Abstract
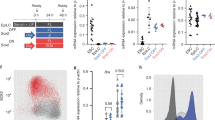
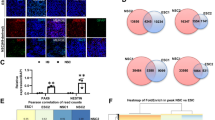
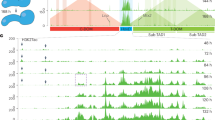
Hox genes controlling motor neuron subtype identity are expressed in rostrocaudal patterns that are spatially and temporally collinear with their chromosomal organization. Here we demonstrate that Hox chromatin is subdivided into discrete domains that are controlled by rostrocaudal patterning signals that trigger rapid, domain-wide clearance of repressive histone H3 Lys27 trimethylation (H3K27me3) polycomb modifications. Treatment of differentiating mouse neural progenitors with retinoic acid leads to activation and binding of retinoic acid receptors (RARs) to the Hox1–Hox5 chromatin domains, which is followed by a rapid domain-wide removal of H3K27me3 and acquisition of cervical spinal identity. Wnt and fibroblast growth factor (FGF) signals induce expression of the Cdx2 transcription factor that binds and clears H3K27me3 from the Hox1–Hox9 chromatin domains, leading to specification of brachial or thoracic spinal identity. We propose that rapid clearance of repressive modifications in response to transient patterning signals encodes global rostrocaudal neural identity and that maintenance of these chromatin domains ensures the transmission of positional identity to postmitotic motor neurons later in development.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Ensini, M., Tsuchida, T.N., Belting, H.G. & Jessell, T.M. The control of rostrocaudal pattern in the developing spinal cord: specification of motor neuron subtype identity is initiated by signals from paraxial mesoderm. Development 125, 969–982 (1998).
Dasen, J.S., Tice, B.C., Brenner-Morton, S. & Jessell, T.M. A Hox regulatory network establishes motor neuron pool identity and target-muscle connectivity. Cell 123, 477–491 (2005).
Jung, H. et al. Global control of motor neuron topography mediated by the repressive actions of a single hox gene. Neuron 67, 781–796 (2010).
Wu, Y., Wang, G., Scott, S.A. & Capecchi, M.R. Hoxc10 and Hoxd10 regulate mouse columnar, divisional and motor pool identity of lumbar motoneurons. Development 135, 171–182 (2008).
Liu, J.P., Laufer, E. & Jessell, T.M. Assigning the positional identity of spinal motor neurons: rostrocaudal patterning of Hox-c expression by FGFs, Gdf11, and retinoids. Neuron 32, 997–1012 (2001).
Simeone, A. et al. Sequential activation of HOX2 homeobox genes by retinoic acid in human embryonal carcinoma cells. Nature 346, 763–766 (1990).
Papalopulu, N., Lovell-Badge, R. & Krumlauf, R. The expression of murine Hox-2 genes is dependent on the differentiation pathway and displays a collinear sensitivity to retinoic acid in F9 cells and Xenopus embryos. Nucleic Acids Res. 19, 5497–5506 (1991).
Akasaka, T. et al. Mice doubly deficient for the polycomb group genes Mel18 and Bmi1 reveal synergy and requirement for maintenance but not initiation of Hox gene expression. Development 128, 1587–1597 (2001).
Beuchle, D., Struhl, G. & Muller, J. Polycomb group proteins and heritable silencing of Drosophila Hox genes. Development 128, 993–1004 (2001).
Chambeyron, S. & Bickmore, W.A. Chromatin decondensation and nuclear reorganization of the HoxB locus upon induction of transcription. Genes Dev. 18, 1119–1130 (2004).
Boyer, L.A. et al. Polycomb complexes repress developmental regulators in murine embryonic stem cells. Nature 441, 349–353 (2006).
Bernstein, B.E. et al. A bivalent chromatin structure marks key developmental genes in embryonic stem cells. Cell 125, 315–326 (2006).
Rinn, J.L. et al. Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell 129, 1311–1323 (2007).
Soshnikova, N. & Duboule, D. Epigenetic temporal control of mouse Hox genes in vivo. Science 324, 1320–1323 (2009).
Kashyap, V. & Gudas, L.J. Epigenetic regulatory mechanisms distinguish retinoic acid–mediated transcriptional responses in stem cells and fibroblasts. J. Biol. Chem. 285, 14534–14548 (2010).
Peljto, M., Dasen, J.S., Mazzoni, E.O., Jessell, T.M. & Wichterle, H. Functional diversity of ESC-derived motor neuron subtypes revealed through intraspinal transplantation. Cell Stem Cell 7, 355–366 (2010).
Nordström, U., Maier, E., Jessell, T.M. & Edlund, T. An early role for WNT signaling in specifying neural patterns of Cdx and Hox gene expression and motor neuron subtype identity. PLoS Biol. 4, e252 (2006).
Bel-Vialar, S., Itasaki, N. & Krumlauf, R. Initiating Hox gene expression: in the early chick neural tube differential sensitivity to FGF and RA signaling subdivides the HoxB genes in two distinct groups. Development 129, 5103–5115 (2002).
Mahony, S. et al. Ligand-dependent dynamics of retinoic acid receptor binding during early neurogenesis. Genome Biol. 12, R2 (2011).
Shimizu, T., Bae, Y.K. & Hibi, M. Cdx-Hox code controls competence for responding to Fgfs and retinoic acid in zebrafish neural tissue. Development 133, 4709–4719 (2006).
Skromne, I., Thorsen, D., Hale, M., Prince, V.E. & Ho, R.K. Repression of the hindbrain developmental program by Cdx factors is required for the specification of the vertebrate spinal cord. Development 134, 2147–2158 (2007).
Chawengsaksophak, K., de Graaff, W., Rossant, J., Deschamps, J. & Beck, F. Cdx2 is essential for axial elongation in mouse development. Proc. Natl. Acad. Sci. USA 101, 7641–7645 (2004).
Taylor, J.K., Levy, T., Suh, E.R. & Traber, P.G. Activation of enhancer elements by the homeobox gene Cdx2 is cell line specific. Nucleic Acids Res. 25, 2293–2300 (1997).
Wichterle, H., Lieberam, I., Porter, J.A. & Jessell, T.M. Directed differentiation of embryonic stem cells into motor neurons. Cell 110, 385–397 (2002).
Mazzoni, E.O. et al. Embryonic stem cell–based mapping of developmental transcriptional programs. Nat. Methods 8, 1056–1058 (2011).
Eskeland, R. et al. Ring1B compacts chromatin structure and represses gene expression independent of histone ubiquitination. Mol. Cell 38, 452–464 (2010).
Krogan, N.J. et al. The Paf1 complex is required for histone H3 methylation by COMPASS and Dot1p: linking transcriptional elongation to histone methylation. Mol. Cell 11, 721–729 (2003).
Surface, L.E., Thornton, S.R. & Boyer, L.A. Polycomb group proteins set the stage for early lineage commitment. Cell Stem Cell 7, 288–298 (2010).
Montgomery, N.D. et al. The murine polycomb group protein Eed is required for global histone H3 lysine-27 methylation. Curr. Biol. 15, 942–947 (2005).
Shen, X. et al. EZH1 mediates methylation on histone H3 lysine 27 and complements EZH2 in maintaining stem cell identity and executing pluripotency. Mol. Cell 32, 491–502 (2008).
Pasini, D., Bracken, A.P., Hansen, J.B., Capillo, M. & Helin, K. The polycomb group protein Suz12 is required for embryonic stem cell differentiation. Mol. Cell Biol. 27, 3769–3779 (2007).
Rousso, D.L., Gaber, Z.B., Wellik, D., Morrisey, E.E. & Novitch, B.G. Coordinated actions of the forkhead protein Foxp1 and Hox proteins in the columnar organization of spinal motor neurons. Neuron 59, 226–240 (2008).
Dasen, J.S., De Camilli, A., Wang, B., Tucker, P.W. & Jessell, T.M. Hox repertoires for motor neuron diversity and connectivity gated by a single accessory factor, FoxP1. Cell 134, 304–316 (2008).
Moreno, E. & Morata, G. Caudal is the Hox gene that specifies the most posterior Drosophile segment. Nature 400, 873–877 (1999).
Niwa, H. et al. Interaction between Oct3/4 and Cdx2 determines trophectoderm differentiation. Cell 123, 917–929 (2005).
Berger, M.F. et al. Variation in homeodomain DNA binding revealed by high-resolution analysis of sequence preferences. Cell 133, 1266–1276 (2008).
Mazzoni, E.O. et al. Synergistic recruitment of transcription factors to cell type specific enhancers programs motor neuron identity. Nat. Neurosci. doi:10.1038/nn.3467 (21 July 2013).
Chambeyron, S., Da Silva, N.R., Lawson, K.A. & Bickmore, W.A. Nuclear re-organisation of the Hoxb complex during mouse embryonic development. Development 132, 2215–2223 (2005).
Liu, J.P. The function of growth/differentiation factor 11 (Gdf11) in rostrocaudal patterning of the developing spinal cord. Development 133, 2865–2874 (2006).
Bickmore, W.A., Mahy, N.L. & Chambeyron, S. Do higher-order chromatin structure and nuclear reorganization play a role in regulating Hox gene expression during development? Cold Spring Harb. Symp. Quant. Biol. 69, 251–257 (2004).
Golden, M.G. & Dasen, J.S. Polycomb repressive complex 1 activities determine the columnar organization of motor neurons. Genes Dev. 26, 2236–2250 (2012).
Montavon, T. et al. A regulatory archipelago controls Hox genes transcription in digits. Cell 147, 1132–1145 (2011).
Fullwood, M.J. et al. An oestrogen-receptor-α–bound human chromatin interactome. Nature 462, 58–64 (2009).
Wettenhall, J.M., Simpson, K.M., Satterley, K. & Smyth, G.K. affylmGUI: a graphical user interface for linear modeling of single channel microarray data. Bioinformatics 22, 897–899 (2006).
Wu, Z.J., Irizarry, R.A., Gentleman, R., Martinez-Murillo, F. & Spencer, F. A model based background adjustment for oligonucleotide expression arrays. J. Am. Stat. Assoc. 99, 909–917 (2004).
Smyth, G.K. Linear models and empirical bayes methods for assessing differential expression in microarray experiments. Stat. Appl. Genet. Mol. Biol. 3, Article3 (2004).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. Series B Stat. Methodol. 57, 289–300 (1995).
Bolstad, B.M., Irizarry, R.A., Astrand, M. & Speed, T.P. A comparison of normalization methods for high density oligonucleotide array data based on variance and bias. Bioinformatics 19, 185–193 (2003).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S.L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Guo, Y. et al. Discovering homotypic binding events at high spatial resolution. Bioinformatics 26, 3028–3034 (2010).
Acknowledgements
We thank V. Korinek (Institute of Molecular Genetics, Prague) for purified Wnt3A protein, G. Daley (Harvard Medical School) for the Cdx2 construct, M. Kyba (University of Minnesota) for sharing reagents for construction of the inducible Cdx2 ESC line and T. Jessell (Columbia University) and J. Dasen (New York University) for sharing antibodies and for constructive discussion of our findings. E.O.M. was the David and Sylvia Lieb Fellow of the Damon Runyon Cancer Research Foundation (DRG-1937-07), and this work was supported by The Leona M. and Harry B. Helmsley Charitable Trust, US National Institutes of Health grants P01 NS055923 (D.K.G. and H.W.) and R01 NS058502 and NS078097 (H.W.) and The Richard and Susan Smith Family Foundation, Chestnut Hill, Massachusetts (L.A.B.).
Author information
Authors and Affiliations
Contributions
E.O.M., S. Mahony, D.K.G. and H.W. conceived the experiments, analyzed the data and wrote the manuscript. E.O.M. generated and validated inducible cell lines and performed cell differentiations and expression analyses. S. Mahony and C.R. performed all computational and statistical analyses of genomic, expression and sequencing data. M.P. and T.P. performed and optimized caudalization experiments. S.R.T. and L.A.B. performed analysis of Prc2-null and hypomorph cell lines. S. McCuine and R.A.Y. performed ChIP-chip experiments.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 (PDF 1313 kb)
Rights and permissions
About this article
Cite this article
Mazzoni, E., Mahony, S., Peljto, M. et al. Saltatory remodeling of Hox chromatin in response to rostrocaudal patterning signals. Nat Neurosci 16, 1191–1198 (2013). https://doi.org/10.1038/nn.3490
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nn.3490
This article is cited by
-
BAF45D-binding to HOX genes was differentially targeted in H9-derived spinal cord neural stem cells
Scientific Reports (2024)
-
Generation of functional posterior spinal motor neurons from hPSCs-derived human spinal cord neural progenitor cells
Cell Regeneration (2023)
-
Sequential and directional insulation by conserved CTCF sites underlies the Hox timer in stembryos
Nature Genetics (2023)
-
Transcriptional dynamics of murine motor neuron maturation in vivo and in vitro
Nature Communications (2022)
-
A robust TDP-43 knock-in mouse model of ALS
Acta Neuropathologica Communications (2020)