Abstract
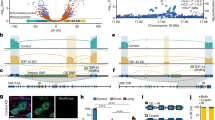
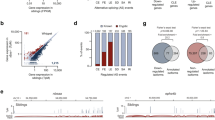
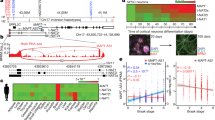
We used cross-linking and immunoprecipitation coupled with high-throughput sequencing to identify binding sites in 6,304 genes as the brain RNA targets for TDP-43, an RNA binding protein that, when mutated, causes amyotrophic lateral sclerosis. Massively parallel sequencing and splicing-sensitive junction arrays revealed that levels of 601 mRNAs were changed (including Fus (Tls), progranulin and other transcripts encoding neurodegenerative disease–associated proteins) and 965 altered splicing events were detected (including in sortilin, the receptor for progranulin) following depletion of TDP-43 from mouse adult brain with antisense oligonucleotides. RNAs whose levels were most depleted by reduction in TDP-43 were derived from genes with very long introns and that encode proteins involved in synaptic activity. Lastly, we found that TDP-43 autoregulates its synthesis, in part by directly binding and enhancing splicing of an intron in the 3′ untranslated region of its own transcript, thereby triggering nonsense-mediated RNA degradation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
References
Neumann, M. et al. Ubiquitinated TDP-43 in frontotemporal lobar degeneration and amyotrophic lateral sclerosis. Science 314, 130–133 (2006).
Arai, T. et al. TDP-43 is a component of ubiquitin-positive tau-negative inclusions in frontotemporal lobar degeneration and amyotrophic lateral sclerosis. Biochem. Biophys. Res. Commun. 351, 602–611 (2006).
Lagier-Tourenne, C., Polymenidou, M. & Cleveland, D.W. TDP-43 and FUS/TLS: emerging roles in RNA processing and neurodegeneration. Hum. Mol. Genet. 19, R46–R64 (2010).
Gitcho, M.A. et al. TDP-43 A315T mutation in familial motor neuron disease. Ann. Neurol. 63, 535–538 (2008).
Kabashi, E. et al. TARDBP mutations in individuals with sporadic and familial amyotrophic lateral sclerosis. Nat. Genet. 40, 572–574 (2008).
Sreedharan, J. et al. TDP-43 mutations in familial and sporadic amyotrophic lateral sclerosis. Science 319, 1668–1672 (2008).
Van Deerlin, V.M. et al. TARDBP mutations in amyotrophic lateral sclerosis with TDP-43 neuropathology: a genetic and histopathological analysis. Lancet Neurol. 7, 409–416 (2008).
Buratti, E. et al. Nuclear factor TDP-43 can affect selected microRNA levels. FEBS J. 277, 2268–2281 (2010).
Cooper, T.A., Wan, L. & Dreyfuss, G. RNA and Disease. Cell 136, 777–793 (2009).
Kwiatkowski, T.J. Jr. et al. Mutations in the FUS/TLS gene on chromosome 16 cause familial amyotrophic lateral sclerosis. Science 323, 1205–1208 (2009).
Vance, C. et al. Mutations in FUS, an RNA processing protein, cause familial amyotrophic lateral sclerosis type 6. Science 323, 1208–1211 (2009).
Mortazavi, A., Williams, B.A., McCue, K., Schaeffer, L. & Wold, B. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat. Methods 5, 621–628 (2008).
Ule, J. et al. CLIP identifies Nova-regulated RNA networks in the brain. Science 302, 1212–1215 (2003).
Licatalosi, D.D. et al. HITS-CLIP yields genome-wide insights into brain alternative RNA processing. Nature 456, 464–469 (2008).
Yeo, G.W. et al. An RNA code for the FOX2 splicing regulator revealed by mapping RNA-protein interactions in stem cells. Nat. Struct. Mol. Biol. 16, 130–137 (2009).
Ling, S.C. et al. ALS-associated mutations in TDP-43 increase its stability and promote TDP-43 complexes with FUS/TLS. Proc. Natl. Acad. Sci. USA 107, 13318–13323 (2010).
Zisoulis, D.G. et al. Comprehensive discovery of endogenous Argonaute binding sites in Caenorhabditis elegans. Nat. Struct. Mol. Biol. 17, 173–179 (2010).
Sephton, C.F. et al. Identification of neuronal RNA targets of TDP-43-containing ribonucleoprotein complexes. J. Biol. Chem. 286, 1204–1215 (2011).
Mili, S. & Steitz, J.A. Evidence for reassociation of RNA-binding proteins after cell lysis: implications for the interpretation of immunoprecipitation analyses. RNA 10, 1692–1694 (2004).
Buratti, E. et al. Nuclear factor TDP-43 and SR proteins promote in vitro and in vivo CFTR exon 9 skipping. EMBO J. 20, 1774–1784 (2001).
Chi, S.W., Zang, J.B., Mele, A. & Darnell, R.B. Argonaute HITS-CLIP decodes microRNA-mRNA interaction maps. Nature 460, 479–486 (2009).
Parkhomchuk, D. et al. Transcriptome analysis by strand-specific sequencing of complementary DNA. Nucleic Acids Res. 37, e123 (2009).
Pang, K.C. et al. RNAdb—a comprehensive mammalian noncoding RNA database. Nucleic Acids Res. 33, D125–D130 (2005).
Wang, E.T. et al. Alternative isoform regulation in human tissue transcriptomes. Nature 456, 470–476 (2008).
Hu, F. et al. Sortilin-mediated endocytosis determines levels of the frontotemporal dementia protein, progranulin. Neuron 68, 654–667 (2010).
Carrasquillo, M.M. et al. Genome-wide screen identifies rs646776 near sortilin as a regulator of progranulin levels in human pasma. Am. J. Hum. Genet. 87, 890–897 (2010).
Du, H. et al. Aberrant alternative splicing and extracellular matrix gene expression in mouse models of myotonic dystrophy. Nat. Struct. Mol. Biol. 17, 187–193 (2010).
Ayala, Y.M. et al. TDP-43 regulates its mRNA levels through a negative feedback loop. EMBO J. 30, 277–288 (2011).
Le Hir, H., Moore, M.J. & Maquat, L.E. Pre-mRNA splicing alters mRNP composition: evidence for stable association of proteins at exon-exon junctions. Genes Dev. 14, 1098–1108 (2000).
Wollerton, M.C., Gooding, C., Wagner, E.J., Garcia-Blanco, M.A. & Smith, C.W. Autoregulation of polypyrimidine tract binding protein by alternative splicing leading to nonsense-mediated decay. Mol. Cell 13, 91–100 (2004).
Sureau, A., Gattoni, R., Dooghe, Y., Stevenin, J. & Soret, J. SC35 autoregulates its expression by promoting splicing events that destabilize its mRNAs. EMBO J. 20, 1785–1796 (2001).
Perlick, H.A., Medghalchi, S.M., Spencer, F.A., Kendzior, R.J. Jr. & Dietz, H.C. Mammalian orthologs of a yeast regulator of nonsense transcript stability. Proc. Natl. Acad. Sci. USA 93, 10928–10932 (1996).
Baker, M. et al. Mutations in progranulin cause tau-negative frontotemporal dementia linked to chromosome 17. Nature 442, 916–919 (2006).
Cruts, M. et al. Null mutations in progranulin cause ubiquitin-positive frontotemporal dementia linked to chromosome 17q21. Nature 442, 920–924 (2006).
Fiesel, F.C. et al. Knockdown of transactive response DNA-binding protein (TDP-43) downregulates histone deacetylase 6. EMBO J. 29, 209–221 (2010).
Strong, M.J. et al. TDP43 is a human low molecular weight neurofilament (hNFL) mRNA-binding protein. Mol. Cell. Neurosci. 35, 320–327 (2007).
Bergeron, C. et al. Neurofilament light and polyadenylated mRNA levels are decreased in amyotrophic lateral sclerosis motor neurons. J. Neuropathol. Exp. Neurol. 53, 221–230 (1994).
Hutton, M. et al. Association of missense and 5′-splice-site mutations in tau with the inherited dementia FTDP-17. Nature 393, 702–705 (1998).
MacDonald, M.E. et al. A novel gene containing a trinucleotide repeat that is expanded and unstable on Huntington's disease chromosomes. Cell 72, 971–983 (1993).
Schwab, C., Arai, T., Hasegawa, M., Yu, S. & McGeer, P.L. Colocalization of transactivation-responsive DNA-binding protein 43 and huntingtin in inclusions of Huntington disease. J. Neuropathol. Exp. Neurol. 67, 1159–1165 (2008).
Valdmanis, P.N. et al. A mutation that creates a pseudoexon in SOD1 causes familial ALS. Ann. Hum. Genet. 73, 652–657 (2009).
Birve, A. et al. A novel SOD1 splice site mutation associated with familial ALS revealed by SOD activity analysis. Hum. Mol. Genet. 19, 4201–4206 (2010).
Mercado, P.A., Ayala, Y.M., Romano, M., Buratti, E. & Baralle, F.E. Depletion of TDP 43 overrides the need for exonic and intronic splicing enhancers in the human apoA-II gene. Nucleic Acids Res. 33, 6000–6010 (2005).
Dreumont, N. et al. Antagonistic factors control the unproductive splicing of SC35 terminal intron. Nucleic Acids Res. 38, 1353–1366 (2009).
Lin, S., Coutinho-Mansfield, G., Wang, D., Pandit, S. & Fu, X.D. The splicing factor SC35 has an active role in transcriptional elongation. Nat. Struct. Mol. Biol. 15, 819–826 (2008).
Tollervey, J.R. et al. Characterizing the RNA targets and position-dependent splicing regulation by TDP-43. Nat. Neurosci. advance online publication, 10.1038/nn.2778 (27 February 2011).
Xu, Y.F. et al. Wild-type human TDP-43 expression causes TDP-43 phosphorylation, mitochondrial aggregation, motor deficits, and early mortality in transgenic mice. J. Neurosci. 30, 10851–10859 (2010).
Igaz, L.M. et al. Dysregulation of the ALS-associated gene TDP-43 leads to neuronal death and degeneration in mice. J. Clin. Invest. 121, 726–738 (2011).
Yeo, G.W., Van Nostrand, E.L. & Liang, T.Y. Discovery and analysis of evolutionarily conserved intronic splicing regulatory elements. PLoS Genet. 3, e85 (2007).
Su, A.I. et al. A gene atlas of the mouse and human protein-encoding transcriptomes. Proc. Natl. Acad. Sci. USA 101, 6062–6067 (2004).
Acknowledgements
The authors would like to thank members of B. Ren's laboratory, especially Z. Ye, S. Kuan and L. Edsall for technical help with the Illumina sequencing and U. Wagner for helpful discussions, K. Clutario and J. Boubaker for technical help, as well as all of the members of the Yeo and Cleveland laboratories, M. Ares Jr for generous support, and the neuro-team of ISIS Pharmaceuticals for critical comments and suggestions on this project. M.P. is the recipient of a Human Frontier Science Program Long Term Fellowship. C.L.-T. is the recipient of the Milton-Safenowitz post-doctoral fellowship from the Amyotrophic Lateral Sclerosis Association. D.W.C. receives salary support from the Ludwig Institute for Cancer Research. S.C.H. is funded by a National Science Foundation Graduate Research Fellowship. This work was supported by grants from the US National Institutes of Health (R37 NS27036 and an American Recovery and Reinvestment Act Challenge grant) to D.W.C and partially by grants from the US National Institutes of Health (HG004659 and GM084317) and the Stem Cell Program at the University of California, San Diego to G.W.Y.
Author information
Authors and Affiliations
Contributions
M.P., C.L.-T., J.M. and T.Y.L. performed the experiments. K.R.H., S.C.H. and T.Y.L. conducted the bioinformatics analysis. S.-C.L. developed the monoclonal TDP-43–specific antibody used for CLIP-seq and the tetracycline-inducible GFP–TDP-43–expressing HeLa cells. S.-C.L. and E.S. generated the transgenic myc–TDP-43 mice. J.P.D. and L.S. conducted the preliminary splice-junction microarray analyses. M.P., C.L.-T., E.W., C.M., Y.S., C.F.B. and H.K. conducted the antisense oligonucleotide experiments. M.P., C.L.-T., K.R.H., G.W.Y. and D.W.C. designed the experiments. M.P., C.L.-T., K.R.H., S.C.H., G.W.Y. and D.W.C. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–14 and Tables 1–6 (PDF 6524 kb)
Rights and permissions
About this article
Cite this article
Polymenidou, M., Lagier-Tourenne, C., Hutt, K. et al. Long pre-mRNA depletion and RNA missplicing contribute to neuronal vulnerability from loss of TDP-43. Nat Neurosci 14, 459–468 (2011). https://doi.org/10.1038/nn.2779
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nn.2779
This article is cited by
-
Neuropathogenesis-on-chips for neurodegenerative diseases
Nature Communications (2024)
-
Translating the ALS Genetic Revolution into Therapies: A Review
Current Treatment Options in Neurology (2024)
-
Stathmin-2 loss leads to neurofilament-dependent axonal collapse driving motor and sensory denervation
Nature Neuroscience (2024)
-
Specific vulnerability of iPSC-derived motor neurons with TDP-43 gene mutation to oxidative stress
Molecular Brain (2023)
-
Genetics of amyotrophic lateral sclerosis: seeking therapeutic targets in the era of gene therapy
Journal of Human Genetics (2023)