Abstract
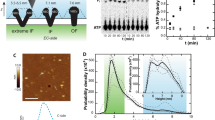
Biological membranes compartmentalize and define physical borders of cells. They are crowded with membrane proteins that fulfill diverse crucial functions. About one-third of all genes in organisms code for, and the majority of drugs target, membrane proteins. To combine structure and function analysis of membrane proteins, we designed a two-chamber atomic force microscopy (AFM) setup that allows investigation of membranes spanned over nanowells, therefore separating two aqueous chambers. We imaged nonsupported surface layers (S layers) of Corynebacterium glutamicum at sufficient resolution to delineate a 15 Å–wide protein pore. We probed the elastic and yield moduli of nonsupported membranes, giving access to the lateral interaction energy between proteins. We combined AFM and fluorescence microscopy to demonstrate the functionality of proteins in the setup by documenting proton pumping by Halobacterium salinarium purple membranes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Neher, E. & Sakmann, B. Single-channel currents recorded from membrane of denervated frog muscle fibres. Nature 260, 799–802 (1976).
Mueller, P., Rudin, D.O., Tien, H.T. & Wescott, W.C. Reconstitution of cell membrane structure in vitro and its transformation into an excitable system. Nature 194, 979–980 (1962).
Huang, C. Studies on phosphatidylcholine vesicles. Formation and physical characteristics. Biochemistry 8, 344–352 (1969).
Caffrey, M. Membrane protein crystallization. J. Struct. Biol. 142, 108–132 (2003).
Fujiyoshi, Y. The structural study of membrane proteins by electron crystallography. Adv. Biophys. 35, 25–80 (1998).
Frank, J. Single-particle imaging of macromolecules by cryo-electron microscopy. Annu. Rev. Biophys. Biomol. Struct. 31, 303–319 (2002).
Binnig, G., Quate, C.F. & Gerber, C. Atomic force microscope. Phys. Rev. Lett. 56, 930–933 (1986).
Müller, D.J., Fotiadis, D., Scheuring, S., Müller, S.A. & Engel, A. Electrostatically balanced subnanometer imaging of biological specimens by atomic force microscopy. Biophys. J. 76, 1101–1111 (1999).
Schabert, F.A., Henn, C. & Engel, A. Native Escherichia coli OmpF porin surfaces probed by atomic force microscopy. Science 268, 92–94 (1995).
Fotiadis, D. et al. Atomic-force microscopy: Rhodopsin dimers in native disc membranes. Nature 421, 127–128 (2003).
Scheuring, S. & Sturgis, J.N. Chromatic adaptation of photosynthetic membranes. Science 309, 484–487 (2005).
Oesterhelt, D. & Stoeckenius, W. Functions of a new photoreceptor membrane. Proc. Natl. Acad. Sci. USA 70, 2853–2857 (1973).
Danelon, C., Perez, J.P., Santschi, C., Brugger, J. & Vogel, H. Cell membranes suspended across nanoaperture arrays. Langmuir 22, 22–25 (2006).
Chami, M. et al. Organization of the outer layers of the cell envelope of Corynebacterium glutamicum: A combined freeze-etch electron microscopy and biochemical study. Biol. Cell 83, 219–229 (1995).
Scheuring, S. et al. Charting and unzipping the surface-layer of Corynebacterium glutamicum with the atomic force microscope. Mol. Microbiol. 44, 675–684 (2002).
Sleytr, U.B. Basic and applied S-layer research: an overview. FEMS Microbiol. Rev. 20, 5–12 (1997).
Scheuring, S., Levy, D. & Rigaud, J-L. Watching the components of photosynthetic bacterial membranes and their “in situ” organization by atomic force microscopy. Biochim. Biophys. Acta 1712, 109–127 (2005).
Chang, G., Spencer, R.H., Lee, A.T., Barclay, M.T. & Rees, D.C. Structure of the MscL homolog from Mycobacterium tuberculosis: a gated mechanosensitive ion channel. Science 282, 2220–2226 (1998).
Landau, L.D. & Lifschitz, E.M. Theory of elasticity (Pergamon Press, New York, 1986).
Yao, X., Jericho, M.H., Pink, D. & Beveridge, T.J. Thickness and elasticity of gram-negative murein sacculi measured by atomic force microscopy. J. Bacteriol. 181, 6865–6875 (1999).
Evans, E. & Rawics, W. Entropy-driven tension and bending elasticity in condensed-fluid membranes. Phys. Rev. Lett. 64, 2094–2097 (1990).
Kano, K. & Fendler, J.H. Pyranine as a sensitive pH probe for liposome interiors and surfaces. pH gradients across phospholipid vesicles. Biochim. Biophys. Acta 509, 289–299 (1978).
Seigneuret, M. & Rigaud, J.L. Use of fluorescent pH probe pyranine to detect heterogeneous directions of proton movement in bacteriorhodopsin reconsituted large liposomes. FEBS Lett. 188, 101–106 (1985).
Stock, D., Leslie, A.G. & Walker, J.E. Molecular architecture of the rotary motor in ATP synthase. Science 286, 1700–1705 (1999).
Oesterhelt, F. et al. Unfolding pathways of individual bacteriorhodopsins. Science 288, 143–146 (2000).
Müller, D.J. et al. Observing membrane protein diffusion at subnanometer resolution. J. Mol. Biol. 327, 925–930 (2003).
Stahlberg, H. et al. Two-dimensional crystals: a powerful approach to assess structure, function and dynamics of membrane proteins. FEBS Lett. 504, 166–172 (2001).
Viani, M.B. et al. Probing protein-protein interactions in real time. Nat. Struct. Biol. 7, 644–647 (2000).
Seifert, U. Dynamics of a bound membrane. Phys. Rev. E Stat. Phys. Plasmas Fluids Relat. Interdiscip. Topics 49, 3124–3127 (1994).
Sorzano, C.O. et al. XMIPP: a new generation of an open-source image processing package for electron microscopy. J. Struct. Biol. 148, 194–204 (2004).
Acknowledgements
We thank F. Oesterhelt for assistance in force curve analysis, A. Martinez-Gil for technical assistance, C. Gueudry for help with fluorescence microscopy and S. Lesko for spring-constant determination. This study was supported by the INSERM and INSERM Avenir, a 'Ministère de l'Education Nationale' scholarship (to R.P.G.) and an Action Concertée Incitative Nanosciences 2004 grant (NR206).
Author information
Authors and Affiliations
Contributions
R.P.G. performed AFM imaging, S-layer preparation with C.H., fluorescence imaging and fluorescence data analysis. G.A. and B.B. prepared the nanopatterned Si(001) surfaces. P.S. and S.S. performed force-curve analysis and physical interpretation. S.S performed image and data analysis, conceived the project and prepared the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Schematic representation of the 2-chamber AFM setup. (PDF 663 kb)
Supplementary Fig. 2
AFM characterization of the holey Si(001) support wafers. (PDF 1512 kb)
Supplementary Video 1
Proton pumping of non-supported purple membranes. Sequence of green fluorescence images over a time-range of 500 seconds documenting fluorescence intensity variations in chambers. (MOV 467 kb)
Supplementary Note
Digital image treatment (PDF 48 kb)
Rights and permissions
About this article
Cite this article
Gonçalves, R., Agnus, G., Sens, P. et al. Two-chamber AFM: probing membrane proteins separating two aqueous compartments. Nat Methods 3, 1007–1012 (2006). https://doi.org/10.1038/nmeth965
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth965
This article is cited by
-
Tuning membrane protein mobility by confinement into nanodomains
Nature Nanotechnology (2017)
-
Glass is a Viable Substrate for Precision Force Microscopy of Membrane Proteins
Scientific Reports (2015)
-
Structure and Permeability of Ion-channels by Integrated AFM and Waveguide TIRF Microscopy
Scientific Reports (2014)
-
Investigation on the folding mode of a polymer chain in a spiral crystal by single molecule force spectroscopy
Chinese Journal of Polymer Science (2014)
-
Atomic force microscopy of model lipid membranes
Analytical and Bioanalytical Chemistry (2013)