Abstract
The shallow water bivalve Codakia orbicularis lives in symbiotic association with a sulfur-oxidizing bacterium in its gills. The endosymbiont fixes CO2 and thus generates organic carbon compounds, which support the host's growth. To investigate the uncultured symbiont's metabolism and symbiont–host interactions in detail we conducted a proteogenomic analysis of purified bacteria. Unexpectedly, our results reveal a hitherto completely unrecognized feature of the C. orbicularis symbiont's physiology: the symbiont's genome encodes all proteins necessary for biological nitrogen fixation (diazotrophy). Expression of the respective genes under standard ambient conditions was confirmed by proteomics. Nitrogenase activity in the symbiont was also verified by enzyme activity assays. Phylogenetic analysis of the bacterial nitrogenase reductase NifH revealed the symbiont's close relationship to free-living nitrogen-fixing Proteobacteria from the seagrass sediment. The C. orbicularis symbiont, here tentatively named ‘Candidatus Thiodiazotropha endolucinida’, may thus not only sustain the bivalve's carbon demands. C. orbicularis may also benefit from a steady supply of fixed nitrogen from its symbiont—a scenario that is unprecedented in comparable chemoautotrophic symbioses.
Similar content being viewed by others
Mutualistic associations between marine invertebrates and sulfur-oxidizing (thiotrophic) bacteria are a well-documented and widespread phenomenon in a variety of sulfidic habitats ranging from hydrothermal vents to shallow-water coastal ecosystems1–3. Thioautotrophic symbionts generate energy through sulfide oxidation and provide their hosts with organic carbon. In the Lucinidae, a diverse family of marine bivalves, all members are obligatorily dependent on their bacterial gill endosymbionts after larval development and metamorphosis4. The shallow-water lucinid Codakia orbicularis, which lives in the sediment beneath the tropical seagrass Thalassia testudinum along the Caribbean and Western Atlantic coast5, harbours a single species of endosymbionts in its gills6. The symbiont has been shown to be newly acquired by each clam generation7,8 from a pool of free-living symbiosis-competent bacteria in the environment9, rather than being inherited from clam parents. C. orbicularis appears not to release its endosymbionts, even under adverse conditions, but can digest them as a source of nutrition10–12. Moreover, bacterial cell division seems to be inhibited inside the host tissue. The majority of the symbiont population was shown to be polyploid (that is, containing multiple genome copies), while dividing symbiont cell stages are very rarely observed in host bacteriocytes13. The host undoubtedly benefits from the symbiont both by way of detoxification of its sulfidic environment and by supply of organic compounds through the bacterial Calvin–Benson cycle. It remains questionable, however, whether the symbiont gains any advantage from this association in evolutionary terms11.
Biological nitrogen fixation (diazotrophy) is the conversion of molecular nitrogen (N2) into ammonia14. It provides the basis for the biosynthesis of organic nitrogen compounds and thus of indispensable cellular components such as proteins and nucleic acids. Diazotrophy occurs exclusively in the prokaryotic kingdom15; nitrogen fixation in eukaryotes is therefore restricted to symbioses with bacteria16. Prokaryotic nitrogen fixers are associated with a diversity of eukaryotic hosts in a wide range of habitats, including the well-studied Rhizobium and Bradyrhizobium genera that form root-nodule symbioses with legumes17 and the actinomycetal Frankia species associated with woody plants18. In marine environments, symbiotic nitrogen-fixers are known to be associated with a variety of invertebrates, such as wood-boring bivalves (shipworms)19, corals, sponges and sea urchins20.
Marine ecosystems are frequently nitrogen-limited and therefore considered to be favourable niches for diazotrophic organisms21. Coastal seagrass beds, such as those inhabited by Codakia species, are areas of particularly high microbial nitrogen fixation activity14, as the plants' rhizosphere is densely populated by a complex diazotrophic bacterial community22,23. The majority of these nitrogen-fixers are anaerobic, heterotrophic sulfate reducers, which decompose decaying seagrass leaves, producing large amounts of toxic sulfide24. Thiotrophic members of the rhizosphere community oxidize and thus detoxify the sulfide25. Unlike most other animals, for which sulfide is toxic, lucinids thrive in this habitat. Moreover, through the sulfide-oxidation activity of their symbionts, the bivalves contribute to sulfide detoxification of the seagrass bed ecosystem and have therefore been suggested to be part of a beneficial association with the seagrass (‘tripartite symbiosis’)26. However, nitrogen fixation in their sulfur-oxidizing symbiont has to our knowledge never been reported for C. orbicularis, nor yet for any other thiotrophic marine invertebrate symbiosis, despite the broad scientific attention these systems have attracted1–3.
Here, we show, in a combined proteogenomic and experimental approach, that the thioautotrophic gill endosymbiont of C. orbicularis, ‘Candidatus Thiodiazotropha endolucinida’, expresses nitrogen fixation proteins and exhibits nitrogenase activity under its natural ambient conditions in the host. This finding adds a novel feature to the picture of the Codakia symbiosis and enhances our understanding of its role in the seagrass ecosystem.
Results and discussion
Detection of nitrogen fixation in the C. orbicularis symbiont
To reconstruct the C. orbicularis symbiont's metabolism, we sequenced the genome of purified bacteria to provide a basis for subsequent global proteomic analyses. As expected, the symbiont's genome encoded all the enzymes necessary for sulfide oxidation via the Dsr-APS-Sat pathway and for carbon fixation through the Calvin–Benson cycle. The respective enzymes were expressed in high concentrations, as detected at the protein level (these results will be the subject of a separate publication). Interestingly, we also detected a complete gene cluster for the fixation of dinitrogen in the symbiont's genome (CODIS_20190–20650), including the highly conserved nitrogenase enzyme complex.
The C. orbicularis symbiont's nitrogen fixation (nif) cluster comprises 47 genes (Fig. 1), 26 of which were found to be expressed as proteins in this study (Supplementary Tables 3 and 4). The actual nitrogenase enzyme complex consists of two components: a dinitrogenase (component I), which is a molybdenum-iron protein and heterotetramer of two NifD and two NifK subunits, and a dinitrogenase reductase (component II), a homodimer of two NifH subunits15. All three proteins were found to be expressed in C. orbicularis symbiont samples isolated from freshly collected clams (see Supplementary Table 4 for protein concentrations). Nitrogen fixation-related electron transfer proteins, regulatory proteins, as well as proteins needed for nitrogenase cofactor biosynthesis are also encoded among the nif genes and many of them were detected as proteins under in situ conditions. In addition to these nif-encoded proteins, we also identified a number of genes and proteins that are apparently indirectly related to nitrogen fixation in the C. orbicularis symbiont (see Supplementary Results and Discussion for details). First, immediately adjacent to the nif genes, the C. orbicularis symbiont's genome encodes a set of Rnf electron transfer proteins (CODIS_20660–20720), which are putatively involved in electron transfer towards the nitrogenase complex. Second, the high abundance of glutamine synthetase (CODIS_03590) and the P-II signal transduction protein GlnK (CODIS_03150) suggests that nitrogen fixation-derived ammonium is assimilated through the GS/GOGAT pathway and that the symbionts analysed in this study may have experienced nitrogen limitation. The two-component nitrogen regulation (Ntr) system, which controls ammonium assimilation in Proteobacteria27, appears also to be present and active in the C. orbicularis symbiont. Third, we also detected a rubrerythrin (Rbr, CODIS_04230) as one of the most abundant proteins in the ‘Ca. T. endolucinida’ proteome. Rubrerythrins provide protection against hydrogen peroxide-mediated oxidative stress28 and have been suggested to play a role in shielding the highly oxygen-sensitive nitrogenase protein from oxidative damage29. We observed that both the nitrogenase proteins and Rbr were detected with significantly lower protein concentrations during diazotrophy-inhibiting conditions (that is, energy limitation caused by sulfide starvation; Supplementary Fig. 5 and Supplementary Results and Discussion). These results may suggest a co-regulation of Rbr and nitrogenase expression and support the idea that Rbr might be involved in protecting the nitrogenase from oxidative stress.
The ‘Ca. T. endolucinida’ COS nif and rnf gene clusters (middle) were compared to the genomes of Allochromatium vinosum DSM 180 (top) and Sedimenticola thiotaurini SIP-G1 (bottom) using Bl2seq (BLASTn, E-value 1e-5). Gene functions: red, nitrogenase and nitrogenase reductase; blue, electron transfer, cofactor biosynthesis and other nitrogen fixation-specific functions; yellow, transcriptional and post-translational regulation of nitrogen fixation; grey, function not necessarily related to nitrogen fixation; green, rnf genes. White arrows with solid outlines indicate unknown function, and white arrows with broken outlines indicate other functions (these genes of A. vinosum and S. thiotaurini do not have a homologue in the C. orbicularis symbiont's nif cluster). Sequence similarities are symbolized by red hues for direct comparisons and blue hues for reversed comparisons. Darker colours correspond to higher identities. For protein functions see Supplementary Table 3. *Identified as a protein in this study. Sequence data and the BLAST comparison files were drawn with the R package genoPlotR (ref. 57) and edited in Inkscape.
The functionality of the nitrogenase enzyme complex was confirmed by an acetylene reduction assay30. We detected the nitrogenase-dependent conversion of acetylene to ethylene in C. orbicularis gill tissue samples and in purified symbionts obtained from freshly collected individuals (Fig. 2). Ethylene accumulation rates in gill samples ranged from 8.65 to 15.41 nmol h−1 mg−1 of tissue under aerobic conditions, while no ethylene production was detected in gills incubated under low oxygen conditions. The latter might be due to the fact that host bacteriocytes probably die quickly in the absence of oxygen, which may also be detrimental for their intracellular symbionts. On the other hand, symbionts that were isolated from their bacteriocytes showed distinctly higher nitrogenase activities (Fig. 2) with an average ethylene production rate of 43 nmol h−1 mg−1 under aerobic conditions. Under microaerobic conditions, the nitrogenase activity of purified symbionts was even higher (57.5 nmol C2H4 h−1 mg−1) than in the aerobic approach, suggesting that the thioautotrophic symbiont's nitrogenase enzyme complex may—as most other nitrogenases—be oxygen-sensitive and work more efficiently under low-oxygen conditions16. Low ethylene production (∼0.24 nmol C2H4 h−1 mg−1) was observed in crude sediment collected from T. testudinum seagrass beds, presumably indicating the presence of free-living nitrogen-fixing bacteria in the sediment. No ethylene production was detected after incubation of sea water with acetylene and after incubation of symbiont-free foot tissue with acetylene, respectively. This clearly indicates that nitrogenase activity is exclusively present in symbiont-containing tissue in C. orbicularis.
The production of ethylene as a consequence of acetylene reduction by the nitrogenase enzyme complex was measured in gill tissue samples from freshly collected C. orbicularis specimens, in purified gill symbionts, in symbiont-free foot tissue, in the water column and in T. testudinum sediment. Gill tissue and isolated symbionts were assayed under aerobic (+O2) conditions and under microaerobic (−O2) conditions (‘+O2’ and ‘−O2’ refer to initial oxygen concentrations at the start of the assay). All other sample types were incubated under aerobic conditions. Purified symbiont fractions contained approximately 5 × 108 to 8 × 108 bacteria per assay and gill tissue samples contained approximately 3 × 108 bacteria per assay (see Methods for details). The ethylene production rate is given in nmol h−1 mg−1 as average values (n = 5), and error bars show standard deviation. No ethylene production was detected in gill tissue samples under microaerobic conditions, in symbiont-free foot tissue and in the water column.
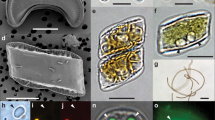
To support this observation, we performed a western blot analysis based on an antibody raised against NifH (∼34 kDa). As displayed in Fig. 3a, hybridization of the NifH antibody was observed in extracts from freshly collected C. orbicularis gills, while no signal could be detected in the negative control, that is, gill tissue from the asymbiotic marine bivalve Arcopagia crassa (Tellinidae). As described by Frenkiel and Mouëza31, gills of freshly collected individuals of C. orbicularis consist of an arrangement of filaments organized in three zones, the lateral of which contains the bacteriocytes (BCs), filled with intracellular bacterial symbionts, and granule-containing cells (GC) without endosymbionts (Fig. 3b). To allocate the expression of the nitrogenase protein NifH to the symbionts in the bacteriocytes, we conducted immunofluorescence microscopy on C. orbicularis gill histological sections using an anti-NifH antibody and 4′,6-diamidino-2-phenylindole (DAPI) counterstaining. The DAPI staining (Fig. 3c) allowed for the localization of host nuclei (large light blue dots) in bacteriocytes and granule cells all along the lateral gill filament zone but also of bacterial DNA (small light blue dots), only within the bacteriocytes. The NifH signals (yellow fluorescence, Fig. 3d) cover the bacteriocyte area but are absent from the granule cells and thus evidently emanate from the bacterial endosymbionts located in the cytoplasm of the bacteriocytes (Fig. 3e), even if not all the bacteria from a single bacteriocyte seem to be positive. In the asymbiotic tellinid A. crassa, which was used as a negative control, no NifH immunofluorescence was observed (Supplementary Fig. 2). Moreover, trophosome tissue sections of the deep-sea tube worm Riftia pachyptila (Vestimentifera, negative control 2), did not hybridize with the NifH antibody either. R. pachyptila harbours a sulfur-oxidizing symbiont in its trophosome, which is closely related to the C. orbicularis symbiont, but which is not capable of diazotrophy32,33. These results clearly reveal that nifH expression in C. orbicularis can be specifically attributed to the symbionts inside the host bacteriocytes.
a, Western blot analysis of NifH expression in gill extracts (50 mg of protein extract per lane) of the asymbiotic tellinid Arcopagia crassa (left, negative control) and of C. orbicularis (right). The NifH band migrates at around 34 kDa. The full western blot is displayed in Supplementary Fig. 6. b, Goldner staining of gill filaments from a freshly collected C. orbicularis bivalve. Each gill filament possesses a ciliated zone (CZ) devoid of bacteria. The lateral zone contains mostly bacteriocytes (BC) harbouring the bacterial symbionts and granule cells (GC, stained in orange), which are free of bacteria. Dark brown dots are C. orbicularis cell nuclei. Scale bar, 30 µm. c, DAPI staining of the lateral gill filament zone. Large bright blue dots indicate the location of nuclei in bacteriocytes and granule cells. Symbiont DNA is visible as small light blue dots all over the bacteriocyte area but not in the granule cell area. Sp: space between the gill filaments. Scale bar, 20 µm. d, Immunolocalization of the C. orbicularis symbiont's nitrogenase protein in the same gill section as in c, hybridized with the Alexa Fluor 546-labelled NifH antibody (yellow fluorescence). NifH fluorescence is visible in bacteriocytes but not in granule cells. Sp: space between gill filaments. Scale bar, 20 µm. e, Magnification of a detail from d (shown by a rectangle), highlighting the localization of NifH labelling in the bacteria. Scale bar, 2 µm.
Availability and use of other nitrogen sources in the seagrass sediment
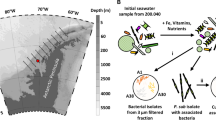
Our sediment pore water analyses show that ambient inorganic nitrogen concentrations are comparatively low in the direct vicinity of C. orbicularis (that is, within a radius of 20 cm from the animals), suggesting that diazotrophy is of particular advantage for the bacterial symbiont in this habitat. In marine systems, the main sources of nitrogen available for bacterial growth (besides N2) are NH4+ and free amino acids34, as well as—to a lesser extent—NO3−, NO2− and urea35. In seagrass bed sediments in particular, NH4+ is considered to be the dominating nitrogen source, with concentrations of up to 175 µM in some areas36. We detected NH4+ levels from 1 to 10 µM in the T. testudinum sediment directly surrounding C. orbicularis, with the highest concentrations measured in the deeper layers of the sediment (Fig. 4). These values are low compared to previously reported NH4+ concentrations of up to 120 µM in T. testudinum sediments in subtropical Florida37. However, sediment pore water ammonium concentrations have been shown to vary substantially between sampling sites and times36,38. In tropical T. testudinum seagrass beds of the Caribbean, NH4+ levels were observed to be particularly low, that is, below 25 µM in the upper 10 cm of sediment39. In this presumably ammonium-limited environment, nitrogen fixation is probably the preferable strategy for bacterial nitrogen acquisition.
Gas concentrations (in µM) in pore water samples from T. testudinum sediments are given as averages of six individual sediment cores (n = 6). Standard deviations are indicated by error bars. The light grey background indicates the sediment depth in which C. orbicularis burrows. Sediment samples were taken within a radius of 20 cm around C. orbicularis specimens.
Interestingly, a number of genes required for uptake and assimilation of nitrate and/or nitrite were found to be expressed in the C. orbicularis symbiont (including a nitrate transporter and assimilatory nitrate and nitrite reductase subunits, Supplementary Table 4). Unlike ammonium, nitrate and nitrite are usually considered not to play a substantial role as nitrogen sources in seagrass sediments, with NO3− + NO2− concentrations ranging from 0 to 10 µM (ref. 36). In accord with previous studies, we detected NO3− and NO2− concentrations of up to 9 µM and below 3 µM, respectively, in our pore water analysis in the immediate surroundings, that is, within a radius of 20 cm around C. orbicularis specimens (Fig. 4). The detection of assimilatory nitrate/nitrite reductase subunits indicates that, despite the relatively low ambient concentration, nitrate (and nitrite) may be relevant as nitrogen sources for the C. orbicularis symbiont after all, in addition to N2 fixation. This seems to be surprising at first, as nitrogen fixation is often repressed in the presence of alternative nitrogen sources in free-living diazotrophs40. The simultaneous expression of both nif genes and nitrate assimilation genes might be due to less stringent regulation in C. orbicularis symbionts, as also suggested for other symbiotic nitrogen fixers, which provide for their hosts' nitrogen supply41. However, the most likely explanation would be that the environmental nitrate and nitrite levels are too low to trigger repression of the N2 fixation genes.
Overall, in view of the specific nitrogen-limited ambient conditions in the seagrass bed habitat, nitrogen fixation in the C. orbicularis symbiont emerges as a most advantageous feature. Moreover, employing bacterial endosymbionts that can fix N2 in this environment might also be highly profitable for C. orbicularis. Nitrogen incorporated by the symbiont through N2 fixation most probably supplements the bivalve's nitrogen diet after symbiont digestion. Thus supplied, the host would circumvent competition with seagrass root tissues for other, limited nitrogen sources. (See Supplementary Fig. 3 for a model of interactions between bivalves, symbionts, seagrass and the free-living microbial community.)
Phylogeny of nitrogen fixation in the C. orbicularis symbiont
Our phylogenetic analyses of the nitrogenase reductase NifH and of the nitrogenase subunits NifD and NifK place the C. orbicularis symbiont in a group with free-living diazotrophic Gammaproteobacteria that are known to be sulfur oxidizers, such as Sedimenticola thiotaurini, Thiorhodococcus drewsii and Allochromatium vinosum (Fig. 5 and Supplementary Fig. 4b–d). Moreover, particularly high similarities were observed between the C. orbicularis symbiont's NifH and NifH sequences of a variety of uncultured diazotrophic isolates from seagrass beds and mangrove sediments, some of them classified as presumptive Beta- or Gammaproteobacteria42 (see Supplementary Results and Discussion and Supplementary Table 5 for details). Although these uncultured isolates are not characterized with regard to their 16S rRNA genes (nor their genetic potential for thiotrophy), this pronounced NifH-based identity implies a high degree of physiological similarity to ‘Ca. T. endolucinida’. At the level of 16S rRNA sequence phylogeny, the C. orbicularis symbiont clusters most closely with other yet uncultured thiotrophic lucinid symbionts from seagrass bed habitats, many of which are known Gammaproteobacteria (Supplementary Fig. 4a). None of these related lucinid symbionts has been characterized with regard to potential diazotrophy, but considering their close phylogenetic relatedness and their very similar ecosystems it seems quite likely that these symbionts, too, are capable of nitrogen fixation.
The tree was inferred based on maximum likelihood. Numbers given on the branches are bootstrap proportions as a percentage of 1,000 replicates for values ≥50%. Taxonomic classes of the phylum Proteobacteria are given on the right. Although the original inference used full-length NifH amino acid sequences, partial NifH sequences from uncultured marine isolates were added by maximum parsimony (dotted branches). Note that some of the full-length sequences had up to 80 partial sequences affiliated to them. For clarity's sake, only one partial sequence per full-length sequence is displayed (that is, the one with highest sequence similarity). For a comprehensive list of all partial sequences that were affiliated to the C. orbicularis symbiont ‘Candidatus Thiodiazotropha endolucinida’ COS see Supplementary Table 5. ‘Ca. T. endolucinida’ COS (this study) is shown in bold. Orange dots denote known diazotrophic sulfur oxidizers. (Note that no information regarding potential thiotrophy is available for the uncultured seagrass bed isolates. Some of these clones might be sulfur oxidizers, although they are not marked with orange dots.) Accession numbers or locus tags in parentheses refer to the International Nucleotide Sequence Database Collaboration (INSDC). The sequence marked with an asterisk was only available as a nucleotide sequence and was translated for this analysis (frame 3). The NifH sequences of Anabaena variabilis ATCC 29413 (AAA93020) and Frankia alni ArI3 (AAA96262) were used as outgroup (shown in Supplementary Figure 4d).
These observations suggest the presence of a hitherto unrecognized group of sulfur-oxidizing diazotrophic Gammaproteobacteria in seagrass sediment ecosystems, whose members can be free-living, like the uncultured clones, or live in symbiosis with C. orbicularis and (possibly) other lucinids. The phylogenetic relatedness of thiotrophic lucinid symbionts and seagrass sediment bacteria has been suggested previously23,43. The C. orbicularis symbiont exists as a free-living bacterium in the environment before colonization of its bivalve host9 and therefore presumably belongs to this microbial seagrass community. Moreover, free-living sulfide oxidizers are known to be part of the complex bacterial population in the seagrass rhizosphere (Supplementary Fig. 3)25, although their potential involvement in plant-associated nitrogen fixation has not been addressed so far. It will be highly interesting to investigate this hypothetical gammaproteobacterial community of thiotrophic diazotrophs in seagrass beds in further studies, to verify its existence, abundance and composition.
Conclusion
The results presented in this study provide evidence that the thiotrophic C. orbicularis gill symbiont ‘Ca. T. endolucinida’ is a diazotroph. The bacterium is thus perfectly adapted to the prevalent low NH4+ and NO3− concentrations in its seagrass habitat. As C. orbicularis digests parts of its symbiont population for nutrition, it seems very likely that diazotrophy-derived nitrogen compounds are transferred from the symbiont to the host, supporting the bivalve's nitrogen diet. Future studies will be needed to confirm this proposition. Harbouring a ‘multi-talented’ symbiont, which not only detoxifies sulfide and sustains its host's carbon needs, but which presumably also provides nitrogen compounds, seems to be most advantageous for C. orbicularis and may enable the bivalve to prevail in its highly sulfidic and nitrogen-limited environment. The interesting question of how the symbiont might benefit from this association in evolutionary terms—considering the highly restrictive regime of its host—remains to be elucidated in future studies (Supplementary Discussion). As the specific ambient conditions in sulfidic seagrass beds are a common feature shared by many lucinid symbioses, it seems quite likely that future studies will reveal diazotrophy in other thiotrophic lucinid symbionts as well. Follow-up studies are required to elucidate the composition of a putative population of sulfur-oxidizing diazotrophic Gammaproteobacteria in the seagrass bed sediment and their specific role in the rhizosphere ecosystem.
Marine chemoautrophic symbioses have been studied intensively over the past decades, as has nitrogen fixation in marine and terrestrial habitats. However, the combination of both chemoautotrophy and nitrogen fixation in a microbial population or even in a single organism has rarely been considered. The recent discovery of diazotrophic Thiothrix species living as amphipod ectosymbionts in a sulfidic cave44 and the coexistence of chemoautotrophy and diazotrophy in the microbial community of cold water corals45, along with the results of this study, suggest that chemoautotrophic diazotrophs may be more widespread than previously anticipated. Other invertebrate models colonizing sulfidic environments may rely on chemoautotrophic symbionts that are also capable of diazotrophy.
Methods
Sampling
Adult individuals of C. orbicularis (Linné, 1758) were collected by hand at a depth of 5–10 cm in the sediment of T. testudinum seagrass beds in Guadeloupe (French West-Indies, Caribbean). For details of sampling sites, times and replicate numbers see Supplementary Table 1. Specimens designated for immediate dissection (for proteomics, genome sequencing, acetylene reduction assays, western blot and immunohistochemistry) were transported to the laboratory in seawater at ambient temperature within 1 h after collection. C. orbicularis individuals designated for incubation (starvation experiment, see below) were transported to the laboratory within 5–6 h. Sediment samples were collected off ‘Îlet cochon’, directly adjacent to C. orbicularis individuals (that is, not more than 20 cm away from the animals) using translucent polypropylene sediment corers (80 mm diameter). The cores (up to 16 cm in depth) were sliced at 4 cm intervals in the field, stored at 4 °C in an ice chest for transport to the laboratory and were subjected to pore water extraction within 1 h (see section ‘Chemical analysis of pore water samples’).
Incubation experiments with C. orbicularis
For incubation experiments, freshly collected C. orbicularis specimens were randomly separated into two batches (a control batch and a starvation batch). Specimens in the control batch were dissected immediately, while those in the starvation batch were kept in sterile (0.22 µm-filtered) sea water in 50 l plastic tanks at 26 °C for seven days. The water was oxygenated using an aquarium air pump. To simulate starvation conditions, no organic particles and no reduced sulfur compounds were added throughout the experiment. Bivalves were killed after one week of incubation, and symbionts were purified by density gradient centrifugation (see section ‘Symbiont enrichment’) and stored at −20 °C.
Chemical analysis of pore water samples
Nitrate, nitrite and ammonium concentrations in pore water from sediment cores were analysed as follows. Pore water was extracted from the sliced cores by filtration under vacuum using a 5 µm filter. Collected pore water samples were then passed through 0.45 µm membrane syringe filters, frozen immediately and stored frozen before colorimetric tests. Nitrate and nitrite concentrations were analysed according to the standard colorimetric Griess method46 with the use of vanadium as a reduction agent47, and ammonium was measured by the indophenol blue method48.
Symbiont enrichment
Bacterial symbionts were purified from healthy C. orbicularis gills with a light beige colour using Percoll density gradient centrifugation as described previously13,49. Briefly, after dissection of the animals, gill tissue was homogenized in a Dounce homogenizer in cold sterile seawater (salinity 35 g l−1) and the resulting homogenate was centrifuged at 30g for 1 min (4 °C) to remove crude tissue debris. The supernatant was centrifuged again (400g, 2 min, 4 °C) to pellet the bacterial cells, which were subsequently resuspended in sterile sea water and layered on top of a Percoll cushion (60% Percoll in imidazole-buffered saline, containing 490 mM NaCl, 30 mM MgSO4, 11 mM CaCl2, 3 mM KCl and 50 mM imidazole, pH 7.5). During centrifugation (4,000g, 10 min, 4 °C), the bacteria—because of their elemental sulfur inclusions—accumulate below the cushion, while host tissue fragments stay on top. The bacteria were collected, washed twice in sterile sea water, and stored at −20 °C.
Genome sequencing and genomic analysis
Symbionts for genome sequencing were isolated from one single C. orbicularis host individual immediately after collection of the animal (as described in section ‘Sampling’) in March 2012. Genomic DNA was isolated from the purified bacteria using the MasterPure DNA Purification Kit (Epicentre). The genome was sequenced using Illumina sequencing technology. A Nextera shot-gun library was generated for a 112 bp paired end (PE) sequencing run on a Genome Analyzer IIx. For an overview of all subsequent steps involved in assembly, binning and annotation, see Supplementary Fig. 1. Sequencing resulted in 2 × 6,108,111 raw reads, which were checked with FastQC version 0.11.3 (http://www.bioinformatics.babraham.ac.uk/projects/fastqc/) and subsequently trimmed with Trimmomatic version 0.35 (ref. 50) (settings: remove Nextera adapters and leading/trailing low quality bases, trim using MAXINFO:40:0.8 and keep sequences ≥36 nt). As preliminary assemblies had shown some eukaryotic contaminations in the sequenced data, trimmed reads (2 × 5,729,978 reads + 355,335 single reads) were assembled in a two-step process of binning and mapping to obtain a clean symbiont draft genome. First, Ray version 2.3.1 (ref. 51; k = 37) was used and the resulting 1,018 contigs were visually binned with VizBin (ref. 52; Bin ‘COS’: 83 contigs; Bin ‘Other’: 935 contigs). The trimmed reads used for assembly were then mapped against Bin COS with the help of BBSplit version 35.85. In a second step, SPAdes version 3.7.0 (ref. 53) used those mapped reads (2 × 5,131,806 reads + 336,019 single reads) to perform the final assembly (k = 21, 33, 55, 77, ‘careful’ option and minimum coverage of 30). This time, contigs were binned with MetaWatt version 3.5.2 (ref. 54), basically splitting contigs into bins with known and unknown taxon assignments. The final draft genome comprises 55 contigs (total length = 4,489,993 bp; largest contig = 297,909 bp; N50 = 164,003 bp), as evaluated by QUAST version 3.2 (ref. 55). The rapid annotation tool Prokka version 1.11 (ref. 56) was used for automatic open reading frame (ORF) calling and annotation.
Gene cluster comparison
All genes in the nif gene cluster of ‘Ca. T. endolucinida’ COS (CODIS_20190–20650, Fig. 1) were compared to the genomes of Allochromatium vinosum DSM 180 (NC_013851) and Sedimenticola thiotaurini SIP-G1 (NZ_CP011412) using Bl2seq (BLASTn, E-value 1e-5). Sequence data and the BLAST comparison files were drawn with the R package genoPlotR version 0.8.4 (ref. 57) and edited in Inkscape version 0.91. Blast results were automatically edited, so that short hits contained in longer hits were removed. They were further curated manually, so that each of the ‘Ca. T. endolucinida’ genes matched only one gene in its relatives.
Proteomic analyses
Mass spectrometry (MS) analyses of C. orbicularis symbiont proteins were performed in two parallel approaches: (1) in an identification-centred (that is, qualitative), gel-based approach and (2) in a quantitative gel-free approach (see section ‘Gel-free protein quantification’). Measurements for the identification-centred approach were performed in three biological replicates (that is, for symbionts of three individual C. orbicularis hosts) for (1) soluble symbiont proteins from freshly collected C. orbicularis, (2) symbiont membrane proteins from freshly collected hosts and (3) symbiont membrane proteins from C. orbicularis specimens collected after one week of starvation. For the quantitative approach, three measurements (three technical replicates) were acquired for each of the three biological replicates of (1) symbiont proteins from freshly sampled C. orbicularis specimens and (2) symbiont proteins from starved C. orbicularis after one week of starvation. For a detailed overview of replicate numbers in all MS measurements performed in this study, see Supplementary Table 2.
Protein extraction
Symbiont cell pellets for gel-based MS analysis (approach (1)) of soluble proteins and membrane proteins were resuspended in lysis buffer (10 mM Tris, 1 mM EDTA, Roche complete protease inhibitor cocktail) before cell disruption by sonication (3 × 20 s) under permanent cooling. Cell debris was removed by centrifugation (10 min, 4 °C, 12,000g for soluble protein extraction/8,000g for extraction of membrane proteins), leaving the protein raw extract in the supernatant. Raw extracts designated for the analysis of soluble proteins were subjected to acetone precipitation overnight (−20 °C), the resulting protein pellets were washed with ethanol, and soluble proteins were resuspended in 8 M urea/2 M thiourea solution. For membrane protein analysis, the protein raw extract was subjected to repeated ultracentrifugation and solubilization steps as described by Eymann et al.58, before solubilization of the enriched membrane protein fraction in 50 mM TEAB (triethylammonium bicarbonate) buffer. For quantitative gel-free proteome analysis (approach (2)), symbiont cell pellets were resuspended in 50 mM TEAB buffer and cells were disrupted by sonication (3 × 20 s). In either case, protein concentrations were determined using a Bradford assay59.
Gel-based protein identification
Aliquots of 20 µg soluble protein and 20 µg membrane protein fraction, respectively, were loaded onto 12% polyacrylamide mini gels and subjected to gel electrophoresis at 150 V for 1 h. After Coomassie staining, individual gel lanes were excised and divided into ten equally sized slices (subsamples) each. Gel pieces were destained (200 mM NH4HCO3, 30% acetonitrile) and dried before overnight digestion with trypsin solution (1 µg ml−1, sequencing grade, Promega) at 37 °C. Peptides were eluted from the gel pieces in a sonication bath and purified using ZipTips (Millipore) according to the manufacturer's recommendations. As described previously60, peptide mixes were subjected to separation using reversed-phase C18 column chromatography on a nanoACQUITY-UPLC system (Waters). Mass spectrometry (MS)- and tandem mass spectrometry (MS/MS) data were acquired using an online-coupled LTQ-Orbitrap mass spectrometer (Thermo Fisher Scientific). MS spectra were searched against a target–decoy protein sequence database including all sequences from the C. orbicularis symbiont database (see ‘Genome sequencing and genomic analysis’ section) and common laboratory contaminants using the SEQUEST Sorcerer platform (Sage-N). Scaffold (version 4.2, Proteome Software) was used to validate MS/MS-based peptide and protein identifications using the XCorr-based filter settings described previously61. Proteins were considered identified if they passed the filter criteria in at least one of the three biological replicates. Peptide false discovery rates (FDRs) were <2% and protein FDRs were 0.0% throughout all samples.
Gel-free protein quantification
As described by Muntel et al.62, 250 µg protein (in TEAB buffer) was treated with RapiGest SF Surfactant solution (Waters), before reduction with TCEP (tris(2-carboxyethyl)phosphine, final concentration 5 mM), alkylation with iodacetamide (final concentration 10 mM) and digestion with activated trypsin (5 h, 37 °C). After removal of RapiGest by acidification and repeated centrifugation, peptides were purified using StageTips (Proxeon) as suggested by the manufacturer. Tryptically digested bovine haemoglobin (Waters, MassPREP Bovine Hemoglobin Standard) was added to the protein samples as an internal peptide standard at a final concentration of 50 fmol µl−1. Samples were stored at −80 °C until MS analysis. Peptide mixtures were analysed using the nanoACQUITY-UPLC system (Waters), coupled to a Synapt G2 mass spectrometer (Waters), equipped with a NanoLockSpray source in positive mode. The analyser was set to resolution mode and operated with ion mobility separation as described by Huja and co-authors63. The ProteinLynx Global Server (PLGS) 2.5.3 was used for processing raw data and carrying out the database searches against the randomized C. orbicularis symbiont database with added common laboratory contaminants and the bovine haemoglobin sequence63. Proteins were considered identified if they were detected in one of the respective technical or biological replicates with a minimum of two peptides at a maximum FDR of 5% on the protein level. For quantification, only those proteins were included for which a minimum of two peptides were detected and which were identified in a minimum of two technical replicates in at least two biological replicates. For the proteins that met those criteria, average values of protein concentrations (in fmol ng−1) were calculated in two subsequent steps, that is, first across all available technical replicates for each biological replicate and subsequently across all biological replicates using the average values from step 1 (ref. 64). Protein FDRs were below 1%. To identify significant differences in protein expression between fresh control samples and samples obtained after starvation, mean protein abundances calculated across all three technical replicates of each biological replicate were used for statistical testing in TM4 MeV (TM4 Software Suite's Multi-Experiment Viewer; http://www.tm4.org/mev.html). After z-score transformation, an unpaired permutational t-test with Welch approximation (using all permutations, P value of 0.01, adjusted Bonferroni correction) was applied.
Microscopy and immunohistochemistry
For an overview and histological information about the C. orbicularis gill ultrastructure, a gill (dissected from one C. orbicularis individual) was fixed in Bouin's fluid for 24 h at room temperature and then embedded in Paraplast. Tissue sections (7 µm thick) were stained by Goldner's trichrome65 to allow for clear differentiation between granule cells and bacteriocytes.
For immunolocalization, gills of a freshly collected C. orbicularis specimen were dissected and fixed in 2% paraformaldehyde (in 0.22 µm-filtered seawater) and dehydrated in ethanol before being embedded in paraffin. Gill tissue of the asymbiotic bivalve Arcopagia crassa (familiy Tellinidae) was used as a negative control and treated accordingly. As an additional negative control, pieces of trophosome tissue of the vestimentiferan tube worm Riftia pachyptila were fixed in 4% paraformaldehyde immediately after collection (in November 2014, Supplementary Table 1) and stored in 70% ethanol before embedding in paraffin. Histological sections (7 µm thick) were deparaffinized before rehydration. After non-specific binding site saturation, sections were incubated with anti-NifH (nitrogenase iron protein) antibody from Agrisera, at a dilution of 1:500 in blocking solution containing 3% bovine serum albumin (BSA) in 1× PBS (phosphate-buffered saline) for 2 h at room temperature. Subsequently, sections were incubated in the secondary antibody, horseradish peroxidase-conjugated anti-hen IgY (Santa Cruz Biotechnology) at a 1:200 dilution in blocking solution for 1 h at room temperature. Slides were incubated in amplification buffer with Alexa Fluor 546, mounted with Vectashield Mounting Medium with 4′,6-diamidino-2-phenylindole (DAPI, VectorLabs) and observed under an epifluorescence microscope (Nikon Eclipse 80i).
Western blot analysis
Gill tissues from fresh C. orbicularis and A. crassa specimens (negative control) were crushed on ice before protein extraction in lysis buffer as described in the ‘Proteomic analyses’ section. The total protein raw extract was homogenized in Laemmli buffer (Sigma-Aldrich), and denatured at 95 °C for 10 min before storage at −20 °C. Proteins were separated on precast 4–12% NuPage polyacrylamide mini gels (Invitrogen) and blotted to PVDF membranes (Immobilon) for 1.5 h. The proteins were probed with a hen anti-NifH antibody (Agrisera) at a dilution of 1:10,000. Blots were incubated in the secondary antibody (horse radish peroxidase-conjugated anti-hen IgY) diluted to 1:50,000. HRP activity was detected using the ECL kit (GE Healthcare) according to the manufacturer's instructions and documented on Kodak X-OMAT autoradiography film (Fisher-Scientific). The specificity of the antibody raised against NifH was verified using C. orbicularis whole gill protein extracts.
Nitrogenase activity assay
Nitrogenase activity was tested in both dissected gills and in purified gill endosymbiont fractions obtained by the Percoll cushion method (see ‘Symbiont enrichment’ section). The isolated symbiont fractions contained ∼5 × 108 bacteria (per assay), while the number of bacterial cells in the gill samples was approximately 3 × 108 per assay (assuming a bacterial density of 106 bacteria per mg of gill tissue; 242–352 mg of gill tissue was used for each assay). Tissue and cell samples, respectively, were incubated in 22 ml glass tubes containing 2 ml of 0.22 µm-filtered sea water in contact with a headspace (20 ml) of air supplemented with 20 µmoles of acetylene (aerobic conditions at the beginning of the incubation). A parallel incubation of gill tissue and purified symbionts was started under low-oxygen conditions, for which air in the head space was replaced by O2-free argon and supplemented with the same quantity of acetylene. The samples were incubated at 24 °C and shaken continuously (60 r.p.m.) using a shaking water bath (SWB25 Thermo Scientific) throughout the experiment to equilibrate the gas and bacteria/tissue samples. After 24 h of incubation, ethylene production was measured as follows. A 500 µl gas sample was withdrawn from the headspace using a gas-tight hypodermic glass syringe (Interchim) and analysed using a CP-3800 gas chromatograph (Varian). The apparatus was equipped with a flame ionization detector (FID) for acetylene and ethylene detection, and thermal conductivity detectors (TCDs) for air nitrogen detection. Gas analysis was performed on a Rt-QS-Bond capillary PLOT fused-silica column (15 cm × 0.53 mm × 20 µm; Restek) under the following conditions: column input flow rate 9.1 ml min−1, oven temperature 40 °C, FID temperature 150 °C, helium as gas carrier (30 ml min−1 flow) and make-up (20 ml min−1 flow), hydrogen at 4.5 bar and air at 300 ml min−1 flow. The column retention time for acetylene and ethylene was monitored with standards throughout the experiments. Sea water and symbiont-free C. orbicularis foot tissue were assayed as negative controls, as was a sediment sample from the T. testudinum bed (larger seagrass roots were removed before analysis). In addition, gill and foot tissue samples, purified symbionts, sea water and sediment samples were incubated without acetylene and assayed for ethylene production to exclude false positive results. No ethylene production was detected in any of these samples incubated without acetylene.
Phylogenetic analysis
NifH phylogeny
The NifH tree was created in two consecutive steps. First, full-length NifH amino acid sequences of the C. orbicularis symbiont and of related cultured diazotrophic organisms were aligned and the phylogeny was inferred based on maximum likelihood. Second, partial NifH sequences of environmental clones from various marine habitats were added using ARB's interactive parsimony (see later in this paragraph). The NifH protein of ‘Ca. T. endolucinida’ was identified by searching the Pfam66 model Fer4_NifH (PF00142, available from http://pfam.xfam.org/family/PF00142/hmm) against all annotated proteins of the C. orbicularis symbiont's draft genome with hmmsearch (Expect value 1e-10) from HMMER version 3.1b2 (ref. 67). Its sequence was subsequently searched with HMMER's phmmer (Expect value 1e-10) against a local version of NCBI's nr database (version 2015-11-17), and the top 100 hits, that is, the 100 most closely related NifH sequences from other organisms, were retrieved. Additional (partial) NifH sequences of bacterial isolates from tropical seagrass beds42, oligotrophic open ocean68, salt marsh ecosystems69,70, mangrove root sediments71,72 and sulfidic cave water44 (1,007 sequences in total) were included. Some of these NifH sequences71 were only available as nucleotide sequences and were therefore translated using EMBOSS Transeq73 and the correct reading frame was selected manually. Four additional NifH protein sequences from known thiotrophs and two sequences from the outgroup taxa Anabaena variabilis ATCC 29413 (NifH: AAA93020) and Frankia alni ArI3 (NifH: AAA96262) complemented the data set, giving a total of 1,114 sequences for the NifH-based phylogenetic analysis. Sequence entries from the PDB and PRF databases as well as sequences containing stop codons or ambiguous amino acids (X) were removed, leaving 1,073 sequences that were clustered at 100% identity with CD-HIT (refs 74,75) to remove redundancies. All 834 representative non-redundant sequences (93 full length, 741 partial length) were automatically aligned with MAFFT version 7.221 (2014/04/16)76 using the L-INS-i algorithm and the ‘leave gappy region’ option. The alignment was imported into ARB version 5.5 (ref. 77) using a custom filter, and a protein database was created. A total of 55 full-length amino-acid sequences were selected and exported. ProtTest version 3 (refs 78,79) was employed to select the best-fit model of amino-acid replacement from 120 models (15 matrices: +G, +I or + G+I; +F) with a starting topology based on maximum likelihood. As the Bayesian information criterion (BIC; also known as the Schwarz criterion, or SC)80, the corrected Akaike information criterion (AICc)81,82 and the decision theory framework (DT)83 favoured LG+I+G, the phylogeny was then inferred using RAxML version 8.2.4 (ref. 84) with the LG amino acid matrix85, a gamma model of rate heterogeneity and an estimate of proportion of invariable sites. The best tree was chosen from 1,000 independent inferences that were executed on the original alignment (169 alignment patterns) using 1,000 distinct randomized maximum parsimony (MP) trees and was imported into ARB version 6.0.3. All 741 partial length NifH sequences were added by ARB's interactive parsimony. Sequences that could not be unambiguously inserted at a specific position in the tree were removed. To identify the partial length NifH sequences with the highest identity to the full-length sequences, a BLAST+ (ref. 86) protein database of all partial sequences was created and the full-length sequences that had partial sequences assigned by parsimony were searched against this database using BlastP version 2.2.31+. For each full-length sequence, only the best matching partial sequence (that is, the one with highest identity) was kept in the tree, and only if sequence identity between both was 95% or higher. The final tree (Fig. 5) thus comprises the C. orbicularis symbiont's NifH, 54 full-length NifH sequences of cultured organisms and 21 partial NifH sequences mostly from uncultured clones.
NifDK phylogeny
The NifD and NifK trees were constructed based on maximum likelihood as described above and include the respective amino acid sequences of the C. orbicularis symbiont and of the same cultured organisms that are displayed in the NifH tree, but with two exceptions: Methylogaea oryzae JCM 16910 is not part of the NifK tree and Agarivorans gilvus WH0801 could not be included in the NifD tree because the respective genes (locus tags JCM16910_RS02630 and AR383_RS01715) are disrupted. Anabaena variabilis ATCC 29413 (NifD: AAA93021; NifK: AAA93022) and Frankia alni ArI3 (NifD: AAA96263; NifK: AAA96264) were used as outgroup taxa. Note that the uncultured clones from seagrass and mangrove sediments (dotted branches in the NifH tree) are not included in the NifD and NifK trees, because no NifD or NifK sequence information was available for these isolates.
16S rRNA phylogeny
The 16S rRNA-coding region of the ‘Ca. T. endolucinida’ genome was identified using barrnap version 0.6 (available from https://github.com/tseemann/barrnap) and automatically aligned against the SILVA SSU Ref NR 99 Release 123 database (available from http://www.arb-silva.de)87 using the SILVA Incremental Aligner (SINA) version 1.2.11 (ref. 88). The SINA-aligned sequence was imported into the ARB software package version 5.5 (ref. 77). Besides the C. orbicularis symbiont's 16S ribosomal RNA nucleotide sequence and its closest homologues (by NCBI blastn), the data set contained additional 16S rRNA sequences of bivalve and gastropod symbionts, free-living sulfur oxidizers and some diazotrophic bacteria, whose NifH sequences were also included in the NifH tree. The alignment of these 58 sequences was manually refined taking into account the secondary structure information of the rRNA. Phylogenetic reconstruction was performed using a maximum likelihood method. The final tree was calculated with RAxML version 8.2.4 (GTRGAMMA model)84 and was based on an alignment with 750 distinct alignment patterns. The best tree from 1,000 independent inferences using 1,000 distinct randomized maximum parsimony trees is presented. Frankia alni ACN14a (CT573213) was used as outgroup.
Accession codes and data availability
Sequence data that support the findings of this study, that is, the ‘Ca. T. endolucinida’ Whole Genome Shotgun project, have been deposited at DDBJ/ENA/GenBank under project accession no. MARB00000000. The version described in this paper is version MARB01000000 (NCBI BioSample SAMN03435122, BioProject PRJNA284177). All nitrogen metabolism-related protein identifications obtained in this study are available in Supplementary Table 4. Additional data (for example, MS raw data) are available on request from the corresponding author (S.Ma.).
References
Cavanaugh, C. M., McKiness, Z. P., Newton, I. L. & Stewart, F. J. in The Prokaryotes: Symbiotic Associations, Biotechnology, Applied Microbiology Vol. 1 (ed. Dworkin, M. M. ) 475–507 (Springer, 2006).
Fisher, C. R. Chemoautotrophic and methanotrophic symbioses in marine invertebrates. Rev. Aquat. Sci. 2, 399–436 (1990).
Felbeck, H. & Distel, D. L. in The Prokaryotes Vol. 4 (eds Balows, A. et al.) 3891–3906 (Springer, 1992).
Taylor, J. D. & Glover, E. A. Lucinidae (Bivalvia)—the most diverse group of chemosymbiotic molluscs. Zool. J. Linn. Soc. 148, 421–438 (2006).
Abbott, R. T. American Seashells 2nd edn (Van Nostrand Reinhold, 1974).
Berg, C. J. & Alatalo, P. Potential of chemosynthesis in molluscan mariculture. Aquaculture 39, 165–179 (1984).
Gros, O., Darrasse, A., Durand, P., Frenkiel, L. & Moueza, M. Environmental transmission of a sulfur-oxidizing bacterial gill endosymbiont in the tropical lucinid bivalve Codakia orbicularis. Appl. Environ. Microbiol. 62, 2324–2330 (1996).
Gros, O., Frenkiel, L. & Mouëza, M. Embryonic, larval, and post-larval development in the symbiotic clam Codakia orbicularis (Bivalvia: Lucinidae). Invertebr. Biol. 116, 86–101 (1997).
Gros, O., Liberge, M., Heddi, A., Khatchadourian, C. & Felbeck, H. Detection of the free-living forms of sulfide-oxidizing gill endosymbionts in the lucinid habitat (Thalassia testudinum environment). Appl. Environ. Microbiol. 69, 6264–6267 (2003).
Caro, A., Got, P., Bouvy, M., Troussellier, M. & Gros, O. Effects of long-term starvation on a host bivalve (Codakia orbicularis, Lucinidae) and its symbiont population. Appl. Environ. Microbiol. 75, 3304–3313 (2009).
Brissac, T., Gros, O. & Merçot, H. Lack of endosymbiont release by two Lucinidae (Bivalvia) of the genus Codakia: consequences for symbiotic relationships. FEMS Microbiol. Ecol. 67, 261–267 (2009).
König, S., Le Guyader, H. & Gros, O. Thioautotrophic bacterial endosymbionts are degraded by enzymatic digestion during starvation: case study of two lucinids Codakia orbicularis and C. orbiculata. Microsc. Res. Tech. 78, 173–179 (2015).
Caro, A., Gros, O., Got, P., De Wit, R. & Troussellier, M. Characterization of the population of the sulfur-oxidizing symbiont of Codakia orbicularis (Bivalvia, Lucinidae) by single-cell analyses. Appl. Environ. Microbiol. 73, 2101–2109 (2007).
Welsh, D. T. Nitrogen fixation in seagrass meadows: regulation, plant–bacteria interactions and significance to primary productivity. Ecol. Lett. 3, 58–71 (2000).
Raymond, J., Siefert, J. L., Staples, C. R. & Blankenship, R. E. The natural history of nitrogen fixation. Mol. Biol. Evol. 21, 541–554 (2004).
Kneip, C., Lockhart, P., Voß, C. & Maier, U.-G. Nitrogen fixation in eukaryotes—new models for symbiosis. BMC Evol. Biol. 7, 55 (2007).
Pawlowski, K., Ribeiro, A. & Bisseling, T. in Biotechnol. Ann. Rev. Vol. 2 (ed. Raafat El-Gewely, M. ) 151–184 (Elsevier, 1996).
Santi, C., Bogusz, D. & Franche, C. Biological nitrogen fixation in non-legume plants. Ann. Bot. (Lond.) 111, 743–767 (2013).
Distel, D. L., Beaudoin, D. J. & Morrill, W. Coexistence of multiple proteobacterial endosymbionts in the gills of the wood-boring Bivalve Lyrodus pedicellatus (Bivalvia: Teredinidae). Appl. Environ. Microbiol. 68, 6292–6299 (2002).
Fiore, C. L., Jarett, J. K., Olson, N. D. & Lesser, M. P. Nitrogen fixation and nitrogen transformations in marine symbioses. Trends Microbiol. 18, 455–463 (2010).
Riemann, L., Farnelid, H. & Steward, G. F. Nitrogenase genes in non-cyanobacterial plankton: prevalence, diversity and regulation in marine waters. Aquat. Microb. Ecol. 61, 235–247 (2010).
Ingemann Jensen, S., Kühl, M. & Priemé, A. Different bacterial communities associated with the roots and bulk sediment of the seagrass Zostera marina. FEMS Microbiol. Ecol. 62, 108–117 (2007).
Crump, B. C. & Koch, E. W. Attached bacterial populations shared by four species of aquatic angiosperms. Appl. Environ. Microbiol. 74, 5948–5957 (2008).
Jørgensen, B. B. Mineralization of organic matter in the sea bed—the role of sulphate reduction. Nature 296, 643–645 (1982).
Küsel, K., Trinkwalter, T., Drake, H. L. & Devereux, R. Comparative evaluation of anaerobic bacterial communities associated with roots of submerged macrophytes growing in marine or brackish water sediments. J. Exp. Mar. Biol. Ecol. 337, 49–58 (2006).
van der Heide, T. et al. A three-stage symbiosis forms the foundation of seagrass ecosystems. Science 336, 1432–1434 (2012).
Merrick, M. J. & Edwards, R. A. Nitrogen control in bacteria. Microbiol. Rev. 59, 604–622 (1995).
Kurtz, D. M. Jr . Avoiding high-valent iron intermediates: superoxide reductase and rubrerythrin. J. Inorg. Biochem. 100, 679–693 (2006).
Zhao, W., Ye, Z. & Zhao, J. RbrA, a cyanobacterial rubrerythrin, functions as a FNR-dependent peroxidase in heterocysts in protection of nitrogenase from damage by hydrogen peroxide in Anabaena sp. PCC 7120. Mol. Microbiol. 66, 1219–1230 (2007).
Myrold, D. Quantification of Nitrogen Transformations 3rd edn (ASM, 2007).
Frenkiel, L. & Mouëza, M. Gill ultrastructure and symbiotic bacteria in Codakia orbicularis (Bivalvia, Lucinidae). Zoomorphology 115, 51–61 (1995).
Robidart, J. C. et al. Metabolic versatility of the Riftia pachyptila endosymbiont revealed through metagenomics. Environ. Microbiol. 10, 727–737 (2008).
Gardebrecht, A. et al. Physiological homogeneity among the endosymbionts of Riftia pachyptila and Tevnia jerichonana revealed by proteogenomics. ISME J. 6, 766–776 (2012).
Jørgensen, N. O. G., Kroer, N., Coffin, R. B., Yang, X.-H. & Lee, C. Dissolved free amino acids, combined amino acids, and DNA as sources of carbon and nitrogen to marine bacteria. Mar. Ecol. Prog. Ser. 98, 135–148 (1993).
Mulholland, M. R. & Lomas, M. W. in Nitrogen in the Marine Environment 2nd edn (eds Capone, D. G., Bronk, D. A., Mulholland, M. R. & Carpenter, E. J. ) Ch. 7, 303–384 (Academic, 2008).
Touchette, B. W. & Burkholder, J. M. Review of nitrogen and phosphorus metabolism in seagrasses. J. Exp. Mar. Biol. Ecol. 250, 133–167 (2000).
Fourqurean, J. W., Zieman, J. C. & Powell, G. V. N. Relationships between porewater nutrients and seagrasses in a subtropical carbonate environment. Mar. Biol. 114, 57–65 (1992).
Lee, K.-S. & Dunton, K. H. Inorganic nitrogen acquisition in the seagrass Thalassia testudinum: development of a whole-plant nitrogen budget. Limnol. Oceanogr. 44, 1204–1215 (1999).
Williams, S. L. Experimental studies of Caribbean seagrass bed development. Ecol. Monogr. 60, 449–469 (1990).
Masepohl, B. & Forchhammer, K. in Biology of the Nitrogen Cycle 131–145 (Elsevier, 2007).
Dixon, R. & Kahn, D. Genetic regulation of biological nitrogen fixation. Nat. Rev. Microbiol. 2, 621–631 (2004).
Bagwell, C. E. et al. Molecular diversity of diazotrophs in oligotrophic tropical seagrass bed communities. FEMS Microbiol. Ecol. 39, 113–119 (2002).
Green-García, A. M. & Engel, A. S. Bacterial diversity of siliciclastic sediments in a Thalassia testudinum meadow and the implications for Lucinisca nassula chemosymbiosis. Estuar. Coast. Shelf Sci. 112, 153–161 (2012).
Desai, M. S., Assig, K. & Dattagupta, S. Nitrogen fixation in distinct microbial niches within a chemoautotrophy-driven cave ecosystem. ISME J. 7, 2411–2423 (2013).
Middelburg, J. J. et al. Discovery of symbiotic nitrogen fixation and chemoautotrophy in cold-water corals. Sci. Rep. 5, 17962 (2015).
Green, L. C. et al. Analysis of nitrate, nitrite, and [15N]nitrate in biological fluids. Anal. Biochem. 126, 131–138 (1982).
Schnetger, B. & Lehners, C. Determination of nitrate plus nitrite in small volume marine water samples using vanadium(III) chloride as a reduction agent. Mar. Chem. 160, 91–98 (2014).
Solórzano, L. Determination of ammonia in natural waters by the phenolhypochlorite method. Limnol. Oceanogr. 14, 799–801 (1969).
Elisabeth, N. H. et al. Comparative modifications in bacterial gill–endosymbiotic populations of the two bivalves Codakia orbiculata and Lucina pensylvanica during bacterial loss and reacquisition. FEMS Microbiol. Ecol. 89, 646–658 (2014).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Boisvert, S., Laviolette, F. & Corbeil, J. Ray: simultaneous assembly of reads from a mix of high-throughput sequencing technologies. J. Comput. Biol. 17, 1519–1533 (2010).
Laczny, C. C. et al. VizBin—an application for reference-independent visualization and human-augmented binning of metagenomic data. Microbiome 3, 1 (2015).
Bankevich, A. et al. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 19, 455–477 (2012).
Strous, M., Kraft, B., Bisdorf, R. & Tegetmeyer, H. E. The binning of metagenomic contigs for microbial physiology of mixed cultures. Front. Microbiol. 3, 410 (2012).
Gurevich, A., Saveliev, V., Vyahhi, N. & Tesler, G. QUAST: quality assessment tool for genome assemblies. Bioinformatics 29, 1072–1075 (2013).
Seemann, T. Prokka: rapid prokaryotic genome annotation. Bioinformatics 30, 2068–2069 (2014).
Guy, L., Roat Kultima, J. & Andersson, S. G. E. genoPlotR: comparative gene and genome visualization in R. Bioinformatics 26, 2334–2335 (2010).
Eymann, C. et al. A comprehensive proteome map of growing Bacillus subtilis cells. Proteomics 4, 2849–2876 (2004).
Bradford, M. M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein–dye binding. Anal. Biochem. 72, 248–254 (1976).
Xing, P. et al. Niches of two polysaccharide-degrading Polaribacter isolates from the North Sea during a spring diatom bloom. ISME J. 9, 1410–1422 (2015).
Heinz, E. et al. The genome of the obligate intracellular parasite Trachipleistophora hominis: new insights into microsporidian genome dynamics and reductive evolution. PLoS Pathog. 8, e1002979 (2012).
Muntel, J., Hecker, M. & Becher, D. An exclusion list based label-free proteome quantification approach using an LTQ Orbitrap. Rapid Commun. Mass Spectrom. 26, 701–709 (2012).
Huja, S. et al. Fur is the master regulator of the extraintestinal pathogenic Escherichia coli response to serum. mBio 5, e01460 (2014).
Silva, J. C., Gorenstein, M. V., Li, G.-Z., Vissers, J. P. C. & Geromanos, S. J. Absolute quantification of proteins by LCMSE: a virtue of parallel MS acquisition. Mol. Cell. Proteomics 5, 144–156 (2006).
Luna, L. G. Histopathologic Methods and Color Atlas of Special Stains and Tissue Artifacts 1st edn (American Histolabs, 1992).
Finn, R. D. et al. The Pfam protein families database: towards a more sustainable future. Nucleic Acids Res. 44, D279–D285 (2016).
Eddy, S. R. Accelerated profile HMM searches. PLoS Comp. Biol. 7, e1002195 (2011).
Zehr, J. P., Mellon, M. T. & Zani, S. New nitrogen-fixing microorganisms detected in oligotrophic oceans by amplification of nitrogenase (nifH) genes. Appl. Environ. Microbiol. 64, 3444–3450 (1998).
Lovell, C. R., Piceno, Y. M., Quattro, J. M. & Bagwell, C. E. Molecular analysis of diazotroph diversity in the rhizosphere of the smooth cordgrass, Spartina alterniflora. Appl. Environ. Microbiol. 66, 3814–3822 (2000).
Lovell, C. R. & Davis, D. A. Specificity of salt marsh diazotrophs for vegetation zones and plant hosts: results from a North American marsh. Front. Microbiol. 3, 84 (2012).
Flores-Mireles, A. L., Winans, S. C. & Holguin, G. Molecular characterization of diazotrophic and denitrifying bacteria associated with mangrove roots. Appl. Environ. Microbiol. 73, 7308–7321 (2007).
Dias, A. C. F. et al. Abundance and genetic diversity of nifH gene sequences in anthropogenically affected Brazilian mangrove sediments. Appl. Environ. Microbiol. 78, 7960–7967 (2012).
Rice, P., Longden, I. & Bleasby, A. EMBOSS: The European Molecular Biology Open Software Suite. Trends Genet. 16, 276–277 (2000).
Li, W. & Godzik, A. CD-HIT: a fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics 22, 1658–1659 (2006).
Fu, L., Niu, B., Zhu, Z., Wu, S. & Li, W. CD-HIT: accelerated for clustering the next-generation sequencing data. Bioinformatics 28, 3150–3152 (2012).
Katoh, K., Misawa, K., Kuma, K.-i. & Miyata, T. MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res. 30, 3059–3066 (2002).
Ludwig, W. et al. ARB: a software environment for sequence data. Nucleic Acids Res. 32, 1363–1371 (2004).
Guindon, S. & Gascuel, O. A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst. Biol. 52, 696–704 (2003).
Darriba, D., Taboada, G. L., Doallo, R. & Posada, D. ProtTest 3: fast selection of best-fit models of protein evolution. Bioinformatics 27, 1164–1165 (2011).
Schwarz, G. Estimating the dimension of a model. Ann. Stat. 6, 461–464 (1978).
Sugiura, N. Further analysts of the data by Akaike's information criterion and the finite corrections. Commun. Stat. A 7, 13–26 (1978).
Hurvich, C. M. & Tsai, C.-L. Regression and time series model selection in small samples. Biometrika 76, 297–307 (1989).
Minin, V., Abdo, Z., Joyce, P. & Sullivan, J. Performance-based selection of likelihood models for phylogeny estimation. Syst. Biol. 52, 674–683 (2003).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Le, S. Q. & Gascuel, O. An improved general amino acid replacement matrix. Mol. Biol. Evol. 25, 1307–1320 (2008).
Camacho, C. et al. BLAST+: architecture and applications. BMC Bioinformatics 10, 421 (2009).
Yilmaz, P. et al. The SILVA and ‘All-species Living Tree Project (LTP)’ taxonomic frameworks. Nucleic Acids Res. 42, D643–D648 (2013).
Pruesse, E., Peplies, J. & Glöckner, F. O. SINA: accurate high-throughput multiple sequence alignment of ribosomal RNA genes. Bioinformatics 28, 1823–1829 (2012).
Acknowledgements
The authors acknowledge the technical assistance of J. Matulla and S. Grund and thank K. Riedel, L. Westhoff and M. Bengtson for discussions. This work was supported by the EU in the framework of the Marie Curie ITN ‘Symbiomics’ project (no. 264774) and by the German Science Foundation (DFG project no. MA6346/2-1). O.G.'s group was partially supported by the French National Program EC2CO Biohefect/Ecodyn/Dril/MicrobiEn ‘Fixation de l'azote chez les bivalves marins de la famille des Lucinidae: impact sur le fonctionnement de ces organismes et des écosystèmes tropicaux côtiers associés’.
Author information
Authors and Affiliations
Contributions
S.K. and O.G. planned and performed the experiments with C. orbicularis. Sediment chemistry analyses were conducted by J.T., western blots and immunohistochemistry by M.V., and acetylene reduction measurements by O.G. and D.M.-M. Proteome analyses were planned by S.K., T.S., S.Ma. and D.B., and conducted by S.K., S.Me. and D.B. A.T., A.P. and R.D. performed the genome sequencing and submitted the sequences to NCBI. S.K. and S.E.H. assembled and annotated the genome. S.E.H. performed the phylogenetic analyses and gene cluster comparisons. O.G., T.S. and S.Ma. coordinated and supervised the laboratory work. S.K. and S.Ma. analysed the proteome data. S.Ma. wrote the manuscript. T.H. was involved in manuscript writing, figure generation and, together with R.P., in MS data evaluation.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary information
Supplementary Results and Discussion, Supplementary References, Supplementary Tables 1–3, legends for Supplementary Tables 4 and 5, Supplementary Figures 1–6 (PDF 58519 kb)
Supplementary Table 4
Nitrogen-metabolism-related proteins identified by proteome analyses in this study (XLSX 61 kb)
Supplementary Table 5
Alignment scores of C. orbicularis symbiont NifH in BlastP against partial sequences of sediment clones and full-length sequences of cultured organisms. (XLSX 20 kb)
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
König, S., Gros, O., Heiden, S. et al. Nitrogen fixation in a chemoautotrophic lucinid symbiosis. Nat Microbiol 2, 16193 (2017). https://doi.org/10.1038/nmicrobiol.2016.193
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/nmicrobiol.2016.193
This article is cited by
-
Sulfur oxidation and reduction are coupled to nitrogen fixation in the roots of the salt marsh foundation plant Spartina alterniflora
Nature Communications (2024)
-
Gene loss and symbiont switching during adaptation to the deep sea in a globally distributed symbiosis
The ISME Journal (2023)
-
Chemoautotrophy, symbiosis and sedimented diatoms support high biomass of benthic molluscs in the Namibian shelf
Scientific Reports (2022)
-
Partitioning benthic nitrogen cycle processes among three common macrofauna holobionts
Biogeochemistry (2022)
-
Metagenomic analysis suggests broad metabolic potential in extracellular symbionts of the bivalve Thyasira cf. gouldi
Animal Microbiome (2020)