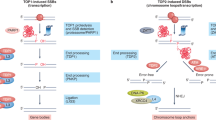
Abstract
It has been unclear whether ischemic stroke induces neurogenesis or neuronal DNA rearrangements in the human neocortex. Using immunohistochemistry; transcriptome, genome and ploidy analyses; and determination of nuclear bomb test–derived 14C concentration in neuronal DNA, we found neither to be the case. A large proportion of cortical neurons displayed DNA fragmentation and DNA repair a short time after stroke, whereas neurons at chronic stages after stroke showed DNA integrity, demonstrating the relevance of an intact genome for survival.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Spalding, K.L., Bhardwaj, R.D., Buchholz, B.A., Druid, H. & Frisen, J. Cell 122, 133–143 (2005).
Bhardwaj, R.D. et al. Proc. Natl. Acad. Sci. USA 103, 12564–12568 (2006).
Jin, K. et al. Proc. Natl. Acad. Sci. USA 103, 13198–13202 (2006).
Kreuzberg, M. et al. Exp. Neurol. 226, 90–99 (2010).
Leker, R.R. et al. Stroke 38, 153–161 (2007).
Li, W. et al. J. Cereb. Blood Flow Metab. 26, 181–198 (2006).
Li, P. et al. Antioxid. Redox Signal. 14, 1905–1918 (2011).
Suberbielle, E. et al. Nat. Neurosci. 16, 613–621 (2013).
Rausch, T. et al. Cell 148, 59–71 (2012).
Bergmann, O. et al. Science 324, 98–102 (2009).
Spalding, K.L. et al. Cell 153, 1219–1227 (2013).
Bergmann, O. et al. Neuron 74, 634–639 (2012).
Brown, T.A. & Southon, J.R. Nucl. Instrum. Methods Phys. Res. B 123, 208–213 (1997).
Klempin, F., Kronenberg, G., Cheung, G., Kettenmann, H. & Kempermann, G. PLoS ONE 6, e25760 (2011).
Benavides, S.H., Monserrat, A.J., Farina, S. & Porta, E.A. Arch. Gerontol. Geriatr. 34, 219–231 (2002).
Osman, A.M., Porritt, M.J., Nilsson, M. & Kuhn, H.G. Stroke 42, 3559–3565 (2011).
Ohira, K. et al. Nat. Neurosci. 13, 173–179 (2010).
Salehpour, M., Hakansson, K. & Possnert, G. Nucl. Instrum. Methods Phys. Res. B. 294, 97–103 (2013).
Hua, Q., Zoppi, U., Williams, A. & Smith, A. Nucl. Instrum. Methods Phys. Res. B. 223–224, 284–292 (2004).
Santos, G.M., Southon, J.R., Griffin, S., Beaupre, S.R. & Druffel, E.R.M. Nucl. Instrum. Methods Phys. Res. B. 259, 293–302 (2007).
Speit, G. & Hartmann, A. Methods Mol. Biol. 314, 275–286 (2006).
Borgström, E., Lundin, S. & Lundeberg, J. PLoS ONE 6, e19119 (2011).
Trapnell, C., Pachter, L. & Salzberg, S.L. Bioinformatics 25, 1105–1111 (2009).
Ge, H. et al. Bioinformatics 27, 1922–1928 (2011).
Li, H. & Durbin, R. Bioinformatics 25, 1754–1760 (2009).
Xie, C. & Tammi, M.T. BMC Bioinformatics 10, 80 (2009).
Acknowledgements
This study was supported by grants from the European Research Council, the Swedish Research Council, The Swedish Heart and Lung Foundation, the Swedish Cancer Society, the Karolinska Institute, Tobias Stiftelsen, AFA Försäkringar, the Strategic Research Programs in Stem Cells and Regenerative Medicine at Karolinska Institutet (StratRegen) and Lund University (StemTherapy), the European Union (project TargetBraIn, 279017), Torsten Söderbergs Stiftelse, and Knut och Alice Wallenbergs Stiftelse. H.B.H. was funded by a grant from the German Research Foundation (Hu 1961/1-1). T.H. was supported by the National Brain Research Programme (NAP), Hungary.
Author information
Authors and Affiliations
Contributions
H.B.H., O.B. and J.F. designed the study and wrote the manuscript. M.S. and G.P. performed the AMS measurements. A.R., T.C., L.C., T.H., G.M., E.E., Z.K., O.L. and S.S. procured post-mortem material and obtained clinical and pathological data. J.T., Z.K. and O.L. performed stroke experiments in mice. B.W.S., B.S., J.L., P.S. and A.D. performed sequencing of RNA and DNA samples. S.B. and S.Z. performed mathematical modeling of cell turnover rates. H.B.H., O.B., E.L., C.S. and L.S. performed experiments and collected data. All authors approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 FACS-based strategy to isolate neuronal and non-neuronal nuclei.
Neuronal nuclei were labeled with an isotype control (a) or an antibody against Fox3 (NeuN-mab) (b) and isolated through subsequent flow cytometry sorting. Re-analysis of sorted NeuN-positve (c) and NeuN-negative nuclei (d) reveals a sorting purity of neuronal, and non-neuronal populations exceeding 95%. DNA-analysis of sorted NeuN-positive nuclei revealed that almost all nuclei were diploid (>99.5%), and no difference in DNA content in control (e) compared to stroke tissue (f and g-merge).
Supplementary Figure 2 NeuN is a specific marker for neurons in the human cortex after ischemic stroke.
(a) All NeuN-positive nuclei in the penumbra of cortical ischemic strokes expressed the pan-neural markers HuD and MAP-2, validating NeuN as a reliable marker for the isolation of neuronal nuclei through FACS both from healthy tissue (n = 3), and from ischemic stroke cortex (n = 3); scale bar indicates 20 μm. (b, c and d) FACS analysis of cortical nuclei demonstrated that almost all NeuN-postive nuclei were immunoreactive for the neuronal marker HuD. Of note, HuD showed also some immunoreactivity to the oligodentrocyte lineage (NeuN-negative/HuD-positve; see Bergmann, O. et al. 2013. Neuron 74:634-9). (e and f) Transcriptional analysis using qRT-PCR revealed that NeuN-positve nuclei showed a high enrichment for neuronal marker genes (FOX3 and MAP2), whereas NeuN-negative sorted nuclei are enriched for non-neuronal marker genes (Iba4, GFAP, CNP, Sox10 and vWf). Error bars are mean ± SD.
Supplementary Figure 3 Analysis of copy number variations in neurons in the post-ischemic cortex.
Comparison of copy number variations (CNVs) of neurons in healthy control cortex and post-ischemic cortex of the same patient analyzed by whole genome sequencing of DNA (reproduced in altogether three patients). There were no significant changes between lesion (upper panel) and healthy tissue (lower panel), and no evidence of CNVs in cortical neurons after ischemic stroke.
Supplementary Figure 4 Neuronal DNA repair is not linked to apoptosis.
(a) DNA repair marker γH2AX (green) does not co-label with cleaved Caspase-3 (red) as a marker for apoptosis in the penumbra of a subacute cortical ischemic stroke lesion (n = 3). Arrow depicts a neuron being positive for γH2AX, but not for cleaved Caspase-3. Of note, arrowheads depict lipofuscin-induced autofluorescence. Scale bar indicates 20μm. γH2AX (green; b) and APE-1 (c) does not co-label with NeuN-immunoreactivity in healthy control cortex (n = 3). Scale bar indicates 20 μm.
Supplementary Figure 5 Number of splice junctions found through RNA sequencing.
Number of splice junctions found with each 5% increment of reads used. The increase in number of splice junctions found diminishes fast with increasing amount of reads used, almost reaching a plateau at 100%. This indicates we have reached saturation in sequencing depth with regards to finding splice junctions in the stroke samples that could lead to the discovery of fusion genes.
Supplementary Figure 6 Genomic 14C concentrations of cortical cells after ischemic stroke.
(a) 14C concentrations of neuronal DNA in acute ischemic cortical lesions (n = 2; light green dots, 3 and 7 days post stroke) were not significantly different from neurons in chronic cortical lesions (red dots), indicating that there is also no generation of new neurons early after stroke the. (b) Genomic 14C concentrations of non-neuronal cells in the control cortex in acute ischemic cortical lesions (dark green dots, 3 and 7 days post stroke) were not significantly different from non-neurons in chronic cortical lesions (orange dots). Error bars indicate two standard deviations in 14C concentration in the respective DNA sample.
Supplementary Figure 7 DCX is expressed in mature neurons in acute cortical ischemic lesions in humans.
(a) The post-mitotic neuronal marker NeuN (red) is already co-expressed with the putative progenitor marker DCX (green) 7 days after infarction, indicating an up-regulation of DCX in pre-existing neurons in humans. Arrows depict the DCX/NeuN positive neuron. Scale bar indicates 20μm (a). (b) show a high magnification images of the white inset; scale bar indicates 10μm.
Supplementary Figure 8 Absence of lipofuscin-negative neurons.
Quantification of neuronal lipofuscin in control cerebral cortex and post-ischemic cerebral cortex (see method section for detailed description of lipufuscin grading). Consistent with the 14C data (Fig. 3) and the lack of new neurons generated upon and surviving cortical stroke, no neurons (identified by NeuN expression) devoid of the age pigment lipofuscin were observed (>1,000 neurons in the penumbra of the ischemic lesion were analyzed in each of the patients 3, 5, 6 and 7, see suppl. table 1). Interestingly, the proportion of neurons with abundant lipofuscin deposits was lower in the penumbra adjacent to the stroke lesion core than in healthy cortex tissue, raising the possibility that high lipofuscin levels may represent a predisposing factor for ischemic neuronal death.
Supplementary Figure 9 Early generation of DCX positive cells after cortical ischemic lesion in mice.
(a) Location of ischemic lesion in cerebral cortex and corpus callosum as shown by NeuN immunohistochemistry. (b-d) Distribution of DCX+, BrdU+, and DCX+/BrdU+ cells in the periinfarct cortex and corpus callosum at 2 weeks after stroke onset (n = 7); scale bar indicates 100 μm (a-d). (e-g) Boxed regions in b-d at higher magnification showing both DCX+/BrdU- (arrows) and DCX+/BrdU+ cells (arrow-heads); scale bar indicates 20 μm. A few DCX-positive neuroblasts were located along the corpus callosum, but no such cells were found in the peri-infarct cortex. BrdU-DCX double-immunostaining showed that DCX-positive cells distributed in the peri-infarct cortex or migrating along the corpus callosum expressed BrdU. No DCX-NeuN double-positive cells were detected in the peri-infarct cortex at 2 weeks, arguing against the possibility that mature, pre-existing neurons expressed DCX at this time-point. In the 4 weeks group (n = 3) no DCX-positive cells were found in the peri-infarct cortex or striatum (data not shown). We also could not detect any NeuN-BrdU double-positive cells in these areas, providing evidence that the newborn DCX-positive neuroblasts, observed in the peri-infarct cortex in the 2 weeks group, had died after 4 weeks and not differentiated to mature neurons.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–9 (PDF 18372 kb)
Supplementary Table 1: List of analyzed human subjects with cortical ischemic stroke.
Description of the included patients, with details on stroke and other medical conditions. (XLS 48 kb)
Supplementary Table 2: List of predicted fusion genes using FusionMap.
RNA fusion gene candidates identified through RNA sequencing in 6 patients with cortical ischemic stroke and 5 healthy controls. Batch 1 and 2 refer to samples included in sequencing round 1 and 2, Unique Cutting Positions is the same as unique mapping location of fusion read, Seed Count specifies the count of reads that were considered a seed read, Rescued Count specifies the count of reads that were considered a rescued read, Chr1 and Chr2 is the chromosomes represented by the first and second gene of the fusion, Position 1 and 2 specifies the cutting position of the first and second gene for the fusion, KnownGene1 and 2 specifies the first and the second gene name, KnownTranscript1 and 2 specifies the known transcript names of the first and second transcript for the fusion, FusionJunctionSequence specifies the full sequence of the detected fusion junction. (XLSX 44 kb)
Rights and permissions
About this article
Cite this article
Huttner, H., Bergmann, O., Salehpour, M. et al. The age and genomic integrity of neurons after cortical stroke in humans. Nat Neurosci 17, 801–803 (2014). https://doi.org/10.1038/nn.3706
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nn.3706
This article is cited by
-
Emergence of task-related spatiotemporal population dynamics in transplanted neurons
Nature Communications (2023)
-
Subventricular zone cytogenesis provides trophic support for neural repair in a mouse model of stroke
Nature Communications (2023)
-
Effect of WiFi signal exposure in utero and early life on neurodevelopment and behaviors of rats
Environmental Science and Pollution Research (2023)
-
IRES-mediated Wnt2 translation in apoptotic neurons triggers astrocyte dedifferentiation
npj Regenerative Medicine (2022)
-
Evidence for postnatal neurogenesis in the human amygdala
Communications Biology (2022)