Abstract
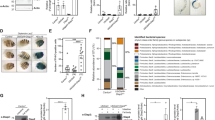
Immune homeostasis is a prerequisite to protective immunity against gastrointestinal infections. In Drosophila, immune deficiency (IMD) signalling (tumour necrosis factor receptor/interleukin-1 receptor, TNFR/IL-1R in mammals) is indispensable for intestinal immunity against invading bacteria. However, how this local antimicrobial immune response contributes to inflammatory regulation remains poorly defined. Here, we show that flies lacking intestinal Bap180 (a subunit of the chromatin-remodelling switch/sucrose non-fermentable (SWI/SNF) complex) are susceptible to infection as a result of hyper-inflammation rather than bacterial overload. Detailed analysis shows that Bap180 is induced by the IMD–Relish response to both enteropathogenic and commensal bacteria. Upregulated Bap180 can feed back to restrain overreactive IMD signalling, as well as to repress the expression of the pro-inflammatory gene eiger (TNF), a critical step to prevent excessive tissue damage and elongate the lifespan of flies, under pathological and physiological conditions, respectively. Furthermore, intestinal targeting of Baf180 renders mice susceptible to a more aggressive infectious colitis caused by Citrobacter rodentium. Together, Bap180 and Baf180 serve as a conserved transcriptional repressor that is critical for the maintenance of innate immune homeostasis in the intestines.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Garrett, W. S., Gordon, J. I. & Glimcher, L. H. Homeostasis and inflammation in the intestine. Cell 140, 859–870 (2010).
Ryu, J. H. et al. Innate immune homeostasis by the homeobox gene caudal and commensal-gut mutualism in Drosophila. Science 319, 777–782 (2008).
Buchon, N., Broderick, N. A. & Lemaitre, B. Gut homeostasis in a microbial world: insights from Drosophila melanogaster. Nat. Rev. Microbiol. 11, 615–626 (2013).
Apidianakis, Y. & Rahme, L. G. Drosophila melanogaster as a model for human intestinal infection and pathology. Dis. Model. Mech. 4, 21–30 (2011).
Kaneko, T. et al. PGRP-LC and PGRP-LE have essential yet distinct functions in the Drosophila immune response to monomeric DAP-type peptidoglycan. Nat. Immunol. 7, 715–723 (2006).
Lemaitre, B. & Hoffmann, J. The host defense of Drosophila melanogaster. Annu. Rev. Immunol. 25, 697–743 (2007).
Erturk-Hasdemir, D. et al. Two roles for the Drosophila IKK complex in the activation of relish and the induction of antimicrobial peptide genes. Proc. Natl Acad. Sci. USA 106, 9779–9784 (2009).
Stoven, S. et al. Caspase-mediated processing of the Drosophila NF-κB factor relish. Proc. Natl Acad. Sci. USA 100, 5991–5996 (2003).
Guo, L., Karpac, J. & Tran, S. L. & Jasper, H. PGRP-SC2 promotes gut immune homeostasis to limit commensal dysbiosis and extend lifespan. Cell 156, 109–122 (2014).
Chen, H., Zheng, X. & Zheng, Y. Age-associated loss of lamin-B leads to systemic inflammation and gut hyperplasia. Cell 159, 829–843 (2014).
Paredes, J. C., Welchman, D. P., Poidevin, M. & Lemaitre, B. Negative regulation by amidase PGRPs shapes the Drosophila antibacterial response and protects the fly from innocuous infection. Immunity 35, 770–779 (2011).
Libert, S., Chao, Y., Chu, X. & Pletcher, S. D. Trade-offs between longevity and pathogen resistance in Drosophila melanogaster are mediated by NFκB signaling. Aging Cell 5, 533–543 (2006).
Maillet, F., Bischoff, V., Vignal, C., Hoffmann, J. & Royet, J. The Drosophila peptidoglycan recognition protein PGRP-LF blocks PGRP-LC and IMD/JNK pathway activation. Cell Host Microbe 3, 293–303 (2008).
Zhao, H. W., Zhou, D. & Haddad, G. G. Antimicrobial peptides increase tolerance to oxidant stress in Drosophila melanogaster. J. Biol. Chem. 286, 6211–6218 (2011).
Mohrmann, L. et al. Differential targeting of two distinct SWI/SNF-related Drosophila chromatin-remodeling complexes. Mol. Cell. Biol. 24, 3077–3088 (2004).
Bonnay, F. et al. Akirin specifies NF-κB selectivity of Drosophila innate immune response via chromatin remodeling. EMBO J. 33, 2349–2362 (2014).
Basset, A. et al. The phytopathogenic bacteria erwinia carotovora infects Drosophila and activates an immune response. Proc. Natl Acad. Sci. USA 97, 3376–3381 (2000).
Carrera, I., Zavadil, J. & Treisman, J. E. Two subunits specific to the PBAP chromatin remodeling complex have distinct and redundant functions during Drosophila development. Mol. Cell. Biol. 28, 5238–5250 (2008).
Brand, A. H. & Perrimon, N. Targeted gene expression as a means of altering cell fates and generating dominant phenotypes. Development 118, 401–415 (1993).
Zeidler, M. P. et al. Temperature-sensitive control of protein activity by conditionally splicing inteins. Nat. Biotechnol. 22, 871–876 (2004).
Bae, Y. S., Choi, M. K. & Lee, W. J. Dual oxidase in mucosal immunity and host–microbe homeostasis. Trends Immunol. 31, 278–287 (2010).
Ha, E. M., Oh, C. T., Bae, Y. S. & Lee, W. J. A direct role for dual oxidase in Drosophila gut immunity. Science 310, 847–850 (2005).
Ha, E. M. et al. Regulation of DUOX by the Gαq-phospholipase Cβ-Ca2+ pathway in Drosophila gut immunity. Dev. Cell 16, 386–397 (2009).
Lemaitre, B. et al. Functional analysis and regulation of nuclear import of dorsal during the immune response in Drosophila. EMBO J. 14, 536–545 (1995).
Igaki, T. & Miura, M. The Drosophila TNF ortholog Eiger: emerging physiological roles and evolution of the TNF system. Semin. Immunol. 26, 267–274 (2014).
Bonnay, F. et al. Big bang gene modulates gut immune tolerance in Drosophila. Proc. Natl Acad. Sci. USA 110, 2957–2962 (2013).
Buchon, N., Broderick, N. A., Chakrabarti, S. & Lemaitre, B. Invasive and indigenous microbiota impact intestinal stem cell activity through multiple pathways in Drosophila. Genes Dev. 23, 2333–2344 (2009).
Igaki, T. et al. Eiger, a TNF superfamily ligand that triggers the Drosophila JNK pathway. EMBO J. 21, 3009–3018 (2002).
Adachi-Yamada, T. et al. P38 mitogen-activated protein kinase can be involved in transforming growth factor beta superfamily signal transduction in Drosophila wing morphogenesis. Mol. Cell. Biol. 19, 2322–2329 (1999).
Hay, B. A., Wolff, T. & Rubin, G. M. Expression of baculovirus P35 prevents cell death in Drosophila. Development 120, 2121–2129 (1994).
Thompson, M. Polybromo-1: the chromatin targeting subunit of the PBAF complex. Biochimie 91, 309–319 (2009).
Zaidman-Remy, A. et al. The Drosophila amidase PGRP-LB modulates the immune response to bacterial infection. Immunity 24, 463–473 (2006).
Lhocine, N. et al. PIMS modulates immune tolerance by negatively regulating Drosophila innate immune signaling. Cell Host Microbe. 4, 147–158 (2008).
Kleino, A. et al. Pirk is a negative regulator of the Drosophila Imd pathway. J. Immunol. 180, 5413–5422 (2008).
Aggarwal, K. et al. Rudra interrupts receptor signaling complexes to negatively regulate the IMD pathway. PLoS Pathog. 4, e1000120 (2008).
Mundy, R., MacDonald, T. T., Dougan, G., Frankel, G. & Wiles, S. Citrobacter rodentium of mice and man. Cell. Microbiol. 7, 1697–1706 (2005).
Georgel, P. et al. Drosophila immune deficiency (IMD) is a death domain protein that activates antibacterial defense and can promote apoptosis. Dev. Cell 1, 503–514 (2001).
Yang, Y., Hou, L., Li, Y., Ni, J. & Liu, L. Neuronal necrosis and spreading death in a Drosophila genetic model. Cell Death Dis. 4, e723 (2013).
Zheng, Y. et al. Interleukin-22 mediates early host defense against attaching and effacing bacterial pathogens. Nat. Med. 14, 282–289 (2008).
Moshkin, Y. M., Mohrmann, L., van Ijcken, W. F. & Verrijzer, C. P. Functional differentiation of SWI/SNF remodelers in transcription and cell cycle control. Mol. Cell. Biol. 27, 651–661 (2007).
Tartey, S. et al. Akirin2 is critical for inducing inflammatory genes by bridging IκB-ζ and the SWI/SNF complex. EMBO J. 33, 2332–2348 (2014).
Fang, F. et al. Proinflammatory stimuli engage Brahma related gene 1 and Brahma in endothelial injury. Circ. Res. 113, 986–996 (2013).
Tian, W. et al. Brahma-related gene 1 bridges epigenetic regulation of proinflammatory cytokine production to steatohepatitis in mice. Hepatology 58, 576–588 (2013).
Jin, Y. et al. Brahma is essential for Drosophila intestinal stem cell proliferation and regulated by hippo signaling. eLife 2, e00999 (2013).
Medzhitov, R. & Horng, T. Transcriptional control of the inflammatory response. Nat. Rev. Immunol. 9, 692–703 (2009).
Wurster, A. L. et al. IL-10 transcription is negatively regulated by BAF180, a component of the SWI/SNF chromatin remodeling enzyme. BMC Immunol. 13, 9 (2012).
Madison, B. B. et al. cis elements of the villin gene control expression in restricted domains of the vertical (crypt) and horizontal (duodenum, cecum) axes of the intestine. J. Biol. Chem. 277, 33275–33283 (2002).
Ryu, J. H. et al. An essential complementary role of NF-κB pathway to microbicidal oxidants in Drosophila gut immunity. EMBO J. 25, 3693–3701 (2006).
Rath, H. C. et al. Normal luminal bacteria, especially bacteroides species, mediate chronic colitis, gastritis, and arthritis in HLA-B27/human β2 microglobulin transgenic rats. J. Clin. Invest. 98, 945–953 (1996).
Rera, M., Clark, R. I. & Walker, D. W. Intestinal barrier dysfunction links metabolic and inflammatory markers of aging to death in Drosophila. Proc. Natl Acad. Sci. USA 109, 21528–21533 (2012).
Gay, C., Collins, J. & Gebicki, J. M. Hydroperoxide assay with the ferric-xylenol orange complex. Anal. Biochem. 273, 149–155 (1999).
Leulier, F. et al. The Drosophila immune system detects bacteria through specific peptidoglycan recognition. Nature Immunol. 4, 478–484 (2003).
Lee, T. I., Johnstone, S. E. & Young, R. A. Chromatin immunoprecipitation and microarray-based analysis of protein location. Nat. Protoc. 1, 729–748 (2006).
Yu, J. et al. MTMR4 attenuates transforming growth factor β (TGFβ) signaling by dephosphorylating R-Smads in endosomes. J. Biol. Chem. 285, 8454–8462 (2010).
Acknowledgements
The authors thank J. Treisman (New York University), J.C. Pastor (Tsinghua University), T. Igaki (Kyoto University), M. Miura (The University of Tokyo), W.-j. Lee (Seoul National University), J.-L. Imler (University of Strasbourg) and F. Shao (National Institute of Biological Sciences, Beijing) for providing numerous reagents and valuable assistance, S. Zheng (Hainan Medical University) for pathological analysis and D. Li for discussions. This work was supported by grants from the National Natural Science Foundation of China to H.T. (81220108018) and L.P. (31370909, 31570897), the Ministry of Science and Technology of China to H.T. (2011CB946104) and L.P. (2012CB518900), the Novo Nordisk-Chinese Academy of Sciences Research Foundation to L.P. (NNCAS-2011-2) and the National Institute of Health of the USA to W.Z. (HL109054). L.P. is a fellow of the CAS Youth Innovation Promotion Association (2012083).
Author information
Authors and Affiliations
Contributions
H.T. and L.P. designed the study. X.H. and J.Y. performed most of the experiments. X.H., J.Y., S.Y., S.-t.G., Y.-X.F. and L.P. collected and analysed the data. M.W., Y.H., Y.C., Y.F. and G.S. performed IHC analysis. L.S. prepared TEM samples. Z.W. helped to generate mice. H.T. and L.P. wrote the paper with contributions from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Figures 1–8, Supplementary Tables 1–7. (PDF 1468 kb)
Rights and permissions
About this article
Cite this article
He, X., Yu, J., Wang, M. et al. Bap180/Baf180 is required to maintain homeostasis of intestinal innate immune response in Drosophila and mice. Nat Microbiol 2, 17056 (2017). https://doi.org/10.1038/nmicrobiol.2017.56
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/nmicrobiol.2017.56