Abstract
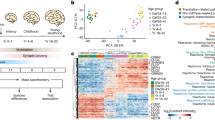
We isolated the postsynaptic density from human neocortex (hPSD) and identified 1,461 proteins. hPSD mutations cause 133 neurological and psychiatric diseases and were enriched in cognitive, affective and motor phenotypes underpinned by sets of genes. Strong protein sequence conservation in mammalian lineages, particularly in hub proteins, indicates conserved function and organization in primate and rodent models. The hPSD is an important structure for nervous system disease and behavior.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
McKusick, V.A. Am. J. Hum. Genet. 80, 588–604 (2007).
Robinson, P.N. et al. Am. J. Hum. Genet. 83, 610–615 (2008).
Smith, C.L., Goldsmith, C.A. & Eppig, J.T. Genome Biol. 6, R7 (2005).
Hurst, L.D. Trends Genet. 18, 486 (2002).
Hedges, S.B., Dudley, J. & Kumar, S. Bioinformatics 22, 2971–2972 (2006).
Wang, H.Y. et al. PLoS Biol. 5, e13 (2007).
Winter, E.E., Goodstadt, L. & Ponting, C.P. Genome Res. 14, 54–61 (2004).
Khaitovich, P. et al. Science 309, 1850–1854 (2005).
Pocklington, A.J., Cumiskey, M., Armstrong, J.D. & Grant, S.G. Mol. Syst. Biol. 2, 2006. 0023 (2006).
Fraser, H.B., Wall, D.P. & Hirsh, A.E. BMC Evol. Biol. 3, 11 (2003).
Fernández, E. et al. Mol. Syst. Biol. 5, 269 (2009).
Miao, H. et al. PLoS ONE 3, e2847 (2008).
Wang, H. et al. J. Proteome Res. 5, 361–369 (2006).
Harris, L.W. et al. PLoS ONE 3, e3964 (2008).
Acknowledgements
We thank C.P. Ponting, P.J. Brophy, R.A.W. Frank, N.H. Komiyama, M.V. Kopanitsa, T.J. Ryan and members of the Genes to Cognition Programme for critical comments on the manuscript and for discussions. We thank the tissue donors, without whom this study would have been impossible. A.B. is supported by EMBO and the European Commission. L.N.v.L., M.O.C., M.D.R.C., J.S.C. and S.G.N.G. are supported by the Wellcome Trust. I.R.W. is supported by Scottish Higher Education Funding Council and by grants from the Medical Research Council, the US National Institutes of Health and the Melville Trust.
Author information
Authors and Affiliations
Contributions
I.R.W. provided brain samples. A.B., M.O.C. and J.S.C. performed proteomic analysis. A.B. performed OMIM, ICD-10 and evolutionary analyses. L.N.v.L. performed network and enrichment analyses. M.D.R.C. integrated data into G2Cdb. L.N.v.L., M.D.R.C., M.O.C., A.B. and S.G.N.G. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–4 and Supplementary Methods (PDF 740 kb)
Supplementary Table 1
Information about Biological Samples. (XLS 28 kb)
Supplementary Table 2
Protein Identifications and Proteomic Data. (XLS 445 kb)
Supplementary Table 3
Summary of OMIM Diseases. (XLS 19 kb)
Supplementary Table 4
OMIM Diseases Identified Among Total hPSD Genes. (XLS 141 kb)
Supplementary Table 5
Human Neural Phenotype gene set enrichment analysis. (XLS 93 kb)
Supplementary Table 6
Brain datasets used in phenotype enrichment analysis. (XLS 327 kb)
Supplementary Table 7
Mammalian Neural Phenotype gene set enrichment Analysis. (XLS 187 kb)
Supplementary Table 8
Comparison of dN/dS between Genome and hPSD. (XLS 10375 kb)
Supplementary Table 9
Mouse and Human Brain Datasets dN/dS Analysis. (XLS 740 kb)
Supplementary Table 10
dN/dS Values for genes expressed in human neurons classified by cellular component. (XLS 847 kb)
Supplementary Table 11
Analysis of dN/dS on Hub, non-Hub and TAP-PSD-95 proteins. (XLS 135 kb)
Supplementary Table 12
Mouse to Human dN/dS Values in hPSD and Other Organelle Proteomes. (XLS 3289 kb)
Rights and permissions
About this article
Cite this article
Bayés, À., van de Lagemaat, L., Collins, M. et al. Characterization of the proteome, diseases and evolution of the human postsynaptic density. Nat Neurosci 14, 19–21 (2011). https://doi.org/10.1038/nn.2719
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nn.2719
This article is cited by
-
Remodeling of the postsynaptic proteome in male mice and marmosets during synapse development
Nature Communications (2024)
-
Transition of allele-specific DNA hydroxymethylation at regulatory loci is associated with phenotypic variation in monozygotic twins discordant for psychiatric disorders
BMC Medicine (2023)
-
Resting-state brain functional alterations and their genetic mechanisms in drug-naive first-episode psychosis
Schizophrenia (2023)
-
Training-induced circuit-specific excitatory synaptogenesis in mice is required for effort control
Nature Communications (2023)
-
Integrative analysis of long noncoding RNAs dysregulation and synapse-associated ceRNA regulatory axes in autism
Translational Psychiatry (2023)