Abstract
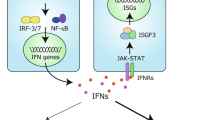
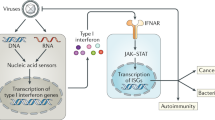
Retinoic acid-inducible gene I (RIG-I) receptor recognizes 5′-triphosphorylated RNA and triggers a signalling cascade that results in the induction of type-I interferon (IFN)-dependent responses. Its precise regulation represents a pivotal balance between antiviral defences and autoimmunity. To elucidate the cellular cofactors that regulate RIG-I signalling, we performed two global RNA interference analyses to identify both positive and negative regulatory nodes operating on the signalling pathway during virus infection. These factors were integrated with experimentally and computationally derived interactome data to build a RIG-I protein interaction network. Our analysis revealed diverse cellular processes, including the unfolded protein response, Wnt signalling and RNA metabolism, as critical cellular components governing innate responses to non-self RNA species. Importantly, we identified K-Homology Splicing Regulatory Protein (KHSRP) as a negative regulator of this pathway. We find that KHSRP associates with the regulatory domain of RIG-I to maintain the receptor in an inactive state and attenuate its sensing of viral RNA (vRNA). Consistent with increased RIG-I antiviral signalling in the absence of KHSRP, viral replication is reduced when KHSRP expression is knocked down both in vitro and in vivo. Taken together, these data indicate that KHSRP functions as a checkpoint regulator of the innate immune response to pathogen challenge.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Tisoncik, J. R. et al. Into the eye of the cytokine storm. Microbiol. Mol. Biol. Rev. 76, 16–32 (2012).
Pichlmair, A. et al. RIG-I-mediated antiviral responses to single-stranded RNA bearing 5′-phosphates. Science 314, 997–1001 (2006).
Yoneyama, M. et al. The RNA helicase RIG-I has an essential function in double-stranded RNA-induced innate antiviral responses. Nat. Immunol. 5, 730–737 (2004).
Takahasi, K. et al. Nonself RNA-sensing mechanism of RIG-I helicase and activation of antiviral immune responses. Mol. Cell 29, 428–440 (2008).
Kato, H. et al. Differential roles of MDA5 and RIG-I helicases in the recognition of RNA viruses. Nature 441, 101–105 (2006).
Saito, T., Owen, D. M., Jiang, F., Marcotrigiano, J. & Gale, M. Jr. Innate immunity induced by composition-dependent RIG-I recognition of hepatitis C virus RNA. Nature 454, 523–527 (2008).
Strahle, L., Garcin, D. & Kolakofsky, D. Sendai virus defective-interfering genomes and the activation of interferon-beta. Virology 351, 101–111 (2006).
Meylan, E. et al. Cardif is an adaptor protein in the RIG-I antiviral pathway and is targeted by hepatitis C virus. Nature 437, 1167–1172 (2005).
Loo, Y.-M. & Gale, M. Jr Immune signaling by RIG-I-like receptors. Immunity 34, 680–692 (2011).
Gack, M. U. et al. TRIM25 RING-finger E3 ubiquitin ligase is essential for RIG-I-mediated antiviral activity. Nature 446, 916–920 (2007).
Zhong, B. et al. The E3 ubiquitin ligase RNF5 targets virus-induced signaling adaptor for ubiquitination and degradation. J. Immunol. 184, 6249–6255 (2010).
Lei, Y. et al. The mitochondrial proteins NLRX1 and TUFM form a complex that regulates type I interferon and autophagy. Immunity 36, 933–946 (2012).
Saitoh, T. et al. Negative regulation of interferon-regulatory factor 3-dependent innate antiviral response by the prolyl isomerase Pin1. Nat. Immunol. 7, 598–605 (2006).
Rothenfusser, S. et al. The RNA helicase Lgp2 inhibits TLR-independent sensing of viral replication by retinoic acid-inducible gene-I. J. Immunol. 175, 5260–5268 (2005).
Choi, S. J. et al. HDAC6 regulates cellular viral RNA sensing by deacetylation of RIG-I. EMBO J. 35, 429–442 (2016).
Crow, Y. J. Type I interferonopathies: Mendelian type I interferon up-regulation. Curr. Opin. Immunol. 32, 7–12 (2015).
Crow, Y. J. Type I interferonopathies: a novel set of inborn errors of immunity. Ann. NY Acad. Sci. 1238, 91–98 (2011).
Mibayashi, M. et al. Inhibition of retinoic acid-inducible gene I-mediated induction of beta interferon by the NS1 protein of influenza A virus. J. Virol. 81, 514–524 (2007).
Gack, M. U. et al. Influenza A virus NS1 targets the ubiquitin ligase TRIM25 to evade recognition by the host viral RNA sensor RIG-I. Cell Host Microbe 5, 439–449 (2009).
Friedman, C. S. et al. The tumour suppressor CYLD is a negative regulator of RIG-I-mediated antiviral response. EMBO Rep. 9, 930–936 (2008).
Konig, R. et al. A probability-based approach for the analysis of large-scale RNAi screens. Nat. Methods 4, 847–849 (2007).
Soulat, D. et al. The DEAD-box helicase DDX3X is a critical component of the TANK-binding kinase 1-dependent innate immune response. EMBO J. 27, 2135–2146 (2008).
Okabe, Y., Sano, T. & Nagata, S. Regulation of the innate immune response by threonine-phosphatase of eyes absent. Nature 460, 520–524 (2009).
You, F. et al. PCBP2 mediates degradation of the adaptor MAVS via the HECT ubiquitin ligase AIP4. Nat. Immunol. 10, 1300–1308 (2009).
Jia, Y. et al. Negative regulation of MAVS-mediated innate immune response by PSMA7. J. Immunol. 183, 4241–4248 (2009).
Konig, R. et al. Human host factors required for influenza virus replication. Nature 463, 813–817 (2010).
Kondo, T., Kawai, T. & Akira, S. Dissecting negative regulation of toll-like receptor signaling. Trends Immunol. 33, 449–458 (2012).
Yasukawa, H., Sasaki, A. & Yoshimura, A. Negative regulation of cytokine signaling pathways. Annu. Rev. Immunol. 18, 143–164 (2000).
Komuro, A., Bamming, D. & Horvath, C. M. Negative regulation of cytoplasmic RNA-mediated antiviral signaling. Cytokine 43, 350–358 (2008).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Seth, R. B., Sun, L. & Chen, Z. J. Antiviral innate immunity pathways. Cell Res. 16, 141–147 (2006).
Wang, Y. Y., Li, L., Han, K. J., Zhai, Z. & Shu, H. B. A20 is a potent inhibitor of TLR3- and Sendai virus-induced activation of NF-κB and ISRE and IFN-β promoter. FEBS Lett. 576, 86–90 (2004).
Lin, R. et al. Negative regulation of the retinoic acid-inducible gene I-induced antiviral state by the ubiquitin-editing protein A20. J. Biol. Chem. 281, 2095–2103 (2006).
King, P. H. & Chen, C. Y. Role of KSRP in control of type I interferon and cytokine expression. J. Interferon Cytokine Res. 34, 267–274 (2014).
Lin, W. J. et al. Posttranscriptional control of type I interferon genes by KSRP in the innate immune response against viral infection. Mol. Cell Biol. 31, 3196–3207 (2011).
Patel, J. R. et al. ATPase-driven oligomerization of RIG-I on RNA allows optimal activation of type-I interferon. EMBO Rep. 14, 780–787 (2013).
Gardner, B. M. & Walter, P. Unfolded proteins are Ire1-activating ligands that directly induce the unfolded protein response. Science 333, 1891–1894 (2011).
Wies, E. et al. Dephosphorylation of the RNA sensors RIG-I and MDA5 by the phosphatase PP1 is essential for innate immune signaling. Immunity 38, 437–449 (2013).
Tripathi, S. et al. Meta- and orthogonal integration of influenza ‘OMICs’ data defines a role for UBR4 in virus budding. Cell Host Microbe 18, 723–735 (2015).
Pickup, J. C. Inflammation and activated innate immunity in the pathogenesis of type 2 diabetes. Diabetes Care 27, 813–823 (2004).
Allam, R. & Anders, H.-J. The role of innate immunity in autoimmune tissue injury. Curr. Opin. Rheumatol. 20, 538–544 (2008).
Tardif, K. D., Mori, K. & Siddiqui, A. Hepatitis C virus subgenomic replicons induce endoplasmic reticulum stress activating an intracellular signaling pathway. J. Virol. 76, 7453–7459 (2002).
Hampton, R. Y. ER stress response: getting the UPR hand on misfolded proteins. Curr. Biol. 10, R518–R521 (2000).
Urano, F. et al. Coupling of stress in the ER to activation of JNK protein kinases by transmembrane protein kinase IRE1. Science 287, 664–666 (2000).
Cho, J. A. et al. The unfolded protein response element IRE1α senses bacterial proteins invading the ER to activate RIG-I and innate immune signaling. Cell Host Microbe 13, 558–569 (2013).
Eckard, S. C. et al. The SKIV2L RNA exosome limits activation of the RIG-I-like receptors. Nat. Immunol. 15, 839–845 (2014).
Demand, J., Alberti, S., Patterson, C. & Höhfeld, J. Cooperation of a ubiquitin domain protein and an E3 ubiquitin ligase during chaperone/proteasome coupling. Curr. Biol. 11, 1569–1577 (2001).
Reina, C. P., Nabet, B. Y., Young, P. D. & Pittman, R. N. Basal and stress-induced Hsp70 are modulated by ataxin-3. Cell Stress Chaperon. 17, 729–742 (2012).
Cabral Miranda, F. et al. CHIP, a carboxy terminus HSP-70 interacting protein, prevents cell death induced by endoplasmic reticulum stress in the central nervous system. Front. Cell. Neurosci. 8, 438 (2015).
Cui, S. et al. The C-terminal regulatory domain is the RNA 5′-triphosphate sensor of RIG-I. Mol. Cell 29, 169–179 (2008).
Baum, A., Sachidanandam, R. & García-Sastre, A. Preference of RIG-I for short viral RNA molecules in infected cells revealed by next-generation sequencing. Proc. Natl Acad. Sci. USA 107, 16303–16308 (2010).
Alamares-Sapuay, J. G. et al. Serum- and glucocorticoid-regulated kinase 1 is required for nuclear export of the ribonucleoprotein of influenza A virus. J. Virol. 87, 6020–6026 (2013).
Ran, F. A. et al. Genome engineering using the CRISPR-Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Goujon, C. et al. Human MX2 is an interferon-induced post-entry inhibitor of HIV-1 infection. Nature 502, 559–562 (2013).
Jager, S. et al. Global landscape of HIV–human protein complexes. Nature 481, 365–370 (2012).
Jager, S. et al. Purification and characterization of HIV–human protein complexes. Methods 53, 13–19 (2011).
Verschueren, E. et al. Scoring large-scale affinity purification mass spectrometry datasets with MiST. Curr. Protoc. Bioinformatics 49, 8 19 11–18 19 16 (2015).
Sowa, M. E., Bennett, E. J., Gygi, S. P. & Harper, J. W. Defining the human deubiquitinating enzyme interaction landscape. Cell 138, 389–403 (2009).
Chiang, C. Y. et al. Cofactors required for TLR7- and TLR9-dependent innate immune responses. Cell Host Microbe 11, 306–318 (2012).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Dang, O., Navarro, L., Anderson, K. & David, M. Cutting edge: anthrax lethal toxin inhibits activation of IFN-regulatory factor 3 by lipopolysaccharide. J. Immunol. 172, 747–751 (2004).
Runge, S. et al. In vivo ligands of MDA5 and RIG-I in measles virus-infected cells. PLoS Pathog. 10, e1004081 (2014).
Weber, M. & Weber, F. Monitoring activation of the antiviral pattern recognition receptors RIG-I and PKR by limited protease digestion and native PAGE. J. Vis. Exp. 89, e51415 (2014).
Matrosovich, M., Matrosovich, T., Garten, W. & Klenk, H. D. New low-viscosity overlay medium for viral plaque assays. Virol. J. 3, 63 (2006).
Acknowledgements
The authors acknowledge the Krogan laboratory for running and analysis of the AP-MS experiments, M. Schneider for help with editing the manuscript and the Chanda laboratory for discussions and advice. The authors thank R. Albrecht for providing the H1N1 PR8/34 IAV stocks used in the in vivo mouse experiments and C. Lässig and K.-P. Hopfner for providing recombinant RIG-I protein for in vitro assays. The authors acknowledge support from the National Institutes of Health (U19 AI106754 and P50 GM085764). This work was also supported by a grant from the James B. Pendleton Charitable Trust and by CRIP (Center for Research on Influenza Pathogenesis), an NIAID funded Center of Excellence for Influenza Research and Surveillance (CEIRS, contract no. HHSN272201400008C).
Author information
Authors and Affiliations
Contributions
S.S., S.M.Y. and S.K.C. wrote the manuscript. S.S., R.K., S.M.Y. and S.K.C. designed experiments and interpreted data. Y.Z. and A.G.-S. interpreted data. S.S., A.R.-F., S.M.Y., F.G., N.J.H., S.T., V.R.M.T.B., A.I., M.T.S.-A., E.d.C., P.D.D.J., Q.N. and N.J.K. performed experiments. H.M. and D.A.S. provided reagents.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Figures 1–12, Supplementary Discussion, Supplementary Methods, Supplementary References. (PDF 2297 kb)
Supplementary Table 1
RSA analysis of RIG-I positive regulator screen. (XLSX 3747 kb)
Supplementary Table 2
RSA analysis of RIG-I negative regulator screen. (XLSX 2425 kb)
Supplementary Table 3
Summary of RIG-I negative regulator selection strategy. (XLSX 102 kb)
Supplementary Table 4
AP-MS analysis of RIG-I signalling factors. (XLSX 373 kb)
Supplementary Table 5
Gene ontology and functional enrichment of the RIG-I protein network. (XLSX 53 kb)
Supplementary Table 6
Binary interactions of the RIG-I pathway interaction map (Fig. 1d). (XLSX 51 kb)
Rights and permissions
About this article
Cite this article
Soonthornvacharin, S., Rodriguez-Frandsen, A., Zhou, Y. et al. Systems-based analysis of RIG-I-dependent signalling identifies KHSRP as an inhibitor of RIG-I receptor activation. Nat Microbiol 2, 17022 (2017). https://doi.org/10.1038/nmicrobiol.2017.22
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/nmicrobiol.2017.22
This article is cited by
-
Regulation of RIG-I-like receptor-mediated signaling: interaction between host and viral factors
Cellular & Molecular Immunology (2021)
-
Intracellular sensing of viral genomes and viral evasion
Experimental & Molecular Medicine (2019)