Abstract
Feeding can be inhibited by multiple cues, including those associated with satiety, sickness or unpalatable food. How such anorexigenic signals inhibit feeding at the neural circuit level is not completely understood. Although some inhibitory circuits have been identified, it is not yet clear whether distinct anorexigenic influences are processed in a convergent or parallel manner. The amygdala central nucleus (CEA) has been implicated in feeding control, but its role is controversial. The lateral subdivision of CEA (CEl) contains a subpopulation of GABAergic neurons that are marked by protein kinase C-δ (PKC-δ). We found that CEl PKC-δ+ neurons in mice were activated by diverse anorexigenic signals in vivo, were required for the inhibition of feeding by such signals and strongly suppressed food intake when activated. They received presynaptic inputs from anatomically distributed neurons activated by different anorexigenic agents. Our data suggest that CEl PKC-δ+ neurons constitute an important node that mediates the influence of multiple anorexigenic signals.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Sternson, S.M. Hypothalamic survival circuits: blueprints for purposive behaviors. Neuron 77, 810–824 (2013).
Morton, G.J., Cummings, D.E., Baskin, D.G., Barsh, G.S. & Schwartz, M.W. Central nervous system control of food intake and body weight. Nature 443, 289–295 (2006).
Saper, C.B., Chou, T.C. & Elmquist, J.K. The need to feed: homeostatic and hedonic control of eating. Neuron 36, 199–211 (2002).
Woods, S.C., Seeley, R.J., Porte, D. Jr. & Schwartz, M.W. Signals that regulate food intake and energy homeostasis. Science 280, 1378–1383 (1998).
Carter, M.E., Soden, M.E., Zweifel, L.S. & Palmiter, R.D. Genetic identification of a neural circuit that suppresses appetite. Nature 503, 111–114 (2013).
Wu, Q., Clark, M.S. & Palmiter, R.D. Deciphering a neuronal circuit that mediates appetite. Nature 483, 594–597 (2012).
Kishi, T. & Elmquist, J.K. Body weight is regulated by the brain: a link between feeding and emotion. Mol. Psychiatry 10, 132–146 (2005).
Betley, J.N., Cao, Z.F., Ritola, K.D. & Sternson, S.M. Parallel, redundant circuit organization for homeostatic control of feeding behavior. Cell 155, 1337–1350 (2013).
King, B.M. Amygdaloid lesion-induced obesity: relation to sexual behavior, olfaction and the ventromedial hypothalamus. Am. J. Physiol. Regul. Integr. Comp. Physiol. 291, R1201–R1214 (2006).
Kask, A. & Schioth, H.B. Tonic inhibition of food intake during inactive phase is reversed by the injection of the melanocortin receptor antagonist into the paraventricular nucleus of the hypothalamus and central amygdala of the rat. Brain Res. 887, 460–464 (2000).
Beckman, T.R., Shi, Q., Levine, A.S. & Billington, C.J. Amygdalar opioids modulate hypothalamic melanocortin-induced anorexia. Physiol. Behav. 96, 568–573 (2009).
Fekete, E., Vigh, J., Bagi, E.E. & Lenard, L. Gastrin-releasing peptide microinjected into the amygdala inhibits feeding. Brain Res. 955, 55–63 (2002).
Fekete, E.M., Bagi, E.E., Toth, K. & Lenard, L. Neuromedin C microinjected into the amygdala inhibits feeding. Brain Res. Bull. 71, 386–392 (2007).
Kovács, A. et al. Microinjection of RFRP-1 in the central nucleus of amygdala decreases food intake in the rat. Brain Res. Bull. 88, 589–595 (2012).
Bovetto, S. & Richard, D. Lesion of central nucleus of amygdala promotes fat gain without preventing effect of exercise on energy balance. Am. J. Physiol. 269, R781–R786 (1995).
Lenard, L. & Hahn, Z. Amygdalar noradrenergic and dopaminergic mechanisms in the regulation of hunger and thirst-motivated behavior. Brain Res. 233, 115–132 (1982).
Box, B.M. & Mogenson, G.J. Alterations in ingestive behaviors after bilateral lesions of the amygdala in the rat. Physiol. Behav. 15, 679–688 (1975).
Kemble, E.D., Studelska, D.R. & Schmidt, M.K. Effects of central amygdaloid nucleus lesions on ingestion, taste reactivity, exploration and taste aversion. Physiol. Behav. 22, 789–793 (1979).
Petrovich, G.D., Ross, C.A., Mody, P., Holland, P.C. & Gallagher, M. Central, but not basolateral, amygdala is critical for control of feeding by aversive learned cues. J. Neurosci. 29, 15205–15212 (2009).
Haubensak, W. et al. Genetic dissection of an amygdala microcircuit that gates conditioned fear. Nature 468, 270–276 (2010).
Day, H.E., Curran, E.J., Watson, S.J. Jr. & Akil, H. Distinct neurochemical populations in the rat central nucleus of the amygdala and bed nucleus of the stria terminalis: evidence for their selective activation by interleukin-1beta. J. Comp. Neurol. 413, 113–128 (1999).
Marchant, N.J., Densmore, V.S. & Osborne, P.B. Coexpression of prodynorphin and corticotrophin-releasing hormone in the rat central amygdala: evidence of two distinct endogenous opioid systems in the lateral division. J. Comp. Neurol. 504, 702–715 (2007).
Moran, T.H. Cholecystokinin and satiety: current perspectives. Nutrition 16, 858–865 (2000).
Yamamoto, T. & Ueji, K. Brain mechanisms of flavor learning. Front. Syst. Neurosci. 5, 76 (2011).
Dantzer, R. Cytokine-induced sickness behavior: mechanisms and implications. Ann. NY Acad. Sci. 933, 222–234 (2001).
Haba, R. et al. Lipopolysaccharide affects exploratory behaviors toward novel objects by impairing cognition and/or motivation in mice: possible role of activation of the central amygdala. Behav. Brain Res. 228, 423–431 (2012).
Lee, H.M., Giguere, P.M. & Roth, B.L. DREADDs: novel tools for drug discovery and development. Drug Discov. Today 19, 469–473 (2014).
Krashes, M.J. et al. Rapid, reversible activation of AgRP neurons drives feeding behavior in mice. J. Clin. Invest. 121, 1424–1428 (2011).
Gautron, L., Lazarus, M., Scott, M.M., Saper, C.B. & Elmquist, J.K. Identifying the efferent projections of leptin-responsive neurons in the dorsomedial hypothalamus using a novel conditional tracing approach. J. Comp. Neurol. 518, 2090–2108 (2010).
Gradinaru, V. et al. Molecular and cellular approaches for diversifying and extending optogenetics. Cell 141, 154–165 (2010).
Zhang, F. et al. Multimodal fast optical interrogation of neural circuitry. Nature 446, 633–639 (2007).
Ciocchi, S. et al. Encoding of conditioned fear in central amygdala inhibitory circuits. Nature 468, 277–282 (2010).
Maren, S. & Fanselow, M.S. The amygdala and fear conditioning: has the nut been cracked? Neuron 16, 237–240 (1996).
Pitkänen, A., Savander, V. & LeDoux, J.E. Organization of intra-amygdaloid circuitries in the rat: an emerging framework for understanding functions of the amygdala. Trends Neurosci. 20, 517–523 (1997).
Tye, K.M. et al. Amygdala circuitry mediating reversible and bidirectional control of anxiety. Nature 471, 358–362 (2011).
Panksepp, J. Cross-species affective neuroscience decoding of the primal affective experiences of humans and related animals. PLoS ONE 6, e21236 (2011).
Wall, N.R., Wickersham, I.R., Cetin, A., De La Parra, M. & Callaway, E.M. Monosynaptic circuit tracing in vivo through Cre-dependent targeting and complementation of modified rabies virus. Proc. Natl. Acad. Sci. USA 107, 21848–21853 (2010).
Watabe-Uchida, M., Zhu, L., Ogawa, S.K., Vamanrao, A. & Uchida, N. Whole-brain mapping of direct inputs to midbrain dopamine neurons. Neuron 74, 858–873 (2012).
Petreanu, L., Huber, D., Sobczyk, A. & Svoboda, K. Channelrhodopsin-2–assisted circuit mapping of long-range callosal projections. Nat. Neurosci. 10, 663–668 (2007).
Oh, S.W. et al. A mesoscale connectome of the mouse brain. Nature 508, 207–214 (2014).
Lee, H. et al. Scalable control of mounting and attack by Esr1 neurons in the ventromedial hypothalamus. Nature 509, 627–632 (2014).
Taniguchi, H. et al. A resource of Cre driver lines for genetic targeting of GABAergic neurons in cerebral cortex. Neuron 71, 995–1013 (2011).
Neugebauer, V., Li, W., Bird, G.C. & Han, J.S. The amygdala and persistent pain. Neuroscientist 10, 221–234 (2004).
Jennings, J.H., Rizzi, G., Stamatakis, A.M., Ung, R.L. & Stuber, G.D. The inhibitory circuit architecture of the lateral hypothalamus orchestrates feeding. Science 341, 1517–1521 (2013).
Madisen, L. et al. A robust and high-throughput Cre reporting and characterization system for the whole mouse brain. Nat. Neurosci. 13, 133–140 (2010).
Aravanis, A.M. et al. An optical neural interface: in vivo control of rodent motor cortex with integrated fiberoptic and optogenetic technology. J. Neural Eng. 4, S143–S156 (2007).
Neugebauer, V. & Li, W. Processing of nociceptive mechanical and thermal information in central amygdala neurons with knee-joint input. J. Neurophysiol. 87, 103–112 (2002).
Yizhar, O., Fenno, L.E., Davidson, T.J., Mogri, M. & Deisseroth, K. Optogenetics in neural systems. Neuron 71, 9–34 (2011).
Vrontou, S., Wong, A.M., Rau, K.K., Koerber, H.R. & Anderson, D.J. Genetic identification of C fibres that detect massage-like stroking of hairy skin in vivo. Nature 493, 669–673 (2013).
Acknowledgements
We thank B. Lowell (Beth Israel Deaconess Medical Center) for providing Cre-dependent DREADD constructs and viruses for pilot experiments, C. Saper (Beth Israel Deaconess Medical Center) for providing the original hrGFP construct, Z.J. Huang (Cold Spring Harbor Laboratory) for CRF-Cre mice, the Allen Institute for Brain Science for Tac2-Cre mice, C. Xiao and H. Lester for electrophysiology training and advice, and K. Deisseroth (Stanford University) for providing viruses for optogenetics. We thank J.S. Chang, C. Chiu and A. Chang for technical assistance on histology and behavior scoring, M. Martinez and M. McCardle for tail genotyping, G. Mosconi and C. Chiu for lab management, and G. Mancuso for administrative support. This work was supported by US National Institutes of Health grants R01MH085082 and MH070053, and a grant from the Klarman Foundation to D.J.A. H.C. is supported by a postdoctoral fellowship of the Hilda and Preston Davis Foundation and a NARSAD Young Investigator Award. D.J.A. receives funding from the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
H.C. designed and performed the experiments and wrote the manuscript. W.H. generated BAC constructs for PKC-δ+ transgenic mice. T.E.A. made the virus constructs for Cre-out ChR2 and Cre-out eNpHR. D.J.A. contributed to experimental design and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Diverse anorexigenic signals induce c-Fos expression in CEl PKC-δ+ neurons
a-c. Quantification of CEl c-Fos expression in mice intraperitoneal injected with anorexigenic drugs (a), 24 h fasted with or without 3 hr re-feeding (b), or orally infused with water, quinine or sucrose solution (c). Note: some sections of mice injected with saline (a, right), and mice with oral infusion of water or sucrose (c, right) contained no c-Fos+ cells in CEl; in those cases the values for PKC-δ+/Fos+ were assigned as 0. Box plots in (b) show mean (+), median, quartiles (boxes), and range (whiskers). Values in (a) and (c) are means ± s.e.m.. For (a), n = 6 (saline), 7 (CCK), 7 (LiCl), 8 (LPS) brain sections from 4 animals in each condition. One-way ANOVA (left, F(3, 24) = 29.1, p < 0.0001; right, F(3, 24) = 39.8, p < 0.0001) with post-hoc Bonferroni t-test. For (b), n = 15 brain sections from 4 animals (24 h fasted) and 17 brain sections from 5 animals (re-fed). Unpaired t-test (left, t (30) = 7.88, p < 0.0001; right, t (30) = 3.95, p = 0.0004). For (c), n = 12 (water), 15 (quinine), 12 (sucrose) sections from 3 animals in each condition. One-way ANOVA (left, F(2, 36) = 34.1, p < 0.0001; right, F(2, 36) = 6.97, p = 0.0028) with post-hoc Bonferroni t-test. * p<0.05, ** p < 0.01, *** p < 0.001.
Supplementary Figure 2 CEl c-Fos analysis after silencing CEl PKC-δ+ neurons in response to CCK, LiCl or LPS
a. Expression of c-Fos (antibody staining) and PKC-δ (indicated by Cre-dependent virus expression of fluorescent proteins) after injecting CCK, LiCl or LPS. Experimental procedure is shown on the top. The insets show the enlarged boxed areas. b-d. Quantification of c-Fos and PKC-δ+ neurons in each CEl section from mice injected with CCK (b), LiCl (c), or LPS (d). n = 4 animals in each group. Values are means ± s.e.m.; Unpaired t-test. t(6) = 1.2, p = 0.28 (b, total Fos); t(6) = 8.6, p < 0.0001 (b, in PKC-δ+); t(6) = 5.9, p = 0.0011 (b, in PKC-δ–); t(6) = 6.4, p = 0.0007 (b, Fos/PKC-δ); t(6) = 9.8, p < 0.0001 (b, PKC-δ/Fos); t(6) = 3.1, p = 0.021 (c, total Fos); t(6) = 5.8, p < 0.0012 (c, in PKC-δ+); t(6) = 12.7, p < 0.0001 (c, in PKC-δ–); t(6) = 8.9, p < 0.0001 (c, Fos/PKC-δ); t(6) = 11.5, p < 0.0001 (c, PKC-δ/Fos); t(6) = 1.5, p = 0.20 (d, total Fos); t(6) = 3.9, p < 0.0076 (d, in PKC-δ+); t(6) = 4.4, p = 0.0045 (d, in PKC-δ–); t(6) = 3.4, p = 0.0014 (d, Fos/PKC-δ); t(6) = 12.5, p < 0.0001 (d, PKC-δ/Fos). * p < 0.05, ** p < 0.01, *** p < 0.001.
Supplementary Figure 3 Taste sensitivity, drinking and mounting behaviors
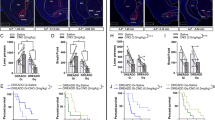
a-b, Taste sensitivity is not affected by hM4Di-DREADDs silencing of CEl PKC-δ+ neurons. Mice in a lickometer with two bottles (a): one contains water and the other contains quinine solution (see concentrations in (b)). Bitter preference was defined by the number of licks of quinine solution divided by the total number of licks on both bottles. Mice were water-deprived for 24 hours and injected with CNO (5 mg/kg) 20 min before the drinking test. n = 7 mice expressing control protein and n = 5 mice expressing hM4Di. Two-way ANOVA, F(1, 35) = 0.051, p = 0.82. c-g, Photo-stimulation of mice expressing ChR2 in CEl PKC-δ+ neurons reduces the number of licks (binned in 2 min) in the first few min of the drinking test (c) in 24 h water deprived mice. 5 Hz, 10 ms 473 nm laser pulse. Two-way ANOVA (F(1, 130) = 4.67, p = 0.033)) with post-hoc Bonferroni t-test indicated a significant difference between animals expressing with ChR2 and controls expressing hrGFP. Total number of licks during the 20 min test is not significantly reduced (d). Total amount of water intake is reduced in animals with photostimulation (e). Cumulative licks (f) and cumulative licks normalized to total licks (g) during the test. n = 8 mice expressing control protein hrGFP and n = 7 mice expressing ChR2. Unpaired t-test, t(13) = 1.13, p = 0.28 (d); t(13) = 2.42, p = 0.031 (e). h. Feeding inhibition by photo activation of CEl PKC-δ+ neurons is not rescued when drinking water is available. n = 5 animals in each group. One-way ANOVA (F(2, 14) = 99.5, p < 0.0001) with post-hoc Bonferroni t-test. i. Mounting towards a female intruder mouse (red dashed outline) was not obviously affected by photostimulation of the male mice expressing ChR2 (yellow dashed outline). Right, latency to stop mounting in response to control light (571 nm) or 473 nm light stimulation. 10 sec 5 Hz, 10 ms light pulses were triggered 1-2 seconds after mounting was started. n = 10 (control) and 8 (activation) trial events from 3 animals. Unpaired t-test, t(16) = 0.429, p = 0.67. Box plots (d, e, i) show mean (+), median, quartiles (boxes), and range (whiskers). Values in (b, c, h) are means ± s.e.m.. n.s., not significant; * p < 0.05, *** p < 0.001.
Supplementary Figure 4 Freezing, place preference and conditioned taste aversion of activating CEl PKC-δ+ neurons
a-b. The level of freezing (a) and automatically scored motion index (b) are not significantly affected by light activation. 473 nm, 5 Hz, 10 ms laser pulses were delivered for 1 min. n = 18 stimulation events from 11 animals. Unpaired t-test, t(34) = 1.21, p = 0.24 (a); t(34) = 0.342, p = 0.73 (b). c-e. Place preference is not affected by photo activating CEl PKC-δ+ neurons in the 3-chamber place preference assay. Mice received light stimulation (20 min of 473 nm, 5 Hz, 10 ms light pulses) either in chamber A or chamber B (c). Percent of time that the mice spent in each chamber on day 1 pre-test (d) or day 4 post-test (e) following training on days 2 and 3 is not affected. hrGFP, n = 8 animals, ChR2, n = 10 animals. Two-way ANOVA, F(1, 48) = 0.005, p = 0.98 (d); F(1, 48) = 0, p = 1.0 (e). f, Three groups of 24 h fasted mice were fed with novel tasting food pellets for 20 min on day 1. Group A mice (gray) expressing control protein (mCherry) and group B mice (red) expressing the activating DREADD hM3Dq-mCherry in their CEl PKC-δ+ neurons were intraperitoneal injected with CNO (2 mg/kg) after feeding; positive control group C mice (blue) expressing hrGFP in CEl PKC-δ+ neurons were intraperitoneal injected with LiCl (150 mg/kg) after feeding. All mice were then put back into their home cage with access to food for 24 h, and then food was deprived on day 2. Feeding was tested again on day 3 and day 7 without CNO injection. hM3Dq pharmacogenetic activation of CEl PKC-δ+ neurons inhibits food intake (day 9, CNO, 2 mg/kg injected 20 min before test), consistent with the results of optogenetic activation (Fig. 4b). Positive control mice (blue) show a strong reduction in food intake on day 3. Food intake on day 3 (test) is not decreased in mice expressing hM3Dq (red), indicating that no conditioned taste aversion has occurred. Increased feeding in these mice compared with control mice expressing mCherry (gray) may reflect the slow wash-out kinetics of CNO, resulting in decreased feeding during the 24 h ad libitum feeding recovery period on day 2. Consequently, on the test day (day 3), these mice may be hungrier than mCherry expressing mice, resulting in increased feeding. n = 6 animals in each group. Two-way ANOVA (F(2, 15) = 13.5, p = 0.0004) with post-hoc Bonferroni t-test (tests on day 1 and day 3); Two-way ANOVA (F(1, 10) = 9.76, p = 0.011) with post-hoc Bonferroni t-test (tests on day 7 and day 9). Values are means ± s.e.m.. * p < 0.05, ** p < 0.01, *** p < 0.001.
Supplementary Figure 5 Monosynaptic upstream inputs of CEl PKC-δ+ neurons revealed by Rabies tracing
a. Dense labeling of monosynaptic inputs to CEl PKC-δ+ neurons (GFP expressing neurons) were observed in several ipisilateral brain regions as illustrated in the diagrams. bar, 400 μm. Arrows point to high-magnification images of the corresponding regions. b. Table showing quantification of the number of GFP labeled neurons in each of these brain regions. Note that sparse labelling was also observed in several other brain regions (such as contralateral insula) that are not included here. Pir, piriform cortex; PLCo, posterolateral cortical amygdaloid area; PVT, paraventricular thalamic nucleus; Ent, Entorhinal cortex; CA1, field CA1 of the hippocampus; PoT, posterior thalamic nucleus group. n = 3 - 6 sections. Values are means ± s.e.m.
Supplementary Figure 6 Upstream inputs of CEl PKC-δ+ neurons that are activated by CCK and LiCl
a. Diagrams illustrate alternative pathways for convergent activation of CEl PKC-δ+ neurons by anorexigenic agents. Red circles indicate first-order pre-synaptic inputs to PKC-δ+ neurons. b-d. Ratio of observed number of overlapping neurons (GFP+ and c-Fos+) divided by the number of c-Fos neurons predicted by random chance (calibrated by the number of c-Fos+ neurons and GFP+ neurons divided by the total number of neurons in the nucleus). A ratio value larger than 1 means the overlapping is higher than random overlapping caused by increased number of c-Fos+ and GFP+ neurons. n = 6 sections from two injection experiments in each group. Unpaired t-test. t(10) = 3.37, p = 0.0056 ((d) saline vs CCK), t(10) = 2.57, p = 0.028 ((d) saline vs LiCl), t(10) = 4.14, p = 0.0014 ((e) saline vs CCK), t(10) = 3.14, p = 0.011 ((e) saline vs LiCl), t(10) = 1.08, p = 0.30 ((f) saline vs CCK), t(10) = 2.66, p = 0.024 ((f) saline vs CCK). Values are means ± s.e.m.. e-g. Monosynaptic rabies-traced (GFP labeled neurons) and c-Fos expressing neurons in LPB (a), BLA (b), and insula (c), in mice intraperitoneal injected with saline, CCK, or LiCl.
Supplementary Figure 7 Upstream inputs to CEl PKC-δ+ neurons that are activated by bitter tastant
a-o. c-Fos expression and quantification in LPB (a-e), BLA (f-j), and insula (k-o) in animals that received oral infusion of water, quinine solution, or sucrose solution 1-1.5 hrs prior to sacrifice . The ratio (d, i, n) indicates the observed number of double-labeled cells relative to the predicted number based on random overlap, and is calculated by dividing the actual counted number of overlapping neurons (GFP+ and c-Fos+) by the number of c-Fos neurons predicted by random chance (calibrated by the number of c-Fos+ neurons and GFP+ neurons divided by the total number of neurons in the nucleus). Values are means ± s.e.m.. For (b-d) n = 5 (H2O), 8 (quinine), 6 (sucrose) sections from two injection experiments in each group. One-way ANOVA (F(2, 16) = 15.8, p = 0.0002 (b); F(2, 16) = 8.99, p = 0.0024 (c); F(2, 16) = 3.64, p = 0.049 (d)) with post-hoc Bonferroni t-test. For (g-i) n = 8 (H2O), 8 (quinine), 5 (sucrose) sections from two injection experiments in each group. One-way ANOVA (F(2, 18) = 16.1, p < 0.0001 (g); F(2, 18) = 26.9, p < 0.0001 (h); F(2, 18) = 3.90, p = 0.0011 (i)) with post-hoc Bonferroni t-test. For (l-m) n = 6 (H2O), 5 (quinine), 6 (sucrose) sections from two injection experiments in each group. One-way ANOVA (F(2, 14) = 49.9, p < 0.0001 (l); F(2, 14) = 10.9, p = 0.0014 (m); F(2, 14) = 10.9, p = 0.045 (n)) with post-hoc Bonferroni t-test. n.s., not significant, * p< 0.05, ** p < 0.01, *** p < 0.001.
Supplementary Figure 8 Monosynaptic inputs and outputs of CEl PKC-δ+ neurons
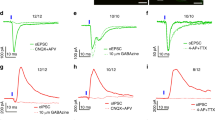
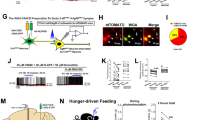
a. Fluorescent images of anti-PKC-δ antibody staining and tdTomato expression after crossing PKC-δ-Cre transgenic mice with AI14 Cre-reporter mice (Allen Institute). The quantification of overlapping (tdTomato/PKC-δ, 0.99 ± 0.45, PKC-δ/tdTomato, 0.97 ± 1.1, means ± s.e.m.) suggests that tdTomator expression can be used to identify PKC-δ expression reliably. b. Fluorescence of tdTomato is used to prospectively identify PKC-δ+ (arrow) or PKC-δ– (arrowhead) neurons in live brain slices for recording. c. Fluorescent nerve terminals in acute amygdala slices derived from ChR2-EYFP neurons in insula or LPB, infected with a non-Cre-dependent AAV. d-e. When CEl PKC-δ+ neurons were voltage-clamped at -70 mV, a monosynaptic excitatory postsynaptic current (EPSC) was triggered by a 2 ms 473 nm light pulse (indicated by blue dots) that activated the nerve terminals from insula (d) or LPB (e). The EPSCs were blocked by bath application of 20 μM CNQX. f. Amplitude of light pulse triggered EPSCs. g. Latency of EPSCs following initiation of the light pulse. n = 16 PKC-δ+ neurons from 3 animals expressing ChR2-EYFP in IC, n = 15 neurons from 2 animals expressing ChR2-EYFP in LPB. Box plots show mean (+), median, quartiles (box), and range (whisker). h. Strong fluorescent terminals from CEl PKC-δ+ neurons expressing ChR2-EYFP were observed in CEA, BNST, and LPB. i. Whole-cell patch clamp recordings in CEA PKC-δ– neurons reveal a large inhibitory postsynaptic current (IPSC) in response to light pulse stimulation of PKC-δ+ neurons (blue dots), while neurons in CEm have small IPSCs. j. Ventral BNST contains mixed neurons that show large IPSCs or small IPSCs in response to optogenetic stimulation of ChR2-expressing terminals projecting from PKC-δ+ neurons in CEl. k-l. Amplitude and latency of IPSCs in CEl and CEm. n = 5 neurons in CEl, n = 6 neurons in CEm. m-n. Amplitude and latency of large and small IPSCs in vBNST. n = 4 neurons for both large and small IPSC. Box plots show mean (+), median, quartiles, and range. Note: 1 out of 6 neurons tested in LPB showed a small IPSC (< 10 pA) in response to a light pulse, 5 out of 6 neurons showed no response. aco, anterior commissure; CEm, medial part of central amygdala; scp, superior cerebral peduncle; MPB, medial parabrachial nucleus.
Supplementary Figure 9 Cre-out eNpHR3.0 expression and quantitative relationship of expression to food intake
a. Representative histology of Cre-out eNpHR3.0-EYFP expression in PKC-δ-Cre mice. b. The reduction in food intake caused by optogenetic inhibition of CEA PKC-δ– neurons is positively correlated with the level of eNpHR3.0 expression in CEl as measured by EYFP fluorescence intensity. Pearson correlation test, Pearson r = -0.71, p = 0.032. Food intake change was calculated as (Food intake (silencing) - Food intake (control))/Food intake (control) x 100. The intensity of EYFP fluorescence was measured using ImageJ (NIH). Note: (a) indicates that there are always a few neurons in BLA and CEm expressing eNpHR3.0, even when a relatively small volume of virus is injected (50-100 nl) to restrict infection to CEl. These small volumes will result in fewer CEl neurons expressing eNpHR3.0, perhaps explaining why the level of feeding inhibition (b) is not as strong as that observed by activation of PKC-δ+ neurons (where larger volumes of virus can be used because of the specificity afforded by Cre recombination in PKC-δ+ neurons; Fig. 4b).
Supplementary Figure 10 Activation of Tac2 or CRF neurons in CEl does not inhibit feeding
a-b. Expression of Tac2 (a) or CRF (b) and PKC-δ (antibody staining) in CEl. The expression of Tac2 or CRF was identified by tdTomato expression after crossing the mice with Ai14 Cre-reporter mice. c. Quantification of the percentage of Tac2, CRF, and PKC-δ+ neurons in CEl, and their degree of overlap. n = 6 to 10 brain section from 3 animals in each group. Values are means ± s.e.m.. d-g. Food intake during photostimulation of Tac2-Cre mice (d-e) and CRF-Cre mice expressing ChR2 (f-g). No significant change was observed. For (d-e), n = 5 animals expressing control protein hrGFP, n = 7 animals expressing ChR2; unpaired t-test, t(10) = 0.634, p = 0.54 (d); t(10) = 0.304, p = 0.77 (e). For (f-g), n = 5 animals in each group; unpaired t-test, t(8) = 0.335, p = 0.75 (f); t(8) = 0.479, p = 0.64 (g). Bar in (a) also applies to (b). Box plots show mean (+), median, quartiles, and range.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–10 (PDF 5212 kb)
Supplementary Methods Checklist
(PDF 554 kb)
Light activation of CEl PKC-δ+ neurons inhibits feeding.
This video shows a 24 h fasted mice approaches and eats a food pellet in its home cage, its feeding is interrupted in a few seconds after photo stimulation of the CEl PKC–δ+ neurons that express ChR2. (WMV 2083 kb)
Rights and permissions
About this article
Cite this article
Cai, H., Haubensak, W., Anthony, T. et al. Central amygdala PKC-δ+ neurons mediate the influence of multiple anorexigenic signals. Nat Neurosci 17, 1240–1248 (2014). https://doi.org/10.1038/nn.3767
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nn.3767
This article is cited by
-
Cocaine and amphetamine regulated transcript (CART) mediates sex differences in binge drinking through central taste circuits
Neuropsychopharmacology (2024)
-
Rhesus infant nervous temperament predicts peri-adolescent central amygdala metabolism & behavioral inhibition measured by a machine-learning approach
Translational Psychiatry (2024)
-
Amygdala electrical stimulation for operant conditioning in rat navigation
Biomedical Engineering Letters (2024)
-
Activation patterns in male and female forebrain circuitries during food consumption under novelty
Brain Structure and Function (2024)
-
Butterflies in the gut: the interplay between intestinal microbiota and stress
Journal of Biomedical Science (2023)