3049 (2012).
Harder, M.N. et al. Type 2 diabetes risk alleles near BCAR1 and in ANK1 associate with decreased beta-cell function whereas risk alleles near ANKRD55 and GRB14 associate with decreased insulin sensitivity in the Danish Inter99 cohort. J. Clin. Endocrinol. Metab. 98, E801–E806 (2013).
Barnes, D.E. & Yaffe, K. The projected effect of risk factor reduction on Alzheimer's disease prevalence. Lancet Neurol. 10, 819–828 (2011).
Lovestone, S. Searching for biomarkers in neurodegeneration. Nat. Med. 16, 1371–1372 (2010).
Hooper, C., Lovestone, S. & Sainz-Fuertes, R. Alzheimer's disease, diagnosis and the need for biomarkers. Biomark. Insights 3, 317–323 (2008).
Fehlbaum-Beurdeley, P. et al. Toward an Alzheimer's disease diagnosis via high-resolution blood gene expression. Alzheimers Dement. 6, 25–38 (2010).
Rye, P.D. et al. A novel blood test for the early detection of Alzheimer's disease. J. Alzheimers Dis. 23, 121–129 (2011).
Booij, B.B. et al. A gene expression pattern in blood for the early detection of Alzheimer's disease. J. Alzheimers Dis. 23, 109–119 (2011).
Lunnon, K. et al. A blood gene expression marker of early Alzheimer's disease. J. Alzheimers Dis. 33, 737–753 (2013).
Fehlbaum-Beurdeley, P. et al. Validation of AclarusDx, a blood-based transcriptomic signature for the diagnosis of Alzheimer's disease. J. Alzheimers Dis. 32, 169–181 (2012).
Wu, Z.C. et al. Association of DAPK1 genetic variations with Alzheimer's disease in Han Chinese. Brain Res. 1374, 129–133 (2011).
Laumet, G. et al. Systematic analysis of candidate genes for Alzheimer's disease in a French, genome-wide association study. J. Alzheimers Dis. 20, 1181–1188 (2010).
Chapuis, J., Vingtdeux, V., Campagne, F., Davies, P. & Marambaud, P. Growth arrest–specific 1 binds to and controls the maturation and processing of the amyloid-beta precursor protein. Hum. Mol. Genet. 20, 2026–2036 (2011).
Lunnon, K. et al. Mitochondrial dysfunction and immune activation are detectable in early Alzheimer's disease blood. J. Alzheimers Dis. 30, 685–710 (2012).
Meaburn, E.L., Schalkwyk, L.C. & Mill, J. Allele-specific methylation in the human genome: implications for genetic studies of complex disease. Epigenetics 5, 578–582 (2010).
Shoemaker, R., Deng, J., Wang, W. & Zhang, K. Allele-specific methylation is prevalent and is contributed by CpG-SNPs in the human genome. Genome Res. 20, 883–889 (2010).
Davies, M.N. et al. Functional annotation of the human brain methylome identifies tissue-specific epigenetic variation across brain and blood. Genome Biol. 13, R43 (2012).
Dempster, E.L. et al. Disease-associated epigenetic changes in monozygotic twins discordant for schizophrenia and bipolar disorder. Hum. Mol. Genet. 20, 4786–4796 (2011).
Wong, C.C. et al. Methylomic analysis of monozygotic twins discordant for autism spectrum disorder and related behavioural traits. Mol. Psychiatry 19, 495–503 (2014).
R Development Core Team. R: a language and environment for statistical computing. (R Foundation for Statistical Computing, Vienna, Austria, 2012).
Gentleman, R.C. et al. Bioconductor: open software development for computational biology and bioinformatics. Genome Biol. 5, R80 (2004).
Davis, S., Du, P., Bilke, S., Triche, J. & Bootwalla, M. Methylumi: Handle Illumina Methylation Data, v2.10.0 (R Graphic Manual, 2014).
Chen, Y.A. et al. Discovery of cross-reactive probes and polymorphic CpGs in the Illumina Infinium HumanMethylation450 microarray. Epigenetics 8, 203–209 (2013).
Acknowledgements
We thank C. Sloan for technical support and I. Bodi and A. King for neuropathological diagnosis of cases. We also thank the Oxford Project to Investigate Memory and Ageing (OPTIMA), the National Institute for Health (NIHR) Biomedical Research Unit in Dementia in the South London and Maudsley NHS Foundation Trust (SLaM), Brains for Dementia Research (Alzheimer Brain Bank, UK), and the donors and families who made this research possible. Blood samples from the London cohort were collected as part of the Alzheimer's Research UK funded study “Biomarkers of AD Neurodegeneration”. This work was funded by US National Institutes of Health grant R01 AG036039 to J.M. and an Equipment Grant from Alzheimer's Research UK. The Oxford Brain Bank is supported in part by the NIHR Oxford Biomedical Research Centre based at Oxford University Hospitals NHS Trust and University of Oxford. Brain banking and neuropathology assessments for the Mount Sinai cohort was supported by US National Institutes of Health grants AG02219, AG05138 and MH064673, and the Department of Veterans Affairs VISN3 MIRECC. Replication work in Boston was supported by US National Institutes of Health grants: R01 AG036042, R01AG036836, R01 AG17917, R01 AG15819, R01 AG032990, R01 AG18023, RC2 AG036547, P30 AG10161, P50 AG016574, U01 ES017155, KL2 RR024151 and K25 AG041906-01.
Author information
Authors and Affiliations
Contributions
K.L., R.S., R.M., M.V., D.C. and J.B. conducted laboratory experiments. J.M. conceived and supervised the project and obtained funding. E.H., L.C.S., R.S. and K.L. undertook data analyses and bioinformatics. C.T., S.L., J.P., S.A.-S., P.K., V.H. and C.J. provided samples for analysis. P.L.D.J., G.S. and D.A.B. provided replication data. Z.K. provided help with cellular heterogeneity correction. L.W.H. provided help with the alternative splicing assays. J.M., K.L. and L.C.S. drafted the manuscript. All of the authors read and approved the final submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Replication of neuropathology-associated ANK1 hypermethylation in the cortex.
Bisulfite-pyrosequencing was used to analyze methylation status across eight CpG sites spanning an extended ANK1 region in the Oxford cohort (N = 62). Methylation betas were adjusted for age and gender, and linear models used to compare between controls (Braak score 0-II) and subjects with AD (Braak score V-VI). Average DNA methylation across this region was significantly elevated in all three cortical regions (EC P = 0.0004; STG P = 0.0008; PFC P = 0.014) in affected individuals, most notably in the EC where six of the eight CpG sites assessed are characterized by significant AD-associated hypermethylation (see Fig. 2d). Source Data is provided in the Supplementary Figure Source Data file.
Supplementary Figure 2 Correlations between the 137 DMPs associated with NP pathology burden at P < 1.20 × 10–7 that were identified by De Jager et al., and our cross-cortex data.
Neuropathology-associated DNA methylation scores for the top-ranked DMPs identified by De Jager and colleagues (X-axis)21 are strongly correlated (r = 0.49, P = 7.8E-10) with cross-cortex Braak-associated differences at the same probes in our samples (Y-axis). Abbreviations: DLFPC; Dorsolateral Prefrontal Cortex. Source Data is provided in the Supplementary Figure Source Data file.
Supplementary Figure 3 ANK1 transcript variants assessed using qRT-PCR.
Primer sequences for the three assays shown in red are given in Supplementary Table S11b.
Supplementary Figure 4 Neuropathology-associated elevation of ANK1 isoform 5-7-10.
Three qPCR assays with specificity to ANK1 isoforms i) 1-2-3-4, ii) 9, and iii) 5-7-10 were run on 36 EC samples in the London cohort from whom RNA was available (see Supplementary Table S2). We found a significant association (ANOVA P = 0.04) between the abundance of isoform 5-7-10 transcripts and AD-associated neuropathology. No significant differences in transcript levels were observed for the other two isoform-specific assays (data not shown). Source Data is provided in the Supplementary Figure Source Data file.
Supplementary Figure 5 Q-Q plot for cross-cortex neuropathology analysis.
Scatterplot shows the negative logarithm of each Brown’s P value from the linear model predicting each corrected beta with Braak stage plotted against P value rank (see Online Methods). The two ANK1 associated probes are shown in red.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5 and Supplementary Tables 1, 7, 8, 10, 11, and 13 (PDF 1485 kb)
41593_2014_BFnn3782_MOESM42_ESM.xlsx
Supplementary Table 2: Detailed demographic information for individual samples in the London 'discovery' cohort. (XLSX 22 kb)
Supplementary Table 3: The 100 top-ranked EC DMPs associated with Braak score in the London 'discovery' cohort.
DMPs were identified within the EC and then cross-tissue (STG, PFC, CER) validated within the same cohort. DMPs were subsequently assessed in cortical regions from the Mount Sinai replication cohort. Presented is the corrected difference in methylation level (Δ) and calculated P value between Braak score 0 and Braak score VI after adjusting for covariates of age and sex. (XLSX 27 kb)
Supplementary Table 4: The 100 top-ranked STG DMPs associated with Braak score in the London 'discovery' cohort.
DMPs were identified within the STG and then cross-tissue (EC, PFC, CER) validated within the same cohort. DMPs were subsequently assessed in cortical regions from the Mount Sinai replication cohort. Presented is the corrected difference in methylation level (Δ) and calculated P value between Braak score 0 and Braak score VI after adjusting for covariates of age and sex. (XLSX 28 kb)
Supplementary Table 5: The 100 top-ranked PFC DMPs associated with Braak score in the London 'discovery' cohort.
DMPs were identified within the PFC and then cross-tissue (EC, STG, CER) validated within the same cohort. DMPs were subsequently assessed in cortical regions from the Mount Sinai replication cohort. Presented is the corrected difference in methylation level (Δ) and calculated P value between Braak score 0 and Braak score VI after adjusting for covariates of age and sex. (XLSX 100 kb)
Supplementary Table 6: The 100 top-ranked CER DMPs associated with Braak score in the London 'discovery' cohort.
DMPs were identified within the PFC and then cross-tissue (EC, STG, PFC) validated within the same cohort. DMPs were subsequently assessed in cortical regions from the Mount Sinai replication cohort. Presented is the corrected difference in methylation level (Δ) and calculated P value between Braak score 0 and Braak score VI after adjusting for covariates of age and sex. (XLSX 25 kb)
Supplementary Table 9: Consistent cross-cortex DMPs associated with Braak score.
Shown are the top 100 DMPs associated with Braak score, across cortical tissue using a cross-cortex meta-analysis. Shown for each DMP are chromosomal location (Hg19), up-/downstream genes, P value from Fisher's test (see Online Methods), P Value from Brown's test (see Online Methods), and estimate (Δ) of methylation difference between individuals with the lowest (score 0) and highest (score VI) from cross-cortex meta-analysis. Also shown are differences (Δ) between Braak 0 and Braak IV and corresponding P values from our individual cortex models in the London discovery cohort (see Online Methods) and the STG and PFC in the Mount Sinai replication cohort. Missing data denotes samples that are not significant. (XLSX 34 kb)
Supplementary Table 12: The 100 top-ranked pre-mortem blood DMPs associated with a diagnosis of AD in the London 'discovery' cohort.
Using a case-control analysis we compared methylation between AD blood samples (in the absence of another dementia) and control blood samples and compared the top 100 p-value ranked DMPs to methylation values in the four brain regions. Presented is the corrected difference in DNA methylation (Δ) and calculated P value after adjusting for covariates of age and sex for control and AD samples. There is little overlap between the top-ranked DMPs identified in blood and nominally-significant (P < 0.05) AD-associated differences in the same direction in EC, STG, PFC or CER. (XLSX 22 kb)
Rights and permissions
About this article
Cite this article
Lunnon, K., Smith, R., Hannon, E. et al. Methylomic profiling implicates cortical deregulation of ANK1 in Alzheimer's disease. Nat Neurosci 17, 1164–1170 (2014). https://doi.org/10.1038/nn.3782
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nn.3782
This article is cited by
-
Quantifying the proportion of different cell types in the human cortex using DNA methylation profiles
BMC Biology (2024)
-
Distinct CSF biomarker-associated DNA methylation in Alzheimer’s disease and cognitively normal subjects
Alzheimer's Research & Therapy (2023)
-
Entorhinal cortex epigenome-wide association study highlights four novel loci showing differential methylation in Alzheimer’s disease
Alzheimer's Research & Therapy (2023)
-
The contribution of DNA methylation to the (dys)function of oligodendroglia in neurodegeneration
Acta Neuropathologica Communications (2023)
-
Step by step: towards a better understanding of the genetic architecture of Alzheimer’s disease
Molecular Psychiatry (2023)