Abstract
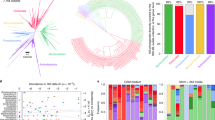
Genome-enabled technologies have supported a dramatic increase in our ability to study microbial communities in environments and hosts. Taking stock of previously funded microbiome research can help to identify common themes, under-represented areas and research priorities to consider moving forward. To assess the status of US microbiome research, a team of government scientists conducted an analysis of federally funded microbiome research. Microbiomes were defined as host-, ecosystem- or habitat-associated communities of microorganisms, and microbiome research was defined as those studies that emphasize community-level analyses using ’omics technologies. Single pathogen, single strain and culture-based studies were not included, except symbiosis studies that served as models for more complex communities. Fourteen governmental organizations participated in the data call. The analysis examined three broad research themes, eight environments and eight microbial categories. Human microbiome research was larger than any other environment studied, and the basic biology research theme accounted for half of the total research activities. Computational biology and bioinformatics, reference databases and biorepositories, standardized protocols and high-throughput tools were commonly identified needs. Longitudinal and functional studies and interdisciplinary research were also identified as needs. This study has implications for the funding of future microbiome research, not only in the United States but beyond.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Breitbart, M. et al. Genomic analysis of uncultured marine viral communities. Proc. Natl Acad. Sci. USA 99, 14250–14255 (2002).
Venter, J. C. et al. Environmental genome shotgun sequencing of the Sargasso Sea. Science 304, 66–74 (2004).
Gill, S. R. et al. Metagenomic analysis of the human distal gut microbiome. Science 312, 1355–1359 (2006).
Fast-Track Action Committee on Mapping the Microbiome (National Science and Technology Council, 2015); https://www.whitehouse.gov/sites/default/files/microsites/ostp/NSTC/ftac-mm_report_final_112015_0.pdf
Schenk, P. M., Carvalhais, L. C. & Kazan, K. Unraveling plant–microbe interactions: can multi-species transcriptomics help? Trends Biotechnol. 30, 177–184 (2012).
Bakker, M. G., Schlatter, D. C., Otto-Hansen, L. & Kinkel, L. L. Diffuse symbioses: roles of plant–plant, plant–microbe and microbe–microbe interactions in structuring the soil microbiome. Mol. Ecol. 23, 1571–1583 (2014).
Virgin, H. W. The virome in mammalian physiology and disease. Cell 157, 142–150 (2014).
Brum, J. R. & Sullivan, M. B. Rising to the challenge: accelerated pace of discovery transforms marine virology. Nature Rev. Microbiol. 13, 147–159 (2015).
Roosinck, M. J. Move over bacteria! Viruses make their mark as mutualistic microbial symbionts. J. Virol. 89, 6532–6535 (2015).
Hewson, I. et al. Densovirus associated with sea-star wasting disease and mass mortality. Proc. Natl Acad. Sci. USA 111, 17278–17283 (2014).
Srinivasiah, S. et al. Dynamics of autochthonous soil viral communities parallels dynamics of host communities under nutrient stimulation. FEMS Micro. Ecol. http://dx.doi.org/10.1093/femsec/fiv063 (2015).
Wu, Z. et al. Deciphering the bat virome catalog to better understand the ecological diversity of bat viruses and the bat origin of emerging infectious diseases. ISME J. http://dx.doi.org/10.1038/ismej.2015.138 (2015).
Forsberg, K. J. et al. The shared antibiotic resistome of soil bacteria and human pathogens. Science 337, 1107–1111 (2012).
Jeng, R. R. et al. Regulation of intestinal inflammation by microbiota following allogeneic bone marrow transplantation. J. Exp. Med. 209, 903–911 (2012).
Vossen, J. M. et al. Complete suppression of the gut microbiome prevents acute graft-versus-host disease following allogenic bone marrow transplantation. PLoS ONE 9, e105706 (2014).
Salas, J. T. & Chang, T. L. Microbiome in human immunodeficiency virus infection. Clin. Lab. Med. 34, 733–745 (2014).
Bulgarelli, G. et al. Structure and functions of the bacterial microbiota of plants. Annu. Rev. Plant Biol. 64, 807–838 (2013).
Ramirez, K. M. et al. Towards a global platform for linking soil biodiversity data. Front. Ecol. Evol. 3, 91 (2015).
Batterman, S. A. et al. Key role of symbiotic N2 fixation in tropical forest secondary succession. Nature 502, 224–227 (2013).
Jansson, J. & Tas, N. The microbial ecology of permafrost. Nature Rev. Microbiol. 12, 414–425 (2014).
Karhu, K. et al. Temperature sensitivity of soil respiration rates enhanced by microbial community response. Nature 513, 81–84 (2014).
Alivisatos, A. P. et al. A unified initiative to harness Earth's microbiomes. Science 350, 507–508 (2015).
The NIH Human Microbiome Project, phase one; www.commonfund.nih.gov/hmp/
The EC MetaHIT programme; http://www.metahit.eu/
The Canadian Human Microbiome Initiative; http://www.cihr-irsc.gc.ca/e/38534.html
The Irish ELDERMET programme; http://eldermet.ucc.ie/
Mullard, A. Microbiology: the inside story. Nature 453, 578–579 (2008).
The International Human Microbiome Consortium; www.human-microbiome.org
The NIH Human Microbiome Project, phase two; http://ihmpdcc.org/
The EC ‘My New Gut’ programme; http://cordis.europa.eu/project/rcn/111044_en.html
The Irish APC Microbiome Institute; http://www.sfi.ie/assets/media/files/downloads/Investments/APC.pdf
The EC ‘Metagenopolis’ programme; http://mgps.eu/index.php?id=accueil
The Canadian ‘Environment, Gene and Chronic Diseases’ programme; https://www.researchnet-recherchenet.ca/rnr16/vwOpprtntyDtls.do?prog=2176&resultCount=25&sponsor=CIHR-10&type=EXACT&view=browseActive&language=E
The Canadian ‘Natural Resources and the Environment: Sector Challenges – Genomic Solutions’ programme; http://www.genomecanada.ca/en/portfolio/research/2015-competition.aspx
The EC ‘Joint Action Intestinal Microbiomics’ programme; https://www.healthydietforhealthylife.eu/index.php/joint-actions/microbiomics
The NSF Microbial Observatories and Microbial Interactions and Processes programme; https://www.nsf.gov/funding/pgm_summ.jsp?pims_id=6166
The NSF Microbial Genome Sequencing Program; http://www.nsf.gov/funding/pgm_summ.jsp?pims_id=5688
The International Census on Marine Microbes; http://icomm.mbl.edu/
The Gordon and Betty Moore Foundation Marine Microbiology Initiative; https://www.moore.org/programs/science/marine-microbiology-initiative
TerraGenome, the International Soil Metagenome Sequencing Consortium; http://www.terragenome.org/
The DOE Microbial Carbon Cycle programme; http://genomicscience.energy.gov/carboncycle/index.shtml
Gilbert, J. A., Jansson, J. K. & Knight, R. The Earth Microbiome Project: successes and aspirations. BMC Biol. 12, 69 (2014).
Kopf, A. et al. The ocean sampling day consortium. GigaScience 4, 66 (2015).
Acknowledgements
The Committee acknowledges the efforts of colleagues in our respective agencies who worked to meet the six-week data call deadline. The authors acknowledge the earlier efforts of the trans-NIH Microbiome Working Group (TMWG), whose FY10-12 portfolio analysis formed the basis of the approach and format for this data call.
Author information
Authors and Affiliations
Contributions
E.S., D.F., L.P. and D.M. conceived the study. All authors jointly planned the data collection effort. All authors except E.S. and J.L. managed the data collection at their agency or department and provided their data to the central database. E.S. and J.L. conducted the analysis, and L.P. and E.S. prepared the initial draft of the manuscript. All authors revised the manuscript and agreed on the final version.
Corresponding author
Ethics declarations
Competing interests
All authors are federal employees and the preparation of this manuscript was done as part of their official duties.
Supplementary information
Supplementary Data 1
An interactive version of the FTACMM Data Call spreadsheet (XLSX 118 kb)
Supplementary Data 2
The logic flow for the FTACMM drop down menu. (XLSX 10 kb)
Rights and permissions
About this article
Cite this article
Stulberg, E., Fravel, D., Proctor, L. et al. An assessment of US microbiome research. Nat Microbiol 1, 15015 (2016). https://doi.org/10.1038/nmicrobiol.2015.15
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/nmicrobiol.2015.15
This article is cited by
-
Einfluss von Konservierungsmitteln in Topika auf die kutane Mikrobiota
Die Dermatologie (2023)
-
Network structure of resource use and niche overlap within the endophytic microbiome
The ISME Journal (2022)
-
A widespread response of Gram-negative bacterial acyl-homoserine lactone receptors to Gram-positive Streptomyces γ-butyrolactone signaling molecules
Science China Life Sciences (2021)
-
Validation and standardization of DNA extraction and library construction methods for metagenomics-based human fecal microbiome measurements
Microbiome (2021)
-
Swabbing Thoroughly People for COVID-19 Positivity. Insights on the Main Bio-analytical and Microbiology Bias and Concerns
Current Microbiology (2020)